I continue with my series on first exploratory steps with CNNs. After all the discussion of CNN basics in the last article,
A simple CNN for the MNIST datasets – I,
we are well prepared to build a very simple CNN with Keras. By simple I mean simple enough to handle the MNIST digit images. The Keras API for creating CNN models, layers and activation functions is very convenient; a simple CNN does not require much code. So, the Jupyter environment is sufficient for our first experiment.
An interesting topic is the use of a GPU. After a somewhat frustrating experience with a MLP on the GPU of a NV 960 GTX in comparison to a i7 6700K CPU I am eager to see whether we get a reasonable GPU acceleration for a CNN. So, we should prepare our code to use the GPU. This requires a bit of preparation.
We should also ask a subtle question: What do we expect from a CNN in comparison to a MLP regarding the MNIST data? A MLP with 2 hidden layers (with 70 and 30 nodes) provided over 99.5% accuracy on the training data and almost 98% accuracy on a test dataset after some tweaking. Even with basic settings for our MLP we arrived at a value over 97.7% after 35 epochs – below 8 secs. Well, a CNN is probably better in feature recognition than a cluster detection algorithm. But we are talking about the last 2 % of remaining accuracy. I admit that I did not know what to expect …
A MLP as an important part of a CNN
At the end of the last article I had discussed a very simple layer structure of convolutional and pooling layers:
- Layer 0: Input layer (tensor of original image data, 3 layers for color channels or one layer for a gray channel)
- Layer 1: Conv layer (small 3×3 kernel, stride 1, 32 filters => 32 maps (26×26), overlapping filter areas)
- Layer 2: Pooling layer (2×2 max pooling => 32 (13×13) maps,
a map node covers 4×4 non overlapping areas per node on the original image) - Layer 3: Conv layer (3×3 kernel, stride 1, 64 filters => 64 maps (11×11),
a map node covers 8×8 overlapping areas on the original image (total effective stride 2)) - Layer 4: Pooling layer (2×2 max pooling => 64 maps (5×5),
a map node covers 10×10 areas per node on the original image (total effective stride 5), some border info lost) - Layer 5: Conv layer (3×3 kernel, stride 1, 64 filters => 64 maps (3×3),
a map node covers 18×18 areas per node (effective stride 5), some border info lost )
This is the CNN structure we are going to use in the near future. (Actually, I followed a suggestion of Francois Chollet; see the literature list in the last article). Let us assume that we somehow have established all these convolution and pooling layers for a CNN. Each layer producse some “feature“-related output, structured in form of a tensors. This led to an open question at the end of the last article:
Where and by what do we get a classification of the resulting data with respect to the 10 digit categories of the MNIST images?
Applying filters and extracting “feature hierarchies” of an image alone does not help without a “learned” judgement about these data. But the answer is very simple:
Use a MLP after the last Conv layer and feed it with data from this Conv layer!
When we think in terms of nodes and artificial neurons, we could say: We just have to connect the “nodes” of the feature maps of layer 5
in our special CNN with the nodes of an input layer of a MLP. As a MLP has a flat input layer we need to prepare 9×64 = 576 receiving “nodes” there. We would use weights with a value of “1.0” along these special connections.
Mathematically, this approach can be expressed in terms of a “flattening” operation on the tensor data produced by the the last Conv data. In Numpy terms: We need to reshape the multidimensional tensor containing the values across the stack of maps at the last Conv2D layer into a long 1D array (= a vector).
From a more general perspective we could say: Feeding the output of the Conv part of our CNN into a MLP for classification is quite similar to what we did when we pre-processed the MNIST data by an unsupervised cluster detection algorithm; also there we used the resulting data as input to an MLP. There is one big difference, however:
The optimization of the network’s weights during training requires a BW propagation of error terms (more precisely: derivatives of the CNN’s loss function) across the MLP AND the convolutional and pooling layers. Error BW propagation should not be stopped at the MLP’s input layer: It has to move from the output layer of the MLP back to the MLP’s input layer and from there to the convolutional and pooling layers. Remember that suitable filter kernels have to be found during (supervised) training.
If you read my PDF on the error back propagation for a MLP
PDF on the math behind Error Back_Propagation
and think a bit about its basic recipes and results you quickly see that the “input layer” of the MLP is no barrier to error back propagation: The “deltas” discussed in the PDF can be back-propagated right down to the MLP’s input layer. Then we just apply the chain rule again. The partial derivatives at the nodes of the input layer with respect to their input values are just “1”, because the activation function there is the identity function. The “weights” between the last Conv layer and the input layer of the MLP are no free parameters – we do not need to care about them. And then everything goes its normal way – we apply chain rule after chain rule for all nodes of the maps to determine the gradients of the CNN’s loss function with respect to the weights there. But you need not think about the details – Keras and TF2 will take proper care about everything …
But, you should always keep the following in mind: Whatever we discuss in terms of layers and nodes – in a CNN these are only fictitious interpretations of a series of mathematical operations on tensor data. Not less, not more ..,. Nodes and layers are just very helpful (!) illustrations of non-cyclic graphs of mathematical operations. KI on the level of my present discussion (MLPs, CNNs) “just” corresponds to algorithms which emerge out of a specific deterministic approach to solve an optimization problem.
Using Tensorflow 2 and Keras
Let us now turn to coding. To be able to use a Nvidia GPU we need a Cuda/Cudnn installation and a working Tensorflow backend for Keras. I have already described the installation of CUDA 10.2 and CUDNN on an Opensuse Linux system in some detail in the article Getting a Keras based MLP to run with Cuda 10.2, Cudnn 7.6 and TensorFlow 2.0 on an Opensuse Leap 15.1 system. You can follow the hints there. In case you run into trouble on your Linux distribution try everything with Cuda 10.1.
Some more hints: TF2 in version 2.2 can be installed by the Pip-mechanism in your virtual Python environment (“pip install –upgrade tensorflow”). TF2 contains already a special Keras version – which is the one we are going to use in our upcoming experiment. So, there is no need to install Keras separately with “pip”. Note also that, in contrast to TF1, it is NOT necessary to install a separate package “tensorflow-gpu”. If all these things are new to you: You find some articles on creating an adequate ML test and development environment based on Python/PyDev/Jupyter somewhere else in this blog.
Imports and settings for CPUs/GPUs
We shall use a Jupyter notebook to perform the basic experiments; but I recommend strongly to consolidate your code in Python files of an Eclipse/PyDev environment afterwards. Before you start your virtual Python environment from a Linux shell you should set the following environment variables:
$>export OPENBLAS_NUM_THREADS=4 # or whatever is reasonable for your CPU (but do not use all CPU cores and/or hyper threads $>export OMP_NUM_THREADS=4 $>export TF_XLA_FLAGS=--tf_xla_cpu_global_jit $>export XLA_FLAGS=--xla_gpu_cuda_data_dir=/usr/local/cuda $>source bin/activate (ml_1) $> jupyter notebook
Required Imports
The following commands in a first Jupyter cell perform the required library imports:
import numpy as np import scipy import time import sys import os import tensorflow as tf from tensorflow import keras as K from tensorflow.python.keras import backend as B from keras import models from keras import layers from keras.utils import to_categorical from keras.datasets import mnist from tensorflow.python.client import device_lib from sklearn.preprocessing import StandardScaler
Do not ignore the statement “from tensorflow.python.keras import backend as B“; we need it later.
The “StandardScaler” of Scikit-Learn will help us to “standardize” the MNIST input data. This is a step which you should know already from MLPs … You can, of course, also experiment with different normalization procedures. But in my opinion using the “StandardScaler” is just convenient. ( I assume that you already have installed scikit-learn in your virtual Python environment).
Settings for CPUs/GPUs
With TF2 the switching between CPU and GPU is a bit of a mess. Not all new parameters and their settings work as expected. As I have explained in the article on the Cuda installation named above, I, therefore, prefer to an old school, but reliable TF1 approach and use the compatibility interface:
#gpu = False
gpu = True
if gpu:
GPU = True; CPU = False; num_GPU = 1; num_CPU = 1
else:
GPU = False; CPU = True; num_CPU = 1; num_GPU = 0
config = tf.compat.v1.ConfigProto(intra_op_parallelism_threads=6,
inter_op_parallelism_threads=1,
allow_soft_placement=True,
device_count = {'CPU' : num_CPU,
'GPU' : num_GPU},
log_device_placement=True
)
config.gpu_options.per_process_gpu_memory_fraction=0.35
config.gpu_options.force_gpu_compatible = True
B.set_session(tf.compat.v1.Session(config=config))
We are brave and try our first runs directly on a GPU. The statement “log_device_placement” will help us to get information about which device – CPU or GP – is actually used.
Loading and preparing MNIST data
We prepare a function which loads and prepares the MNIST data for us. The statements reflect more or less what we did with the MNIST dat when we used them for MLPs.
# load MNIST
# **********
def load_Mnist():
mnist = K.datasets.mnist
(X_train, y_train), (X_test, y_test) = mnist.load_
data()
#print(X_train.shape)
#print(X_test.shape)
# preprocess - flatten
len_train = X_train.shape[0]
len_test = X_test.shape[0]
X_train = X_train.reshape(len_train, 28*28)
X_test = X_test.reshape(len_test, 28*28)
#concatenate
_X = np.concatenate((X_train, X_test), axis=0)
_y = np.concatenate((y_train, y_test), axis=0)
_dim_X = _X.shape[0]
# 32-bit
_X = _X.astype(np.float32)
_y = _y.astype(np.int32)
# normalize
scaler = StandardScaler()
_X = scaler.fit_transform(_X)
# mixing the training indices - MUST happen BEFORE encoding
shuffled_index = np.random.permutation(_dim_X)
_X, _y = _X[shuffled_index], _y[shuffled_index]
# split again
num_test = 10000
num_train = _dim_X - num_test
X_train, X_test, y_train, y_test = _X[:num_train], _X[num_train:], _y[:num_train], _y[num_train:]
# reshape to Keras tensor requirements
train_imgs = X_train.reshape((num_train, 28, 28, 1))
test_imgs = X_test.reshape((num_test, 28, 28, 1))
#print(train_imgs.shape)
#print(test_imgs.shape)
# one-hot-encoding
train_labels = to_categorical(y_train)
test_labels = to_categorical(y_test)
#print(test_labels[4])
return train_imgs, test_imgs, train_labels, test_labels
if gpu:
with tf.device("/GPU:0"):
train_imgs, test_imgs, train_labels, test_labels= load_Mnist()
else:
with tf.device("/CPU:0"):
train_imgs, test_imgs, train_labels, test_labels= load_Mnist()
Some comments:
- Normalization and shuffling: The “StandardScaler” is used for data normalization. I also shuffled the data to avoid any pre-ordered sequences. We know these steps already from the MLP code we built in another article series.
- Image data in tensor form: Something, which is different from working with MLPs is that we have to fulfill some requirements regarding the form of input data. From the last article we know already that our data should have a tensor compatible form; Keras expects data from us which have a certain shape. So, no flattening of the data into a vector here as we were used to with MLPs. For images we, instead, need the width, the height of our images in terms of pixels and also the depth (here 1 for gray-scale images). In addition the data samples are to be indexed along the first tensor axis.
This means that we need to deliver a 4-dimensional array corresponding to a TF tensor of rank 4. Keras/TF2 will do the necessary transformation from a Numpy array to a TF2 tensor automatically for us. The corresponding Numpy shape of the required array is:
(samples, height, width, depth)
Some people also use the term “channels” instead of “depth”. In the case of MNIST we reshape the input array – “train_imgs” to (num_train, 28, 28, 1), with “num_train” being the number of samples. - The use of the function “to_categorical()”, more precisely “tf.keras.utils.to_categorical()”, corresponds to a one-hot-encoding of the target data. All these concepts are well known from our study of MLPs and MNIST. Keras makes life easy regarding this point …
- The statements “with tf.device(“/GPU:0”):” and “with tf.device(“/CPU:0”):” delegate the execution of (suitable) code on the GPU or the CPU. Note that due to the Python/Jupyter environment some code will of course also be executed on the CPU – even if you delegated execution to the GPU.
If you activate the print statements the resulting output should be:
(60000, 28, 28) (10000, 28, 28) (60000, 28, 28, 1) (10000, 28, 28, 1) [0. 0. 0. 0. 0. 0. 0. 1. 0. 0.]
The last line proves the one-hot-encoding.
The CNN architecture – and Keras’ layer API
Now, we come to a central point: We need to build the 5 central layers of our CNN-architecture. When we build our own MLP code we used a special method to build the different weight arrays, which represented the number of nodes via the array dimensions. A simple method was sufficient as we had no major differences between layers. But with CNNs we have to work with substantially different types of layers. So, how are layers to be handled with Keras?
Well, Keras provides a full layer API with different classes for a variety of layers. You find substantial information on this API and different types of layers at
https://keras.io/api/layers/.
The first section which is interesting for our experiment is https://keras.io/api/ layers/ convolution_layers/ convolution2d/.
You do not need to read much to understand that this is exactly what we need for the “convolutional layers” of our simple CNN model. But how do we instantiate the Conv2D class such that the output works seamlessly together with other layers?
Keras makes our life easy again. All layers are to be used in a purely sequential order. (There are much more complicated layer topologies you can build with Keras! Oh, yes …). Well, you guess it: Keras offers you a model API; see:
https://keras.io/api/models/.
And there we find a class for a “sequential model” – see https://keras.io/api/ models/sequential/. This class offers us a method “add()” to add layers (and create an instance of the related layer class).
The only missing ingredient is a class for a “pooling” layer. Well, you find it in the layer API documentation, too. The following image depicts the basic structure of our CNN (see the left side of the drawing), as we designed it (see the list above):
Keras code for the Conv and pooling layers
The convolutional part of the CNN can be set up by the following commands:
Convolutional part of the CNN
# Sequential layer model of our CNN # *********************************** # Build the Conv part of the CNN # ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~ # Choose the activation function for the Conv2D layers conv_act_func = 1 li_conv_act_funcs = ['sigmoid', 'relu', 'elu', 'tanh'] cact = li_conv_act_funcs[conv_act_func] # Build the Conv2D layers cnn = models.Sequential() cnn.add(layers.Conv2D(32, (3,3), activation=cact, input_shape=(28,28,1))) cnn.add(layers.MaxPooling2D((2,2))) cnn.add(layers.Conv2D(64, (3,3), activation=cact)) cnn.add(layers.MaxPooling2D((2,2))) cnn.add(layers.Conv2D(64, (3,3), activation=cact))
Easy, isn’t it? The nice thing about Keras is that it cares about the required tensor ranks and shapes itself; in a sequential model it evaluates the output of a already defined layer to guess the shape of the tensor data entering the next layer. Thus we have to define an “input_shape” only for data entering the first Conv2D layer!
The first Conv2D layer requires, of course, a shape for the input data. We must also tell the layer interface how many filters and “feature maps” we want to use. In our case we produce 32 maps by first Conv2D layer and 64 by the other two Conv2D layers. The (3×3)-parameter defines the filter area size to be covered by the filter kernel: 3×3 pixels. We define no “stride”, so a stride of 1 is automatically used; all 3×3 areas lie close to each other and overlap each other. These parameters result in 32 maps of size 26×26 after the first convolution. The size of the maps of the other layers are given in the layer list at the beginning of this article.
In addition you saw from the code that we chose an activation function via an index of a Python list of reasonable alternatives. You find an explanation of all the different activation functions in the literature. (See also: wikipedia: Activation function). The “sigmoid” function should be well known to you already from my other article series on a MLP.
Now, we have to care about the MLP part of the CNN. The code is simple:
MLP part of the CNN
# Build the MLP part of the CNN # ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~ # Choose the activation function for the hidden layers of the MLP mlp_h_act_func = 0 li_mlp_h_act_funcs = ['relu', 'sigmoid', 'tanh'] mhact = li_mlp_h_act_funcs[mlp_h_act_func] # Choose the output function for the output layer of the MLP mlp_o_act_func = 0 li_mlp_o_act_funcs = ['softmax', 'sigmoid'] moact = li_mlp_o_act_funcs[mlp_o_act_func] # Build the MLP layers cnn.add(layers.Flatten()) cnn.add(layers.Dense(70, activation=mhact)) #cnn.add(layers.Dense(30, activation=mhact)) cnn.add(layers.Dense(10, activation=moact))
This all is very straight forward (with the exception of the last statement). The “Flatten”-layer corresponds to the MLP’s inout layer. It just transforms the tensor output of the last Conv2D layer into the flat form usable for the first “Dense” layer of the MLP. The first and only “Dense layer” (MLP hidden layer) builds up connections from the flat MLP “input layer” and associates it with weights. Actually, it prepares a weight-tensor for a tensor-operation on the output data of the feeding layer. Dense means that all “nodes” of the previous layer are connected to the present layer’s own “nodes” – meaning: setting the right dimensions of the weight tensor (matrix in our case). As a first trial we work with just one hidden layer. (We shall later see that more layers will not improve accuracy.)
I choose the output function (if you will: the activation function of the output layer) as “softmax“. This gives us a probability distribution across the classification categories. Note that this is a different approach compared to what we have done so far with MLPs. I shall comment on the differences in a separate article when I find the time for it. At this point I just want to indicate that softmax combined with the “categorical cross entropy loss” is a generalized version of the combination “sigmoid” with “log loss” as we used it for our MLP.
Parameterization
The above code for creating the CNN would work. However, we want to be able to parameterize our simple CNN. So we include the above statements in a function for which we provide parameters for all layers. A quick solution is to define layer parameters as elements of a Python list – we then get one list per layer. (If you are a friend of clean code design I recommend to choose a more elaborated approach; inject just one parameter object containing all parameters in a structured way. I leave this exercise to you.)
We now combine the statements for layer construction in a function:
# Sequential layer model of our CNN
# ***********************************
# just for illustration - the real parameters are fed later
#
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
li_conv_1 = [32, (3,3), 0]
li_conv_2 = [64, (3,3), 0]
li_conv_3 = [64, (3,3), 0]
li_Conv = [li_conv_1, li_conv_2, li_conv_3]
li_pool_1 = [(2,2)]
li_pool_2 = [(2,2)]
li_Pool = [li_pool_1, li_pool_2]
li_dense_1 = [70, 0]
li_dense_2 = [10, 0]
li_MLP = [li_dense_1, li_dense_2]
input_shape = (28,28,1)
# important !!
# ~~~~~~~~~~~
cnn = None
def build_cnn_simple(li_Conv, li_Pool, li_MLP, input_shape ):
# activation functions to be used in Conv-layers
li_conv_act_funcs = ['relu', 'sigmoid', 'elu', 'tanh']
# activation functions to be used in MLP hidden layers
li_mlp_h_act_funcs = ['relu', 'sigmoid', 'tanh']
# activation functions to be used in MLP output layers
li_mlp_o_act_funcs = ['softmax', 'sigmoid']
# Build the Conv part of the CNN
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
num_conv_layers = len(li_Conv)
num_pool_layers = len(li_Pool)
if num_pool_layers != num_conv_layers - 1:
print("\nNumber of pool layers does not fit to number of Conv-layers")
sys.exit()
rg_il = range(num_conv_layers)
# Define a sequential model
cnn = models.Sequential()
for il in rg_il:
# add the convolutional layer
num_filters = li_Conv[il][0]
t_fkern_size = li_Conv[il][1]
cact = li_conv_act_funcs[li_Conv[il][2]]
if il==0:
cnn.add(layers.Conv2D(num_filters, t_fkern_size, activation=cact, input_shape=input_shape))
else:
cnn.add(layers.Conv2D(num_filters, t_fkern_size, activation=cact))
# add the pooling layer
if il < num_pool_layers:
t_pkern_size = li_Pool[il][0]
cnn.add(layers.MaxPooling2D(t_pkern_size))
# Build the MLP part of the CNN
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
num_mlp_layers = len(li_MLP)
rg_im = range(num_mlp_layers)
cnn.add(layers.Flatten())
for im in rg_im:
# add the dense layer
n_nodes = li_MLP[im][0]
if im < num_mlp_layers - 1:
m_act = li_mlp_h_act_funcs[li_MLP[im][1]]
else:
m_act = li_mlp_o_act_funcs[li_MLP[im][1]]
cnn.add(layers.Dense(n_nodes, activation=m_act))
return cnn
We return the model “cnn” to be able to use it afterwards.
How many parameters does our CNN have?
The layers contribute with the following numbers of weight parameters:
- Layer 1: (32 x (3×3)) + 32 = 320
- Layer 3: 32 x 64 x (3×3) + 64 = 18496
- Layer 5: 64 x 64 x (3×3) + 64 = 36928
- MLP : (576 + 1) x 70 + (70 + 1) x 10 = 41100
Making a total of 96844 weight parameters. Our standard MLP discussed in another article series had (784+1) x 70 + (70 + 1) x 30 + (30 +1 ) x 10 = 57390 weights. So, our CNN is bigger and the CPU time to follow and calculate all the partial derivatives will be significantly higher. So, we should definitely expect some better classification data, shouldn’t we?
Compilation
Now comes a thing which is necessary for models: We have not yet defined the loss function and the optimizer or a learning rate. For the latter Keras can choose a proper value itself – as soon as it knows the loss function. But we should give it a reasonable loss function and a suitable optimizer for gradient descent. This is the main purpose of the “compile()“-function.
cnn.compile(optimizer='rmsprop', loss='categorical_crossentropy', metrics=['accuracy'])
Although TF2 can already analyze the graph of tensor operations for partial derivatives, it cannot guess the beginning of the chain rule sequence.
As we have multiple categories “categorial_crossentropy” is a good choice for the loss function. We should also define which optimized gradient descent method is used; we choose “rmsprop” – as this method works well in most cases. A nice introduction is given here: towardsdatascience: understanding-rmsprop-faster-neural-network-learning-62e116fcf29a. But see the books mentioned in the last article on “rmsprop”, too.
Regarding the use of different metrics for different tasks see machinelearningmastery.com / custom-metrics-deep-learning-keras-python/. In case of a classification problem, we are interested in the categorical “accuracy”. A metric can be monitored during training and will be recorded (besides aother data). We can use it for plotting information on the training process (a topic of the next article).
Training
Training is done by a function model.fit() – here: cnn.fit(). This function accepts a variety of parameters explained here: https://keras.io/ api/ models/ model_training_apis/ #fit-method.
We now can combine compilation and training in one function:
# Training
def train( cnn, build=False, train_imgs, train_labels, reset, epochs, batch_size, optimizer, loss, metrics,
li_Conv, li_Poo, li_MLP, input_shape ):
if build:
cnn = build_cnn_simple( li_Conv, li_Pool, li_MLP, input_shape)
cnn.compile(optimizer=optimizer, loss=loss, metrics=metrics)
cnn.save_weights('cnn_weights.h5') # save the initial weights
# reset weights
if reset and not build:
cnn.load_weights('cnn_weights.h5')
start_t = time.perf_counter()
cnn.fit(train_imgs, train_labels, epochs=epochs, batch_size=batch_size, verbose=1, shuffle=True)
end_t = time.perf_counter()
fit_t = end_t - start_t
return cnn, fit_t # we return cnn to be able to use it by other functions
Note that we save the initial weights to be able to load them again for a new training – otherwise Keras saves the weights as other model data after training and continues with the last weights found. The latter can be reasonable if you want to continue training in defined steps. However, in our forthcoming tests we repeat the training from scratch.
Keras offers a “save”-model and methods to transfer data of a CNN model to files (in two specific formats). For saving weights the given lines are sufficient. Hint: When I specify no path to the file “cnn_weights.h5” the data are – at least in my virtual Python environment – saved in the directory where the notebook is located.
First test
In a further Jupyter cell we place the following code for a test run:
# Perform a training run
# ********************
build = False
if cnn == None:
build = True
batch_size=64
epochs=5
reset = True # we want training to start again with the initial weights
optimizer='rmsprop'
loss='categorical_crossentropy'
metrics=['accuracy']
li_conv_1 = [32, (3,3), 0]
li_conv_2 = [64, (3,3), 0]
li_conv_3 = [64, (3,3), 0]
li_Conv = [li_conv_1, li_conv_2, li_conv_3]
li_pool_1 = [(2,2)]
li_pool_2 = [(2,2)]
li_Pool = [li_pool_1, li_pool_2]
li_dense_1 = [70, 0]
li_dense_2 = [10, 0]
li_MLP = [li_dense_1, li_dense_2]
input_shape = (28,28,1)
try:
if gpu:
with tf.device("/GPU:0"):
cnn, fit_time = train( cnn, build, train_imgs, train_
labels,
reset, epochs, batch_size, optimizer, loss, metrics,
li_Conv, li_Pool, li_MLP, input_shape)
print('Time_GPU: ', fit_time)
else:
with tf.device("/CPU:0"):
cnn, fit_time = train( cnn, build, train_imgs, train_labels,
reset, epochs, batch_size, optimizer, loss, metrics,
li_Conv, li_Pool, li_MLP, input_shape)
print('Time_CPU: ', fit_time)
except SystemExit:
print("stopped due to exception")
You recognize the parameterization of our train()-function. What results do we get ?
Epoch 1/5 60000/60000 [==============================] - 4s 69us/step - loss: 0.1551 - accuracy: 0.9520 Epoch 2/5 60000/60000 [==============================] - 4s 69us/step - loss: 0.0438 - accuracy: 0.9868 Epoch 3/5 60000/60000 [==============================] - 4s 68us/step - loss: 0.0305 - accuracy: 0.9907 Epoch 4/5 60000/60000 [==============================] - 4s 69us/step - loss: 0.0227 - accuracy: 0.9931 Epoch 5/5 60000/60000 [==============================] - 4s 69us/step - loss: 0.0184 - accuracy: 0.9948 Time_GPU: 20.610678611003095
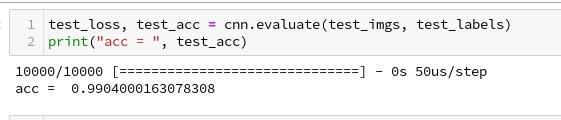
And a successive check on the test data gives us:
We can ask Keras also for a description of the model:
Accuracy at the 99% level
We got an accuracy on the test data of 99%! With 5 epochs in 20 seconds – on my old GPU.
This leaves us a very good impression – on first sight …
Conclusion
We saw today that it is easy to set up a CNN. We used a simple MLP to solve the problem of classification; the data to its input layer are provided by the output of the last convolutional layer. The tensor there has just to be “flattened”.
The level of accuracy reached is impressing. Well, its also a bit frustrating when we think about the efforts we put into our MLP, but we also get a sense for the power and capabilities of CNNs.
In the next post
A simple CNN for the MNIST dataset – III – inclusion of a learning-rate scheduler, momentum and a L2-regularizer
we will care a bit about plotting. We at least want to see the same accuracy and loss data which we used to plot at the end of our MLP tests.