In the last article of this series
Deep Dreams of a CNN trained on MNIST data – I – a first approach based on one selected map of a convolutional layer
I presented some “Deep Dream” images of a CNN trained on MNIST data. Although one cannot produce spectacular images with a simple CNN based on gray scale, low resolution image data, we still got a glimpse of what the big guys do with more advanced CNNs and image data. But we have only started, yet ….
In the forthcoming articles we shall extend our abilities step by step beyond the present level. Our next goal is to work on different length scales within the image, i.e. we go down into sub-tiles of the original image, analyze the sub-structures there and include the results in the CNN’s dreams. But first we need to understand more precisely how we apply the image manipulation techniques that “carve” some additional dream-like optical elements, which correspond to map triggering features, into an arbitrary input image presented to the CNN.
Carving: A combination of low-resolution OIP-pattern creation and a reproduction of high-resolution details
I used the term “carved” intentionally. Artists or craftsmen who create little figures or faces out of wooden blocks by a steady process of carving often say that they only bring to the surface what already was there – namely in the basic fiber structures of the wood. Well, the original algorithm for the creation of pure OIP- or feature patterns systematically amplifies some minimal correlated pixel “structures” detected in an input image with random pixel values. The values of all other pixels outside the OIP pattern are reduced; sooner or later they form a darker and rather homogeneous background. The “detection” is based on (trained) filters which react to certain pixel constellations. We already encoded an adaption of a related method for CNN filter visualization, which Francois Chollet discusses in his book on “Deep learning with Python”, into a Python class in another article series of this blog (see the link section at the end of this article).
There is, however, a major difference regarding our present context: During the image manipulations for the visualization of Deep Dreams we do not want to remove all details of the original image presented to the DeepDream algorithm. Instead, we just modify it at places where the CNN detects traces of patterns which trigger a strong response of one or multiple maps of our CNN – and keep detail information alive elsewhere. I.e., we must counterbalance the tendency of our OIP algorithm to level out information outside the OIP structures.
How do we do this, when both our present OIP algorithm and the MNIST-trained CNN are limited to a resolution of (28×28) pixels, only?
Carving – amplification of certain OIP structures and intermittent replenishment of details of the original image
In this article we follow the simple strategy explained in the last article:
- Preparation:
- We choose a map.
- We turn the colored original input image (about which the CNN shall dream) into a gray one.
- We downscale the original image to a size of (28×28) by interpolation, upscale the result again by interpolating again (with loss) and calculate the difference to the original full resolution image (all interpolations done in a bicubic way). (The difference contains information on detail structures.)
- Loop (4 times or so):
- We apply the OIP-algorithm on the downscaled input image for a fixed amount of epochs with a small epsilon.
The result is a slight emphasis of
pixel structures resembling OIP patterns.
- We upscale the result by bicubic interpolation to the original size.
- We re-add the difference to the original image, i.e. we supply details again.
- We downscale the result again.
- Eventual image optimization: We apply some contrast treatment to the resulting image, whose pixel value range and distribution has gradually been changed in comparison to the original image.
This corresponds to a careful accentuation of an OIP-pattern detected by a certain map in the given structured pixel data of the input image (on coarse scales) – without loosing all details of the initial image. As said – it resembles a process of carving coarse patterns into the original image ..
Note: The above algorithm works on the whole image – a modified version working on a variety of smaller length scales is discussed in further articles.
What does this strategy look like in Python statements for Jupyter cells?
Start with loading required modules and involving your graphics card
We have to load python modules to support our CNN, the OIP algorithm and some plotting. In a second step we need to invoke the graphics card. This is done in two Jupyter cells (1 and 2). Their content was already described in the article
A simple CNN for the MNIST dataset – XI – Python code for filter visualization and OIP detection
of a parallel series in this blog. These initial steps are followed by an instantiation of a class “My_OIP” containing all required methods. The __init__() function of this class restores a Keras model of our basic CNN from a h5-file:
Jupyter cell 3
%matplotlib inline
# Load the CNN-model
# ~~~~~~~~~~~~~~~~~
imp.reload(myOIP)
try:
with tf.device("/GPU:0"):
MyOIP = myOIP.My_OIP(cnn_model_file = 'cnn_best.h5', layer_name = 'Conv2D_3')
except SystemExit:
print("stopped")
Turn the original image into a gray one, downscale it and calculate difference-tensors for coarsely re-enlarged versions
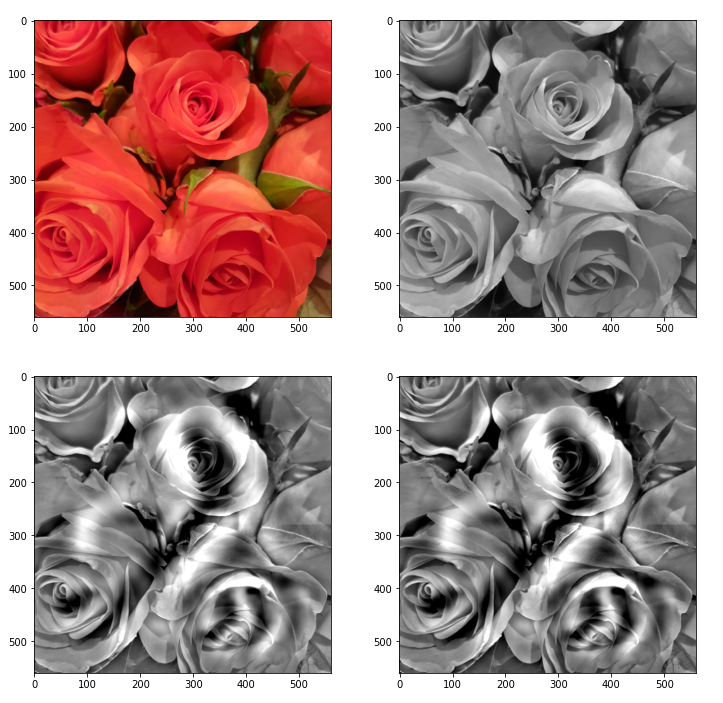
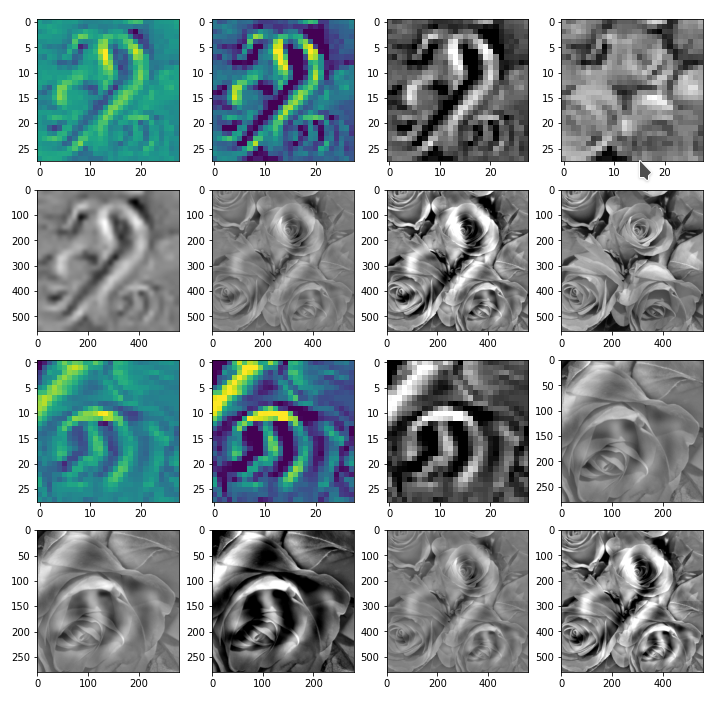
In the next Jupyter cell we deal with the following initial image:

We choose a method (out of 3) to turn it into a gray scaled image. Afterwards we calculate the difference between an image tensor of the original (gray) image and a coarse, but softly rescaled image derived from a (28×28)-pixel version.
Jupyter cell 4
# Down-, gray- and Re-scale images, determine correction differences
# *******************************************************************
import PIL
from PIL import Image, ImageOps
fig_size = plt.rcParams["figure.figsize"]
fig_size[0] = 15
fig_size[1] = 30
fig_1 = plt.figure(1)
ax1_1 = fig_1.add_subplot(531)
ax1_2 = fig_1.add_subplot(532)
ax1_3 = fig_1.add_subplot(533)
ax2_1 = fig_1.add_subplot(534)
ax2_2 = fig_1.add_subplot(535)
ax2_3 = fig_1.add_subplot(536)
ax3_1 = fig_1.add_subplot(537)
ax3_2 = fig_1.add_subplot(538)
ax3_3 = fig_1.add_subplot(539)
ax4_1 = fig_1.add_subplot(5,3,10)
ax4_2 = fig_1.add_subplot(5,3,11)
ax4_3 = fig_1.add_subplot(5,3,12)
# Parameters
# ************
# size of the image to work with
img_wk_size = 560
# method to turn the original picture into a gray scaled one
gray_scaling_
method = 2
# bring the orig img down to (560, 560)
# ***************************************
imgvc = Image.open("rosen_orig_farbe.jpg")
imgvc_g = imgvc.convert("L")
# downsize with PIL
# ******************
imgvc_wk_size = imgvc.resize((img_wk_size,img_wk_size), resample=PIL.Image.BICUBIC)
imgvc_wk_size_g = imgvc_wk_size.convert("L")
imgvc_28 = imgvc.resize( (28,28), resample=PIL.Image.BICUBIC)
imgvc_g_28 = imgvc_g.resize((28,28), resample=PIL.Image.BICUBIC)
# Change to np array
ay_picc = np.array(imgvc_wk_size)
# Display orig and wk_size images
# *********************************
ax1_1.imshow(imgvc)
ax1_2.imshow(imgvc_wk_size)
ax1_3.imshow(imgvc_g, cmap=plt.cm.gray)
# Apply 3 different methods to turn the image into a gray one
# **************************************************************
#Red * 0.3 + Green * 0.59 + Blue * 0.11
#Red * 0.2126 + Green * 0.7152 + Blue * 0.0722
#Red * 0.299 + Green * 0.587 + Blue * 0.114
ay_picc_g1 = ( 0.3 * ay_picc[:,:,0] + 0.59 * ay_picc[:,:,1] + 0.11 * ay_picc[:,:,2] )
ay_picc_g2 = ( 0.299 * ay_picc[:,:,0] + 0.587 * ay_picc[:,:,1] + 0.114 * ay_picc[:,:,2] )
ay_picc_g3 = ( 0.2126 * ay_picc[:,:,0] + 0.7152 * ay_picc[:,:,1] + 0.0722 * ay_picc[:,:,2] )
ay_picc_g1 = ay_picc_g1.astype('float32')
ay_picc_g2 = ay_picc_g2.astype('float32')
ay_picc_g3 = ay_picc_g3.astype('float32')
# Prepare tensors
# *****************
t_picc_g1 = ay_picc_g1.reshape((1, img_wk_size, img_wk_size, 1))
t_picc_g2 = ay_picc_g2.reshape((1, img_wk_size, img_wk_size, 1))
t_picc_g3 = ay_picc_g3.reshape((1, img_wk_size, img_wk_size, 1))
t_picc_g1 = tf.image.per_image_standardization(t_picc_g1)
t_picc_g2 = tf.image.per_image_standardization(t_picc_g2)
t_picc_g3 = tf.image.per_image_standardization(t_picc_g3)
# Display gray-tensor variants
# ****************************
ax2_1.imshow(t_picc_g1[0,:,:,0], cmap=plt.cm.gray)
ax2_2.imshow(t_picc_g2[0,:,:,0], cmap=plt.cm.gray)
ax2_3.imshow(t_picc_g3[0,:,:,0], cmap=plt.cm.gray)
# choose one gray img
# ******************
if gray_scaling_method == 1:
t_picc_g = t_picc_g1
elif gray_scaling_method == 2:
t_picc_g = t_picc_g2
else:
t_picc_g = t_picc_g3
# downsize to (28,28) and standardize
# *****************************************
t_picc_g_28_scd = tf.image.resize(t_picc_g, [28,28], method="bicubic", antialias=True)
t_picc_g_28_std = tf.image.per_image_standardization(t_picc_g_28_scd)
t_picc_g_28 = t_picc_g_28_std
# display
ax3_1.imshow(imgvc_g_28, cmap=plt.cm.gray)
ax3_2.imshow(t_picc_g_28_scd[0,:,:,0], cmap=plt.cm.gray)
ax3_3.imshow(t_picc_g_28_std[0,:,:,0], cmap=plt.cm.gray)
# Upscale and get correction values
# **********************************
t_picc_g_wk_size_scd = tf.image.resize(t_picc_g_28, [img_wk_size,img_wk_size], method="bicubic", antialias=True)
t_picc_g_wk_size_scd_std = tf.image.per_image_standardization(t_picc_g_wk_size_scd)
t_picc_g_wk_size_corr = t_picc_g - t_picc_g_wk_size_scd_std
# Correct and display
# **********************
t_picc_g_wk_size_re = t_picc_g_wk_size_scd_std + t_picc_g_wk_size_corr
# Display
ax4_1.imshow(t_picc_g_wk_size_scd_std[0,:,:,0], cmap=plt.cm.gray)
ax4_2.imshow(t_picc_g[0,:,:,0], cmap=plt.cm.gray)
ax4_3.imshow(t_picc_g_wk_size_re[0,:,:,0], cmap=plt.cm.gray)
The code is straightforward – a lot of visualization steps will later on just show tiny differences. For production you may shorten the code significantly.
In a first step we downscale the image to a work-size of (560×560) px with the help of PIL modules. (Note that 560 can be divided by 2, 4, 8, 16, 28). We then test out three methods to turn the image into a gray one. This step is required, because our MNIST-oriented CNN is trained for gray images, only. We
reshape the image data arrays to prepare for tensor-operations and standardize with the help of Tensorflow – and produce image tensors on the fly. Afterwards, we downscale to a size of (28×28) pixels; this is the size of the MNIST images for which the CNN has been trained. We upscale to the work-size again, compare it to the original detailed image and calculate respective differences of the tensor-components. These differences will later be used to re-add details to our manipulated images.
The following picture displays the resulting output:
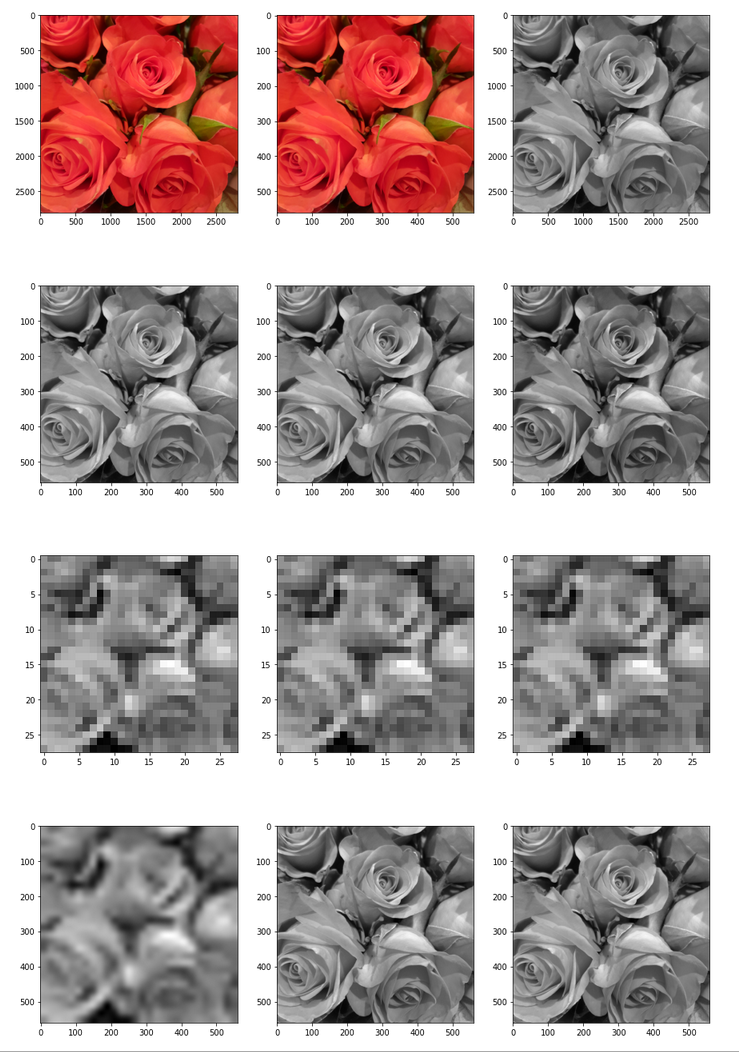

Note: The last two images should not show any difference. The middle row and the displays how coarse the (28×28) resolution really is and how much information we have lost after upscaling to (560×560)px.
OIP-Carving and detail replenishment
We now follow the algorithm described above. This leads to a main loop with steps for downscaling, OIP-carving on the (28×28)-scale, upscaling and a supplementation of original details:
Jupyter cell 5
# **************************************************+
# OIP analysis (to be used after the previous cell)
# **************************************************+
# Prepare plotting
# -------------------
#interactive plotting
#%matplotlib notebook
#plt.ion()
fig_size = plt.rcParams["figure.figsize"]
fig_size[0] = 15
fig_size[1] = 25
fig_2 = plt.figure(2)
ax2_1_1 = fig_2.add_subplot(531)
ax2_1_2 = fig_2.add_subplot(532)
ax2_1_3 = fig_2.add_subplot(533)
ax2_2_1 = fig_2.add_subplot(534)
ax2_2_2 = fig_2.add_subplot(535)
ax2_2_3 = fig_2.add_subplot(536)
ax2_3_1 = fig_2.add_subplot(537)
ax2_3_2 = fig_2.add_subplot(538)
ax2_3_3 = fig_2.add_subplot(539)
ax2_4_1 = fig_2.add_subplot(5,3,10)
ax2_4_2 = fig_2.add_subplot(5,3,11)
ax2_4_3 = fig_2.add_subplot(5,3,12)
fig_size = plt.rcParams["figure.figsize"]
fig_size[0] = 16
fig_size[1] = 8
fig_3 = plt.figure(3)
axa_1 = fig_3.add_subplot(241)
axa_2 = fig_3.add_subplot(242)
axa_3 = fig_3.add_subplot(243)
axa_4 = fig_3.add_subplot(244)
axa_5 = fig_3.add_subplot(245)
axa_6 = fig_3.add_subplot(246)
axa_7 = fig_3.add_subplot(247)
axa_8 = fig_3.add_subplot(248)
li_axa = [axa_1, axa_2, axa_3, axa_4, axa_5, axa_6, axa_7, axa_8]
fig_size = plt.rcParams["figure.figsize"]
fig_size[0] = 16
fig_size[1] = 4
fig_h = plt.figure(50)
ax_h_1 = fig_h.add_subplot(141)
ax_h_2 = fig_h.add_subplot(142)
ax_h_3 = fig_h.add_subplot(143)
ax_h_4 = fig_h.add_subplot(144)
li_axa = [axa_1, axa_2, axa_3, axa_4, axa_5, axa_6, axa_7, axa_8]
# list for img tensors
li_t_imgs = []
# Parameters for the OIP run
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
map_index = 56 # map-index we are interested in
# map_index = -1 # for this value the whole layer is taken; the loss function is averaged over the maps
n_epochs = 20 # should be divisible by 5
n_steps = 6 # number of intermediate reports
epsilon = 0.01 # step size for gradient correction
conv_criterion = 2.e-4 # criterion for a potential stop of optimization
n_iter = 5 # number of iterations for down-, upscaling and supplementation of details
# Initial image
# -----------------
MyOIP._initial_inp_img_data = t_picc_g_28
# Main iteration loop
# ***********************
for i in range(n_iter):
# Perform the OIP analysis
MyOIP._derive_OIP(map_index = map_index,
n_epochs = n_epochs, n_steps = n_steps,
epsilon = epsilon , conv_criterion = conv_criterion,
b_stop_with_convergence=False,
b_print = False,
li_axa = li_axa,
ax1_1 = ax2_1_1, ax1_2 = ax2_1_2,
centre_move = 0.33, fact = 1.0)
t_oip_c_g_28 = MyOIP._inp_img_data
ay_oip_c_g_28 = t_oip_c_g_28[0,:,:,0].numpy()
ay_oip_c_g_28_cont = MyOIP._transform_tensor_to_img(T_img=t_oip_c_g_28[0,:,:,0], centre_move=0.33, fact=1.0)
ax2_1_3.imshow(ay_oip_c_g_28_cont, cmap=plt.cm.gray)
# Rescale to 500
t_oip_c_g_wk_size = tf.image.resize(t_oip_c_g_28, [img_wk_size,img_wk_size], method="bicubic", antialias=True)
#t_oip_c_g_500 = tf.image.per_image_standardization(t_oip_c_g_500)
t_oip_c_g_wk_size_re = t_oip_c_g_wk_size + t_picc_g_wk_size_corr
t_oip_c_g_wk_size_re_std = tf.image.per_image_standardization(t_oip_c_g_wk_size_re)
# remember the img tensors of the iterations
li_t_imgs.append(t_oip_c_g_wk_size_re_std)
# do not change the gray spectrum for iterations
MyOIP._initial_inp_img_data = tf.image.resize(t_oip_c_g_wk_size_re_std, [28,28], method="bicubic", antialias=True)
# Intermediate plotting - if required and if plt.ion() is set
# ------------------------
if b_plot_intermediate and j < n_iter-1:
ay_oip_c_g_wk_size = t_oip_c_g_wk_size[0,:,:,0].numpy()
#display OIPs
ax2_2_1.imshow(t_picc_g_28[0,:,:,0], cmap=plt.cm.gray)
ax2_2_2.imshow(ay_oip_c_g_28, cmap=plt.cm.gray)
ax2_2_3.imshow(ay_oip_c_g_wk_size, cmap=plt.cm.gray)
ax2_3_1.imshow(t_oip_c_g_wk_size[0,:,:,0], cmap=plt.cm.gray)
ax2_3_2.imshow(t_oip_c_g_wk_size_re[0,:,:,0], cmap=plt.cm.gray)
ax2_3_3.imshow(t_picc_g[0,:,:,0], cmap=plt.cm.gray)
# modify the gray spectrum for plotting - cut off by clipping to limit the extend of the gray color map
min1 = tf.reduce_min(t_picc_g)
min2 = tf.reduce_min(t_oip_c_g_wk_size_re)
max1 = tf.reduce_max(t_picc_g)
max2 = tf.reduce_max(t_oip_c_g_wk_size_re)
# tf.print("min1 = ", min1, " :: min2 = ", min2, " :: max1 = ", max1, " :: max2 = ", max2 )
#fac1 = min2/min1 * 0.95
#fac2 = 1.0*fac1
fac2 = 1.18
fac3 = 0.92
cut_lft = fac3*min1
cut_rht = fac3*max1
cut_rht = fac3*abs(min1)
tf.print("cut_lft = ", cut_lft, " cut_rht = ", cut_rht)
t_oip_c_g_wk_size_re_std_plt = fac2 * t_oip_c_g_wk_size_re_std
t_oip_c_g_wk_size_re_std_plt = tf.clip_by_value(t_oip_c_g_wk_size_re_std_plt, cut_lft, cut_rht)
ax2_4_1.imshow(t_oip_c_g_wk_size[0,:,:,0], cmap=plt.cm.gray)
ax2_4_2.imshow(t_oip_c_g_wk_size_re_std[0,:,:,0], cmap=plt.cm.gray)
ax2_4_3.imshow(t_oip_c_g_wk_size_re_std_plt[0,:,:,0], cmap=plt.cm.gray)
# Eventual plotting
# ********************
ay_oip_c_g_wk_size = t_oip_c_g_wk_size[0,:,:,0].numpy()
#display OIPs
ax2_2_1.imshow(t_picc_g_28[0,:,:,0], cmap=plt.cm.gray)
ax2_2_2.imshow(ay_oip_c_g_28, cmap=plt.cm.gray)
ax2_2_3.imshow(ay_oip_c_g_wk_size, cmap=plt.cm.gray)
ax2_3_1.imshow(t_oip_c_g_wk_size[0,:,:,0], cmap=plt.cm.gray)
ax2_3_2.imshow(t_oip_c_g_wk_size_re[0,:,:,0], cmap=plt.cm.gray)
ax2_3_3.imshow(t_picc_g[0,:,:,0], cmap=plt.cm.gray)
# modify the gray spectrum for plotting - cut off by clipping to limit the extend of the gray color map
min1 = tf.reduce_min(t_picc_g)
min2 = tf.reduce_min(t_oip_c_g_wk_size_re)
max1 = tf.reduce_max(t_picc_g)
max2 = tf.reduce_max(t_oip_c_g_wk_size_re)
# tf.print("min1 = ", min1, " :: min2 = ", min2, " :: max1 = ", max1, " :: max2 = ", max2 )
#fac1 = min2/min1 * 0.95
#fac2 = 1.0*fac1
fac2 = 1.10
fac3 = 0.92
cut_lft = fac3*min1
cut_rht = fac3*
max1
#cut_rht = fac3*abs(min1)
tf.print("cut_lft = ", cut_lft, " cut_rht = ", cut_rht)
t_oip_c_g_wk_size_re_std_plt = fac2 * t_oip_c_g_wk_size_re_std
t_oip_c_g_wk_size_re_std_plt = tf.clip_by_value(t_oip_c_g_wk_size_re_std_plt, cut_lft, cut_rht)
ax2_4_1.imshow(t_oip_c_g_wk_size[0,:,:,0], cmap=plt.cm.gray)
ax2_4_2.imshow(t_oip_c_g_wk_size_re_std[0,:,:,0], cmap=plt.cm.gray)
ax2_4_3.imshow(t_oip_c_g_wk_size_re_std_plt[0,:,:,0], cmap=plt.cm.gray)
# Histogram analysis and plotting
# *********************************
# prepare the histograms of greay values for the saved imagetensors
ay0 = np.sort( t_picc_g[0, :, :, 0].numpy().ravel() )
ay1 = np.sort( li_t_imgs[0][0,:,:,0].numpy().ravel() )
ay2 = np.sort( li_t_imgs[1][0,:,:,0].numpy().ravel() )
ay3 = np.sort( li_t_imgs[2][0,:,:,0].numpy().ravel() )
ay4 = np.sort( li_t_imgs[3][0,:,:,0].numpy().ravel() )
#print(ay1)
nx, binsx, patchesx = ax_h_1.hist(ay0, bins='auto')
nx, binsx, patchesx = ax_h_2.hist(ay2, bins='auto')
nx, binsx, patchesx = ax_h_3.hist(ay3, bins='auto')
nx, binsx, patchesx = ax_h_4.hist(ay4, bins='auto')
We first prepare some Matplotlib axes frames. Note that the references should be clearly distinguishable and unique across the cells of the Jupyter notebook.
Parameters
Then we define parameters for a DeepDream run. [The attentive reader has noticed that I now allow for a value of nmap_index = -1. This corresponds to working with a loss function calculated and averaged over all maps (nmap=-1) of a layer. The changed code of the class My_OIP is given at the end of the article. We come back to this option in a later post of this series. For the time being we just work with a chosen single map and its cost function.]
The most interesting parameters are “n_epochs” and “n_iter“. These parameters help to tune a careful balance between OIP-“carving” and “detail replenishment”. These are parameters you absolutely should experiment with!
Iterations
We first pick the (28×28) gray image prepared in the previous Jupyter cell as the initial image presented to the CNN’s input layer. We then start an iteration by invoking the “My_OIP” class discussed in the article named above. (The parameters “centre_move = 0.33” and “fact = 1.0” refer to an image manipulation, namely a contrast enhancement.)
After OIP-carving over the chosen limited number of epochs we apply a special treatment to the data for the OPI-image derived so far. In a side step we enhance the contrast contrast for displaying the intermediate results. We then rescale by bicubic interpolation from 28×28 to the working size 560×560, add the prepared correction data and standardize. We add the results to a list for a later histogram analysis.
The preparation of plotting (of an intermediate and eventual image) comprises a step to reduce the spectrum of pixel values. This step is required as the iterative image manipulation leads to an extension of the maximum and the minimum values, i.e. the flanks of the value spectrum, up to a factor of 1.7 – although we only find a few pixels in that extended range. If we applied a standard gray-scale color map to the whole wider range of values then we would get a strong reduction in contrast – for the sake of including some extreme pixel values. The remedy is to cut the spectrum off at the original maximum/minimum value.
The histogram analysis performed at the end in addition shows that the central spectrum has become a bit narrower – we can compensate this by some factors fac2 and fac3. Play around with them.
Results of the carving process
Results for map 57 are displayed in the following image:
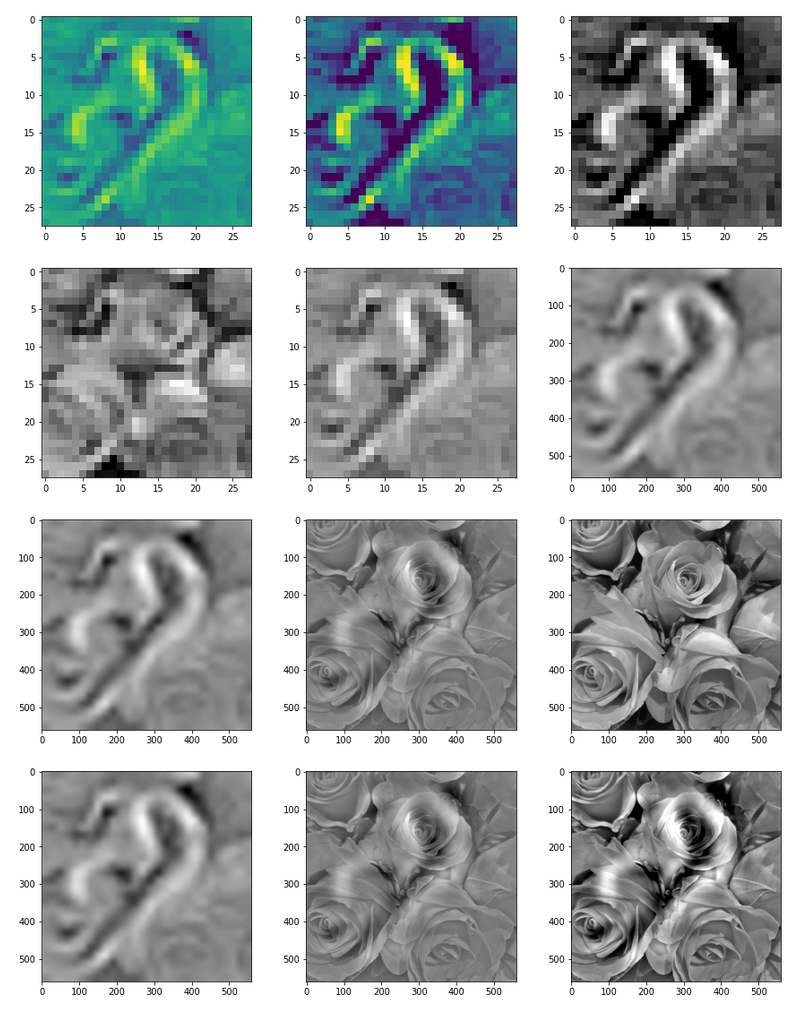
The effect of the “carving” is clearly visible as an overlayed ghost like pattern apparition. Standardization of the final images makes no big difference – however, the discussed compensation for the widened spectrum of the eventual image is important.
The histogram analysis shows the flank widening and the narrowing of the central core of the pixel value spectrum:
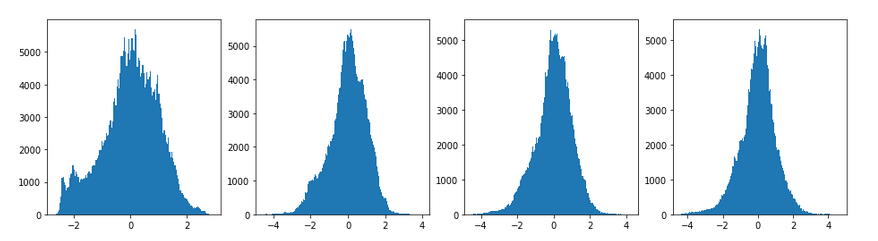
The following image allow a comparison of the original gray image with the carved image for parameters [n_epochs=20, n_iter=5]:
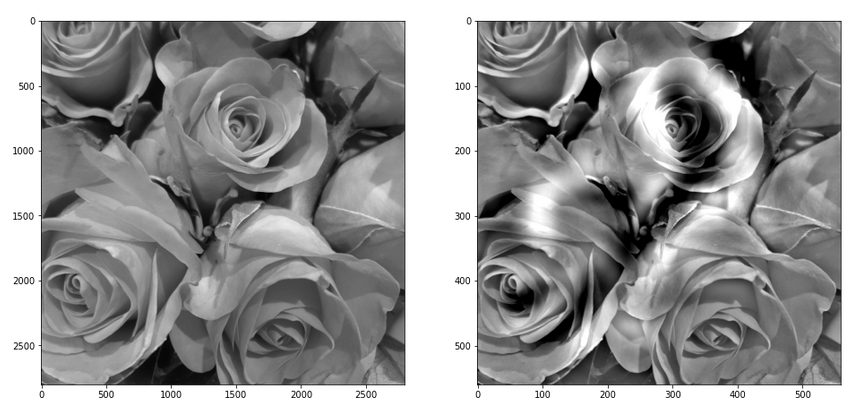
Conclusion
Deep Dreams are based on special image analysis and manipulation techniques. We adapted some of the techniques discussed in literature for high resolution images and advanced CNNs in a special way and for a single deep layer map of a low resolution CNN. We called the required intermittent combination of down- and upscaling, enhancing OIP-patterns and re-adding details “carving“. This helped us to create a “dream” of a CNN – which was trained with a bunch low resolution MNIST images, only; the dream can be based on arbitrary quadratic input images (here: an image of a bunch of roses).
The technique presented in this article can be extended to include a pattern analysis on various sub-parts and length-scales of the original image. In the next blog post we shall have a look at some of the required programming steps.
My_OIP class – V0.65
I have modified the My_OIP code a bit since the last articles on CNNs. You find the new version below.
'''
Module to create OIPs as visualizations of filters of a simple CNN for the MNIST data set
@version: 0.65, 12.12.2020
@change: V0.5: was based on version 0.4 which was originally created in Jupyter cells
@change: V0.6: General revision of class "my_OIP" and its methods
@change: V0.6: Changes to the documentation
@change: V0.65: Added intermediate plotting of selected input fluctuation patterns
@attention: General status: For experimental purposes only!
@requires: A full CNN trained on MNIST data
@requires: A Keras model of the CNN and weight data saved in a h5-file, e.g."cnn_MIST_best.h5".
This file must be placed in the main directory of the Jupyter notebooks.
@requires: A Jupyter environment - from where the class My_OIP is called and where plotting takes place
@note: The description to the interface to the class via the __init__()-method may be incomplete
@note: The use of prefixes li_ and ay_ is not yet consistent. ay_ should indicate numpy arrays, li_ instead normal Python lists
@warning: This version has not been tested outside a Jupyter environment - plotting in GTK/Qt-environment may require substantial changes
@status: Under major development with frequent changes
@author: Dr. Ralph Mönchmeyer
@copyright: Simplified BSD License, 12.12.2020. Copyright (c) 2020, Dr. Ralph Moenchmeyer, Augsburg, Germany
Redistribution and use in source and binary forms, with or without
modification, are permitted provided that the following conditions are met:
1. Redistributions of source code must retain the above copyright notice, this
list of conditions and
the following disclaimer.
2. Redistributions in binary form must reproduce the above copyright notice,
this list of conditions and the following disclaimer in the documentation
and/or other materials provided with the distribution.
THIS SOFTWARE IS PROVIDED BY THE COPYRIGHT HOLDERS AND CONTRIBUTORS "AS IS" AND
ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE IMPLIED
WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE ARE
DISCLAIMED. IN NO EVENT SHALL THE COPYRIGHT OWNER OR CONTRIBUTORS BE LIABLE FOR
ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL, EXEMPLARY, OR CONSEQUENTIAL DAMAGES
(INCLUDING, BUT NOT LIMITED TO, PROCUREMENT OF SUBSTITUTE GOODS OR SERVICES;
LOSS OF USE, DATA, OR PROFITS; OR BUSINESS INTERRUPTION) HOWEVER CAUSED AND
ON ANY THEORY OF LIABILITY, WHETHER IN CONTRACT, STRICT LIABILITY, OR TORT
(INCLUDING NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY OUT OF THE USE OF THIS
SOFTWARE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH DAMAGE.
'''
# Modules to be imported - these libs must be imported in a Jupyter cell nevertheless
# ~~~~~~~~~~~~~~~~~~~~~~~~
import time
import sys
import math
import os
from os import path as path
import numpy as np
from numpy import save # used to export intermediate data
from numpy import load
import scipy
from sklearn.preprocessing import StandardScaler
from itertools import product
import tensorflow as tf
from tensorflow import keras as K
from tensorflow.python.keras import backend as B # this is the only version compatible with TF 1 compat statements
#from tensorflow.keras import backend as B
from tensorflow.keras import models
from tensorflow.keras import layers
from tensorflow.keras import regularizers
from tensorflow.keras import optimizers
from tensorflow.keras.optimizers import schedules
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.datasets import mnist
from tensorflow.python.client import device_lib
import matplotlib as mpl
from matplotlib import pyplot as plt
from matplotlib.colors import ListedColormap
import matplotlib.patches as mpat
from IPython.core.tests.simpleerr import sysexit
class My_OIP:
'''
@summary: This class allows for the creation and the display of OIP-patterns,
to which a selected map of a CNN-model and related filters react maximally
@version: Version 0.6, 10.10.2020
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
@change: Revised methods
@requires: In the present version the class My_OIP requires:
* a CNN-model which works with standardized (!) input images, size 28x28,
* a CNN-Modell which was trained on MNIST digit data,
* exactly 4 length scales for random data fluctations are used to compose initial statistial image data
(the length scales should roughly have a factor of 2 between them)
* Assumption : exactly 1 input image and not a batch of images is assumed in various methods
@note: Main Functions:
0) _init__()
1) _load_cnn_model() => load cnn-model
2) _build_oip_model() => build an oip-model to create OIP-images
3) _setup_gradient_tape_and_iterate_function()
=> Implements TF2 GradientTape to watch input data for eager gradient calculation
=> Creates a convenience function by the help of Keras to iterate and optimize the OIP-adjustments
4) _oip_strat_0_optimization_loop():
=> Method implementing a simple strategy to create OIP-images,
based on superposition of random data on long range data (compared to 28 px)
The optimization uses "gradient ascent"
to get an optimum output of the selected Conv map
6) _derive_OIP(): => Method used to start the creation of an OIP-image for a chosen map
- based on an input image with statistical random date
6) _derive_OIP_for_Prec_Img(): => Method used to start the creation of an OIP-image for a chosen map
- based on an input image with was derived from a PRECURSOR run,
which tests the reaction of the map to large scale fluctuations
7) _build_initial_img_data(): => Builds an input image based on random data for fluctuations on 4 length scales
8) _build_initial_img_from_prec():
=> Reconstruct an input image based on saved random data for long range fluctuations
9) _prepare_precursor(): => Prepare a _precursor run by setting up TF2 GradientTape and the _iterate()-function
10) _precursor(): => Precursor run which systematically tests the reaction of a selected convolutional map
to long range fluctuations based on a (3x3)-grid upscaled to the real image size
11) _display_precursor_imgs(): => A method which plots up to 8 selected precursor images with fluctuations,
which triggered a maximum map reaction
12) _transfrom_tensor_to_img(): => A method which allows to transform tensor data of a standardized (!) image to standard image data
with (gray)pixel valus in [0, 255]. Parameters allow for a contrast enhancement.
Usage hints
~~~~~~~~~~~
@note: If maps of a new convolutional layer are to be investigated then method _build_oip_model(layer_name) has to be rerun
with the layer's name as input parameter
'''
# Method to initialize an instantiation object
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
def __init__(self, cnn_model_file = 'cnn_MNIST_best.h5',
layer_name = 'Conv2D_3',
map_index = 0,
img_dim = 28,
b_build_oip_model = True
):
'''
@summary: Initialization of an object instance - read in a CNN model, build an OIP-Model
@note: Input:
~~~~~~~~~~~~
@param cnn_model_file: Name of a file containing a fully trained CNN-model;
the model can later be overwritten by self._load_cnn_model()
@param layer_name: We can define a layer name, which we are interested in, already when starting;
the layer can later be overwritten by self._build_oip_model()
@param map_index: We can define a map, which we are interested in, already when starting;
A map-index is NOT required for building the OIP-model, but for the GradientTape-object
@param img_dim: The dimension of the assumed quadratic images (2 for MNIST)
@note: Major internal variables:
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
@ivar _cnn_model: A reference to the CNN model object
@ivar _layer_name: The name of convolutional layer
(can be overwritten by method _build_oip_model() )
@ivar _map_index: Index of the map in the chosen layer's output array
(can later be overwritten by other methods)
@ivar _r_cnn_inputs: A reference to the input tensor of the CNN model
Could be a batch of images; but in this class only 1 image is assumed
@ivar _layer_output: Tensor with all maps of a certain layer
@
ivar _oip_submodel: A new model connecting the input of the CNN-model with a certain map's (!) output
@ivar _tape: An instance of TF2's GradientTape-object
Watches input, output, loss of a model
and calculates gradients in TF2 eager mode
@ivar _r_oip_outputs: Reference to the output of the new OIP-model = map-activation
@ivar _r_oip_grads: Reference to gradient tensors for the new OIP-model (output dependency on input image pixels)
@ivar _r_oip_loss: Reference to a loss defined on the OIP-output - i.e. on the activation values of the map's nodes;
Normally chosen to be an average of the nodes' activations
The loss defines a hyperplane on the (28x28)-dim representation space of the input image pixel values
@ivar _val_oip_loss: Loss value for a certain input image
@ivar _iterate Reference toa Keras backend function which invokes the new OIP-model for a given image
and calculates both loss and gradient values (in TF2 eager mode)
This is the function to be used in the optimization loop for OIPs
@note: Internal Parameters controlling the optimization loop:
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
@ivar _oip_strategy: 0, 1 - There are two strategies to evolve OIP patterns out of statistical data
- only the first strategy is supported in this version
Both strategies can be combined with a precursor calculation
0: Simple superposition of fluctuations at different length scales
1: NOT YET SUPPORTED
Evolution over partially evolved images based on longer scale variations
enriched with fluctuations on shorter length scales
@ivar _ay_epochs: A list of 4 optimization epochs to be used whilst
evolving the img data via strategy 1 and intermediate images
@ivar _n_epochs: Number of optimization epochs to be used with strategy 0
@ivar _n_steps: Defines at how many intermediate points we show images and report
during the optimization process
@ivar _epsilon: Factor to control the amount of correction imposed by the gradient values of the oip-model
@note: Input image data of the OIP-model and references to it
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
@ivar _initial_precursor_img: The initial image to start a precursor optimization with.
Would normally be an image of only long range fluctuations.
@ivar _precursor_image: The evolved image created and selected by the precursor loop
@ivar _initial_inp_img_data: A tensor representing the data of the input image
@ivar _inp_img_data: A tensor representig the
@ivar _img_dim: We assume quadratic images to work with
with dimension _img_dim along each axis
For the time being we only support MNIST images
@note: Internal parameters controlling the composition of random initial image data
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
@ivar _li_dim_steps: A list of the intermediate dimensions for random data;
these data are smoothly scaled to the image dimensions
@ivar _ay_facts: A Numpy array of 4 factors to control the amount of
contribution of the statistical variations
on the 4 length scales to the initial image
@note: Internal variables to save data of a precursor run
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
@ivar _list_of_covs: list of long range fluctuation data for a (3x3)-grid covering the image area
@ivar _li_fluct_enrichments: [li_facts, li_dim_steps] data for enrichment with small fluctuations
@note: Internal variables for plotting
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
@ivar _li_axa: A Python list of references to external (Jupyter-) axes-frames for plotting
'''
# Input data and variable initializations
# ****************************************
# the model
# ~~~~~~~~~~
self._cnn_model_file = cnn_model_file
self._cnn_model = None
# the chosen layer of the CNN-model
self._layer_name = layer_name
# the index of the map in the layer array
self._map_index = map_index
# References to objects and the OIP-submodel
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
self._r_cnn_inputs = None # reference to input of the CNN_model, also used in the oip-model
self._layer_output = None
self._oip_submodel = None
# References to watched GradientTape objects
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
self._tape = None # TF2 GradientTape variable
# some "references"
self._r_oip_outputs = None # output of the oip-submodel to be watched
self._r_oip_grads = None # gradients determined by GradientTape
self._r_oip_loss = None # loss function
# loss and gradient values (to be produced ba a backend function _iterate() )
self._val_oip_grads = None
self._val_oip_loss = None
# The Keras function to produce concrete outputs of the new OIP-model
self._iterate = None
# The strategy to produce an OIP pattern out of statistical input images
# ~~~~~~~~~~~~~~~~~~~~~~~~~--------~~~~~~
# 0: Simple superposition of fluctuations at different length scales
# 1: Move over 4 interediate images - partially optimized
self._oip_strategy = 0
# Parameters controlling the OIP-optimization process
# ~~~~~~~~~~~~~~~~~~~~~~~~~--------~~~~~~
# number of epochs for optimization strategy 1
self._ay_epochs = np.array((20, 40, 80, 400), dtype=np.int32)
len_epochs = len(self._ay_epochs)
# number of epochs for optimization strategy 0
self._n_epochs = self._ay_epochs[len_epochs-1]
self._n_steps = 6 # divides the number of n_epochs into n_steps to produce intermediate outputs
# size of corrections by gradients
self._epsilon = 0.01 # step-size for gradient correction
# Input image-typess and references
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
# precursor image
self._initial_precursor_img = None
self._precursor_img = None # output of the _precursor-method
# The input image for the OIP-creation - a superposition of inial random fluctuations
self._initial_inp_img_data = None # The initial data constructed
self._inp_img_data = None # The data used and varied for optimization
# image dimension
self._img_dim = img_dim # = 28 => MNIST images for the time being
# Parameters controlling the setup of an initial image
# ~~~~~~~~~~~~~~~~~~~~~~~~~--------
~~~~~~~~~~~~~~~~~~~
# The length scales for initial input fluctuations
self._li_dim_steps = ( (3, 3), (7,7), (14,14), (28,28) ) # can later be overwritten
# Parameters for fluctuations - used both in strategy 0 and strategy 1
self._ay_facts = np.array( (0.5, 0.5, 0.5, 0.5), dtype=np.float32 )
# Data of a _precursor()-run
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~
self._list_of_covs = None # list of long range fluctuation data for a (3x3)-grid covering the image area
self._li_fluct_enrichments = None # = [li_facts, li_dim_steps] list with with 2 list of data enrichment for small fluctuations
# These data are required to reconstruct the input image to which a map reacted
# List of references to axis subplots
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
# These references may change from Jupyter cell to Jupyter cell and provided by the called methods
self._li_axa = None # will be set by methods according to axes-frames in Jupyter cells
# axis frames for a single image in 2 versions (with contrast enhancement)
self._ax1_1 = None
self._ax1_2 = None
# Variables for Deep Dream Analysis
# ***********************************
# Here we need a stack of tiled sub-images of a large image, as we subdivide on different length scales
self._num_tiles = 1
self._t_image_stack = None
# ********************************************************
# Model setup - load the cnn-model and build the OIP-model
# ************
if path.isfile(self._cnn_model_file):
# We trigger the initial load of a model
self._load_cnn_model(file_of_cnn_model = self._cnn_model_file, b_print_cnn_model = True)
# We trigger the build of a new sub-model based on the CNN model used for OIP search
self._build_oip_model(layer_name = self._layer_name, b_print_oip_model = True )
else:
print("<\nWarning: The standard file " + self._cnn_model_file +
" for the cnn-model could not be found!\n " +
" Please use method _load_cnn_model() to load a valid model")
sys.exit()
return
#
# Method to load a specific CNN model
# **********************************
def _load_cnn_model(self, file_of_cnn_model=None, b_print_cnn_model=True ):
'''
@summary: Method to load a CNN-model from a h5-file and create a reference to its input (image)
@version: 0.2 of 28.09.2020
@requires: filename must already have been saved in _cnn_model_file or been given as a parameter
@requires: file must be a h5-file
@change: minor changes - documentation
@note: A reference to the CNN's input is saved internally
@warning: An input in form of an image - a MNIST-image - is implicitly assumed
@note: Parameters
-----------------
@param file_of_cnn_model: Name of h5-file with the trained (!) CNN-model
@param b_print_cnn_model: boolean - Print some information on the CNN-model
'''
if file_of_cnn_model != None:
self._cnn_model_file = file_of_cnn_model
# Check existence of the file
if not path.isfile(self._cnn_model_file):
print("\nWarning: The file " + file_of_cnn_model +
" for the cnn-model could not be found!\n" +
"Please change the parameter \"file_of_cnn_model\"" +
" to load a valid model")
# load the CNN model
self._cnn_model = models.load_model(self._cnn_model_file)
# Inform about the model and its
file
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~
print("Used file to load a ´ model = ", self._cnn_model_file)
# we print out the models structure
if b_print_cnn_model:
print("Structure of the loaded CNN-model:\n")
self._cnn_model.summary()
# handle/references to the models input => more precise the input image
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
# Note: As we only have one image instead of a batch
# we pick only the first tensor element!
# The inputs will be needed for buildng the oip-model
self._r_cnn_inputs = self._cnn_model.inputs[0] # !!! We have a btach with just ONE image
# print out the shape - it should be known from the original cnn-model
if b_print_cnn_model:
print("shape of cnn-model inputs = ", self._r_cnn_inputs.shape)
return
#
# Method to construct a model to optimize input for OIP creation
# ***************************************
def _build_oip_model(self, layer_name = 'Conv2D_3', b_print_oip_model=True ):
'''
@summary: Method to build a new (sub-) model - the "OIP-model" - of the CNN-model by
connectng the input of the CNN-model with one of its Conv-layers
@version: 0.4 of 28.09.2020
@change: Minor changes - documentation
@note: We need a Conv layer to build a working model for input image optimization
We get the Conv layer by the layer's name
The new model connects the first input element of the CNN to the output maps of the named Conv layer CNN
We use Keras' models.Model() functionality
@note: The layer's name is crucial for all later investigations - if you want to change it this method has to be rerun
@requires: The original, trained CNN-model must be loaded and referenced by self._cnn_model
@warning: Only 1 input image and not a batch is implicitly assumed
@note: Parameters
-----------------
@param layer_name: Name of the convolutional layer of the CNN for whose maps we want to find OIP patterns
@param b_print_oip_model: boolean - Print some information on the OIP-model
'''
# free some RAM - hopefully
del self._oip_submodel
# check for loaded CNN-model
if self._cnn_model == None:
print("Error: CNN-model not yet defined.")
sys.exit()
# get layer name
self._layer_name = layer_name
# We build a new model based on the model inputs and the output
self._layer_output = self._cnn_model.get_layer(self._layer_name).output
# Note: We do not care at the moment about a complex composition of the input
# We trust in that we handle only one image - and not a batch
# Create the sub-model via Keras' models.Model()
model_name = "mod_oip__" + layer_name
self._oip_submodel = models.Model( [self._r_cnn_inputs], [self._layer_output], name = model_name)
# We print out the oip model structure
if b_print_oip_model:
print("Structure of the constructed OIP-sub-model:\n")
self._oip_submodel.summary()
return
#
# Method to set up GradientTape and an iteration function providing loss and gradient values
# *********************************************************************************************
def _setup_gradient_tape_and_iterate_function(self, b_print = True, b_full_layer = False):
'''
@summary: Central method to watch input variables and resulting gradient changes
@version: 0.6 of 12.12.2020
n @change: V0.6: We add the definition of a loss function for a whole layer
@note: For TF2 eager execution we need to watch input changes and trigger automatic gradient evaluation
@note: The normalization of the gradient is strongly recommended; as we fix epsilon for correction steps
we thereby will get changes to the input data of an approximately constant order.
This - together with standardization of the images (!) - will lead to convergence at the size of epsilon !
@param b_full_layer: Boolean; define a loss function for a single map (map_index > -1) or the full layer (map_index = -1)
'''
# Working with TF2 GradientTape
self._tape = None
# Watch out for input, output variables with respect to gradient changes
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
with tf.GradientTape() as self._tape:
# Input
# ~~~~~
self._tape.watch(self._r_cnn_inputs)
# Output
# ~~~~~~
self._r_oip_outputs = self._oip_submodel(self._r_cnn_inputs)
# only for testing
# *******************
#t_shape = self._r_oip_outputs[0, :, :, :].shape # the 0 is important, else "Noen" => problems
#scaling_factor = tf.cast(tf.reduce_prod(t_shape), tf.float32)
#tf.print("scaling_factor: ", scaling_factor)
#t_loss = tf.reduce_sum(tf.math.square(self._r_oip_outputs[0, :, :, :])) / scaling_factor
# Define Loss
# ***********
if (b_full_layer):
# number of neurons over all layer maps
t_shape = self._r_oip_outputs[0, :, :, :].shape # the 0 is important, else "Noen" => problems
scaling_factor = tf.cast(tf.reduce_prod(t_shape), tf.float32) # shape! eg. 3x3x128 for layer 3
tf.print("scaling_factor: ", scaling_factor)
# The scaled total loss of the layer
self._r_oip_loss = tf.reduce_sum(tf.math.square(self._r_oip_outputs[0, :, :, :])) / scaling_factor
# self._r_oip_loss = tf.reduce_mean(self._r_oip_outputs[0, :, :, self._map_index])
else:
self._r_oip_loss = tf.reduce_mean(self._r_oip_outputs[0, :, :, self._map_index])
#self._loss = B.mean(oip_output[:, :, :, map_index])
#self._loss = B.mean(oip_outputs[-1][:, :, map_index])
#self._loss = tf.reduce_mean(oip_outputs[-1][ :, :, map_index])
if b_print:
print(self._r_oip_loss)
print("shape of oip_loss = ", self._r_oip_loss.shape)
# Gradient definition and normalization
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
self._r_oip_grads = self._tape.gradient(self._r_oip_loss, self._r_cnn_inputs)
print("shape of grads = ", self._r_oip_grads.shape)
# normalization of the gradient - required for convergence
t_tiny = tf.constant(1.e-7, tf.float32)
self._r_oip_grads /= (tf.sqrt(tf.reduce_mean(tf.square(self._r_oip_grads))) + t_tiny)
#self._r_oip_grads /= (B.sqrt(B.mean(B.square(self._r_oip_grads))) + 1.e-7)
#self._r_oip_grads = tf.image.per_image_standardization(self._r_oip_grads)
# Define an abstract recallable Keras function as a convenience function for iterations
# The _iterate()-function produces loss and gradient values for corrected img data
# The first list of addresses points to the input data, the last to the output data
self._iterate = B.function( [self._r_cnn_inputs], [self._r_oip_loss, self._r_oip_grads] )
#
# Method to optimize an emerging OIP out of statistical input
image data
# (simple strategy - just optimize, no precursor, no intermediate pattern evolution, ..)
# ********************************
def _oip_strat_0_optimization_loop(self, conv_criterion = 5.e-4,
b_stop_with_convergence = False,
b_show_intermediate_images = True,
b_print = True):
'''
@summary: Method to control the optimization loop for OIP reconstruction of an initial input image
with a statistical distribution of pixel values.
@version: 0.4 of 28.09.2020
@changes: Minor changes - eliminated some unused variables
@note: The function also provides intermediate output in the form of printed data and images !.
@requires: An input image tensor must already be available at _inp_img_data - created by _build_initial_img_data()
@requires: Axis-objects for plotting, typically provided externally by the calling functions
_derive_OIP() or _precursor()
@note: Parameters:
-----------------
@param conv_criterion: A small threshold number for (difference of loss-values / present loss value )
@param b_stop_with_convergence: Booelan which decides whether we stop a loop if the conv-criterion is fulfilled
@param b_show_intermediate_image: Boolean which decides whether we show up to 8 intermediate images
@param b_print: Boolean which decides whether we print intermediate loss values
@note: Intermediate information is provided at _n_steps intervals,
which are logarithmically distanced with respect to _n_epochs
Reason: Most changes happen at the beginning
@note: This method produces some intermediate output during the optimization loop in form of images.
It uses an external grid of plot frames and their axes-objects. The addresses of the
axis-objects must provided by an external list "li_axa[]" to self._li_axa[].
We need a seqence of _n_steps+2 axis-frames (or more) => length(_li_axa) >= _n_steps + 2).
@todo: Loop not optimized for TF 2 - but not so important here - a run takes less than a second
'''
# Check that we already an input image tensor
if ( (self._inp_img_data == None) or
(self._inp_img_data.shape[1] != self._img_dim) or
(self._inp_img_data.shape[2] != self._img_dim) ) :
print("There is no initial input image or it does not fit dimension requirements (28,28)")
sys.exit()
# Print some information
if b_print:
print("*************\nStart of optimization loop\n*************")
print("Strategy: Simple initial mixture of long and short range variations")
print("Number of epochs = ", self._n_epochs)
print("Epsilon = ", self._epsilon)
print("*************")
# some initial value
loss_old = 0.0
# Preparation of intermediate reporting / img printing
# --------------------------------------
# Logarithmic spacing of steps (most things happen initially)
n_el = math.floor(self._n_epochs / 2**(self._n_steps) )
li_int = []
if n_el != 0:
for j in range(self._n_steps):
li_int.append(n_el*2**j)
else: # linear spacing
n_el = math.floor(self._n_epochs / (self._n_steps+1) )
for j in range(self._n_steps+1):
li_int.append(n_el*j)
if b_print:
print("li_int = ", li_int)
# A counter for intermediate reporting
n_rep = 0
#
Convergence? - A list for steps meeting the convergence criterion
# ~~~~~~~~~~~~
li_conv = []
# optimization loop
# *******************
# counter for steps with zero loss and gradient values
n_zeros = 0
for j in range(self._n_epochs):
# Get output values of our Keras iteration function
# ~~~~~~~~~~~~~~~~~~~
self._val_oip_loss, self._val_oip_grads = self._iterate([self._inp_img_data])
# loss difference to last step - shuold steadily become smaller
loss_diff = self._val_oip_loss - loss_old
#if b_print:
# print("j: ", j, " :: loss_val = ", self._val_oip_loss, " :: loss_diff = ", loss_diff)
# # print("loss_diff = ", loss_diff)
loss_old = self._val_oip_loss
if j > 10 and (loss_diff/(self._val_oip_loss + 1.-7)) < conv_criterion:
li_conv.append(j)
lenc = len(li_conv)
# print("conv - step = ", j)
# stop only after the criterion has been met in 4 successive steps
if b_stop_with_convergence and lenc > 5 and li_conv[-4] == j-4:
return
grads_val = self._val_oip_grads
#grads_val = normalize_tensor(grads_val)
# The gradients average value
avg_grads_val = (tf.reduce_mean(grads_val)).numpy()
# Check if our map reacts at all
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
if self._val_oip_loss == 0.0 and avg_grads_val == 0.0 and b_print :
print( "0-values, j= ", j,
" loss = ", self._val_oip_loss, " avg_loss = ", avg_grads_val )
n_zeros += 1
if n_zeros > 10 and b_print:
print("More than 10 times zero values - Try a different initial random distribution of pixel values")
return
# gradient ascent => Correction of the input image data
# ~~~~~~~~~~~~~~~
self._inp_img_data += self._val_oip_grads * self._epsilon
# Standardize the corrected image - we won't get a convergence otherwise
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
self._inp_img_data = tf.image.per_image_standardization(self._inp_img_data)
# Some output at intermediate points
# Note: We us logarithmic intervals because most changes
# appear in the intial third of the loop's range
if (j == 0) or (j in li_int) or (j == self._n_epochs-1) :
if b_print or (j == self._n_epochs-1):
# print some info
print("\nstep " + str(j) + " finalized")
#print("Shape of grads = ", grads_val.shape)
print("present loss_val = ", self._val_oip_loss)
print("loss_diff = ", loss_diff)
# show the intermediate image data
if b_show_intermediate_images:
imgn = self._inp_img_data[0,:,:,0].numpy()
# print("Shape of intermediate img = ", imgn.shape)
self._li_axa[n_rep].imshow(imgn, cmap=plt.cm.get_cmap('viridis'))
# counter
n_rep += 1
return
#
# Standard UI-method to derive OIP from a given initial input image
# ********************
def _derive_OIP(self, map_index = 1,
n_epochs = None,
n_steps = 6,
epsilon =
0.01,
conv_criterion = 5.e-4,
li_axa = [],
ax1_1 = None, ax1_2 = None,
b_stop_with_convergence = False,
b_show_intermediate_images = True,
b_print = True,
centre_move = 0.33, fact = 1.0):
'''
@summary: Method to create an OIP-image for a given initial input image
This is the standard user interface for finding an OIP
@warning: This method should NOT be used for finding an initial precursor image
Use _prepare_precursor() to define the map first and then _precursor() to evaluate initial images
@version: V0.5, 12.12.2020
@change: V0.4: Minor changes - added internal _li_axa for plotting, added documentation
This method starts the process of producing an OIP of statistical input image data
@change: V0.5 : map_index can now have the value "-". Then a loss function for the whole layer (instead of a single map) is prepared
@requires: A map index (or -1) should be provided to this method
@requires: An initial input image with statistical fluctuations of pixel values must have been created.
@warning: This version only supports the most simple strategy - "strategy 0"
------------- Optimize in one loop - starting from a superposition of fluctuations
No precursor, no intermediate evolutions
@note: Parameters:
-----------------
@param map_index: We can and should chose a map here (overwrites previous settings)
If map_index == -1 => We take a loss for the whole layer
@param n_epochs: Number of optimization steps (overwrites previous settings)
@param n_steps: Defines number of intervals (with length n_epochs/ n_steps) for reporting
standard value: 6 => 8 images - start image, end image + 6 intermediate
This number also sets a requirement for providing (n_step + 2) external axis-frames
to display intermediate images of the emerging OIP
=> see _oip_strat_0_optimization_loop()
@param epsilon: Size for corrections by gradient values
@param conv_criterion: A small threshold number for convegenc (checks: difference of loss-values / present loss value )
@param b_stop_with_convergence:
Booelan which decides whether we stop a loop if the conv-criterion is fulfilled
@param _li_axa: A Python list of references to external (Jupyter-) axes-frames for plotting
@note: Preparations for plotting:
We need n_step + 2 axis-frames which must be provided externally
With Jupyter this can externally be done by statements like
# figure
# -----------
#sizing
fig_size = plt.rcParams["figure.figsize"]
fig_size[0] = 16
fig_size[1] = 8
fig_a = plt.figure()
axa_1 = fig_a.add_subplot(241)
axa_2 = fig_a.add_subplot(242)
axa_3 = fig_a.add_subplot(243)
axa_4 = fig_a.add_subplot(244)
axa_5 = fig_a.add_subplot(245)
axa_6 = fig_a.add_subplot(246)
axa_7 = fig_a.add_subplot(247)
axa_8 = fig_a.add_subplot(248)
li_axa = [axa_1, axa_2, axa_3, axa_4, axa_5, axa_6, axa_7, axa_8]
'''
# Get input parameters
self._map_index = map_index
self._n_epochs = n_epochs
self._n_steps = n_steps
self._epsilon = epsilon
# references to plot frames
self._li_axa = li_axa
num_frames = len(li_
axa)
if num_frames < n_steps+2:
print("The number of available image frames (", num_frames, ") is smaller than required for intermediate output (", n_steps+2, ")")
sys.exit()
# 2 axes frames to display the final OIP image (with contrast enhancement)
self._ax1_1 = ax1_1
self._ax1_2 = ax1_2
# number of epochs for optimization strategy 0
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
if n_epochs == None:
len_epochs = len(self._ay_epochs)
self._n_epochs = self._ay_epochs[len_epochs-1]
else:
self._n_epochs = n_epochs
# Reset some variables
self._val_oip_grads = None
self._val_oip_loss = None
self._iterate = None
# Setup the TF2 GradientTape watch and a Keras function for iterations
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
# V0.5 => map_index == "-1"
if self._map_index == -1:
self._setup_gradient_tape_and_iterate_function(b_print = b_print, b_full_layer=True)
if b_print:
print("GradientTape watch activated (map-based: map = ", self._map_index, ")")
else:
self._setup_gradient_tape_and_iterate_function(b_print = b_print, b_full_layer=False)
if b_print:
print("GradientTape watch activated (layer-based: layer = ", self._layer_name, ")")
'''
# Gradient handling - so far we only deal with addresses
# ~~~~~~~~~~~~~~~~~~
self._r_oip_grads = self._tape.gradient(self._r_oip_loss, self._r_cnn_inputs)
print("shape of grads = ", self._r_oip_grads.shape)
# normalization of the gradient
self._r_oip_grads /= (B.sqrt(B.mean(B.square(self._r_oip_grads))) + 1.e-7)
#self._r_oip_grads = tf.image.per_image_standardization(self._r_oip_grads)
# define an abstract recallable Keras function
# producing loss and gradients for corrected img data
# the first list of addresses points to the input data, the last to the output data
self._iterate = B.function( [self._r_cnn_inputs], [self._r_oip_loss, self._r_oip_grads] )
'''
# get the initial image into a variable for optimization
self._inp_img_data = self._initial_inp_img_data
# Start optimization loop for strategy 0
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
if self._oip_strategy == 0:
self._oip_strat_0_optimization_loop( conv_criterion = conv_criterion,
b_stop_with_convergence = b_stop_with_convergence,
b_show_intermediate_images = b_show_intermediate_images,
b_print = b_print
)
# Display the last OIP-image created at the end of the optimization loop
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
# standardized image
oip_img = self._inp_img_data[0,:,:,0].numpy()
# transformed image
oip_img_t = self._transform_tensor_to_img(T_img=self._inp_img_data[0,:,:,0],
centre_move = centre_move,
fact = fact)
# display
ax1_1.imshow(oip_img, cmap=plt.cm.get_cmap('viridis'))
ax1_2.imshow(oip_img_t, cmap=plt.cm.get_cmap('viridis'))
return
#
# Method to derive OIP from a given initial input image if map_index is already defined
# ********************
def _derive_OIP_for_Prec_
Img(self,
n_epochs = None,
n_steps = 6,
epsilon = 0.01,
conv_criterion = 5.e-4,
li_axa = [],
ax1_1 = None, ax1_2 = None,
b_stop_with_convergence = False,
b_show_intermediate_images = True,
b_print = True):
'''
@summary: Method to create an OIP-image for an already given map-index and a given initial input image
This is the core of OIP-detection, which starts the optimization loop
@warning: This method should NOT be used directly for finding an initial precursor image
Use _prepare_precursor() to define the map first and then _precursor() to evaluate initial images
@version: V0.4, 28.09.2020
@changes: Minor changes - added internal _li_axa for plotting, added documentation
This method starts the process of producing an OIP of statistical input image data
@note: This method should only be called after _prepare_precursor(), _precursor(), _build_initial_img_prec()
For a trial of different possible precursor images rerun _build_initial_img_prec() with a different index
@requires: A map index should be provided to this method
@requires: An initial input image with statistical fluctuations of pixel values must have been created.
@warning: This version only supports the most simple strategy - "strategy 0"
------------- Optimize in one loop - starting from a superposition of fluctuations
no intermediate evolutions
@note: Parameters:
-----------------
@param n_epochs: Number of optimization steps (overwrites previous settings)
@param n_steps: Defines number of intervals (with length n_epochs/ n_steps) for reporting
standard value: 6 => 8 images - start image, end image + 6 intermediate
This number also sets a requirement for providing (n_step + 2) external axis-frames
to display intermediate images of the emerging OIP
=> see _oip_strat_0_optimization_loop()
@param epsilon: Size for corrections by gradient values
@param conv_criterion: A small threshold number for convegenc (checks: difference of loss-values / present loss value )
@param b_stop_with_convergence:
Booelan which decides whether we stop a loop if the conv-criterion is fulfilled
@param _li_axa: A Python list of references to external (Jupyter-) axes-frames for plotting
@note: Preparations for plotting:
We need n_step + 2 axis-frames which must be provided externally
With Jupyter this can externally be done by statements like
# figure
# -----------
#sizing
fig_size = plt.rcParams["figure.figsize"]
fig_size[0] = 16
fig_size[1] = 8
fig_a = plt.figure()
axa_1 = fig_a.add_subplot(241)
axa_2 = fig_a.add_subplot(242)
axa_3 = fig_a.add_subplot(243)
axa_4 = fig_a.add_subplot(244)
axa_5 = fig_a.add_subplot(245)
axa_6 = fig_a.add_subplot(246)
axa_7 = fig_a.add_subplot(247)
axa_8 = fig_a.add_subplot(248)
li_axa = [axa_1, axa_2, axa_3, axa_4, axa_5, axa_6, axa_7, axa_8]
'''
# Get input parameters
self._n_epochs = n_epochs
self._n_steps = n_steps
self._epsilon = epsilon
# references to plot frames
self._li_axa = li_axa
num_frames
= len(li_axa)
if num_frames < n_steps+2:
print("The number of available image frames (", num_frames, ") is smaller than required for intermediate output (", n_steps+2, ")")
sys.exit()
# 2 axes frames to display the final OIP image (with contrast enhancement)
self._ax1_1 = ax1_1
self._ax1_2 = ax1_2
# number of epochs for optimization strategy 0
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
if n_epochs == None:
len_epochs = len(self._ay_epochs)
self._n_epochs = self._ay_epochs[len_epochs-1]
else:
self._n_epochs = n_epochs
# Note: No setup of GradientTape and _iterate(required) - this is done by _prepare_precursor
# get the initial image into a variable for optimization
self._inp_img_data = self._initial_inp_img_data
# Start optimization loop for strategy 0
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
if self._oip_strategy == 0:
self._oip_strat_0_optimization_loop( conv_criterion = conv_criterion,
b_stop_with_convergence = b_stop_with_convergence,
b_show_intermediate_images = b_show_intermediate_images,
b_print = b_print
)
# Display the last OIP-image created at the end of the optimization loop
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
# standardized image
oip_img = self._inp_img_data[0,:,:,0].numpy()
# transfored image
oip_img_t = self._transform_tensor_to_img(T_img=self._inp_img_data[0,:,:,0])
# display
ax1_1.imshow(oip_img, cmap=plt.cm.get_cmap('viridis'))
ax1_2.imshow(oip_img_t, cmap=plt.cm.get_cmap('viridis'))
return
#
# Method to build an initial image from a superposition of random data on different length scales
# ***********************************
def _build_initial_img_data( self,
strategy = 0,
li_epochs = [20, 50, 100, 400],
li_facts = [0.5, 0.5, 0.5, 0.5],
li_dim_steps = [ (3,3), (7,7), (14,14), (28,28) ],
b_smoothing = False,
ax1_1 = None, ax1_2 = None):
'''
@summary: Standard method to build an initial image with random fluctuations on 4 length scales
@version: V0.2 of 29.09.2020
@note: This method should NOT be used for initial images based on a precursor images.
See _build_initial_img_prec(), instead.
@note: This method constructs an initial input image with a statistical distribution of pixel-values.
We use 4 length scales to mix fluctuations with different "wave-length" by a simple approach:
We fill four square with a different numer of cells below the number of pixels
in each dimension of the real input image; e.g. (4x4), (7x7, (14x14), (28,28) <= (28,28).
We fill the cells with random numbers in [-1.0, 1.]. We smootly scale the resulting pattern
up to (28,28) (or what ever the input image dimensions are) by bicubic interpolations
and eventually add up all values. As a final step we standardize the pixel value distribution.
@warning: This version works with 4 length scales, only.
@warning: In the present version th eparameters "strategy " and li_epochs have no effect
@note: Parameters:
-----------------
r
@param strategy: The strategy, how to build an image (overwrites previous settings) - presently only 0 is supported
@param li_epochs: A list of epoch numbers which will be used in strategy 1 - not yet supported
@param li_facts: A list of factors which the control the relative strength of the 4 fluctuation patterns
@param li_dim_steps: A list of square dimensions for setting the length scale of the fluctuations
@param b_smoothing: Parameter which builds a control image
@param ax1_1: matplotlib axis-frame to display the built image
@param ax1_2: matplotlib axis-frame to display a second version of the built image
'''
# Get input parameters
# ~~~~~~~~~~~~~~~~~~
self._oip_strategy = strategy # no effect in this version
self._ay_epochs = np.array(li_epochs) # no effect in this version
# factors by which to mix the random number fluctuations of the different length scales
self._ay_facts = np.array(li_facts)
# dimensions of the squares which simulate fluctuations
self._li_dim_steps = li_dim_steps
# A Numpy array for the eventual superposition of random data
fluct_data = None
# Strategy 0: Simple superposition of random patterns at 4 different wave-length
# ~~~~~~~~~~
if self._oip_strategy == 0:
dim_1_1 = self._li_dim_steps[0][0]
dim_1_2 = self._li_dim_steps[0][1]
dim_2_1 = self._li_dim_steps[1][0]
dim_2_2 = self._li_dim_steps[1][1]
dim_3_1 = self._li_dim_steps[2][0]
dim_3_2 = self._li_dim_steps[2][1]
dim_4_1 = self._li_dim_steps[3][0]
dim_4_2 = self._li_dim_steps[3][1]
fact1 = self._ay_facts[0]
fact2 = self._ay_facts[1]
fact3 = self._ay_facts[2]
fact4 = self._ay_facts[3]
# print some parameter information
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
print("\nInitial image composition - strategy 0:\n Superposition of 4 different wavelength patterns")
print("Parameters:\n",
fact1, " => (" + str(dim_1_1) +", " + str(dim_1_2) + ") :: ",
fact2, " => (" + str(dim_2_1) +", " + str(dim_2_2) + ") :: ",
fact3, " => (" + str(dim_3_1) +", " + str(dim_3_2) + ") :: ",
fact4, " => (" + str(dim_4_1) +", " + str(dim_4_2) + ")"
)
# fluctuations
fluct1 = 2.0 * ( np.random.random((1, dim_1_1, dim_1_2, 1)) - 0.5 )
fluct2 = 2.0 * ( np.random.random((1, dim_2_1, dim_2_2, 1)) - 0.5 )
fluct3 = 2.0 * ( np.random.random((1, dim_3_1, dim_3_2, 1)) - 0.5 )
fluct4 = 2.0 * ( np.random.random((1, dim_4_1, dim_4_2, 1)) - 0.5 )
# Scaling with bicubic interpolation to the required image size
fluct1_scale = tf.image.resize(fluct1, [28,28], method="bicubic", antialias=True)
fluct2_scale = tf.image.resize(fluct2, [28,28], method="bicubic", antialias=True)
fluct3_scale = tf.image.resize(fluct3, [28,28], method="bicubic", antialias=True)
fluct4_scale = fluct4
# superposition
fluct_data = fact1*fluct1_scale + fact2*fluct2_scale + fact3*fluct3_scale + fact4*fluct4_scale
# get the standardized plus smoothed and unsmoothed image
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
# TF2 provides a function performing standardization of image data function
fluct_data_unsmoothed = tf.image.per_image_
standardization(fluct_data)
fluct_data_smoothed = tf.image.per_image_standardization(
tf.image.resize( fluct_data, [28,28], method="bicubic", antialias=True) )
if b_smoothing:
self._initial_inp_img_data = fluct_data_smoothed
else:
self._initial_inp_img_data = fluct_data_unsmoothed
# Display of both variants => there should be no difference
# ~~~~~~~~~~~~~~~~~~~~~~~~
img_init_unsmoothed = fluct_data_unsmoothed[0,:,:,0].numpy()
img_init_smoothed = fluct_data_smoothed[0,:,:,0].numpy()
ax1_1.imshow(img_init_unsmoothed, cmap=plt.cm.get_cmap('viridis'))
ax1_2.imshow(img_init_smoothed, cmap=plt.cm.get_cmap('viridis'))
print("Initial images plotted")
return self._initial_inp_img_data
#
# Method to build an initial image from a superposition of a PRECURSOR image with random data on different length scales
# ***********************************
def _build_initial_img_from_prec( self,
prec_index = -1,
strategy = 0,
li_epochs = (20, 50, 100, 400),
li_facts = (1.0, 0.0, 0.0, 0.0, 0.0),
li_dim_steps = ( (3,3), (7,7), (14,14), (28,28) ),
b_smoothing = False,
b_print = True,
b_display = False,
ax1_1 = None, ax1_2 = None):
'''
@summary: Method to build an initial image based on a Precursor image => Input for _derive_OIP_for_Prec_Img()
@version: V0.3, 03.10.2020
@changes: V0.2: Minor- only documentation and comparison of index to length of _li_of_flucts[]
@changes: V0.3: Added Booleans to control the output and display of images
@changes: V0.4: Extended the reconstruction part / extended documentation
@note: This method differs from _build_initial_img_data() as follows:
* It uses a Precursor image as the fundamental image
* The data of the Precursor Image will be reconstructed from a (3x3) fluctuation pattern and enrichments
* This method adds even further fluctuations if requested
@note: This method should be called manually from a Jupyter cell
@note: This method saves the reconstructed input image into self._initial_inp_img_data
@note: This method should be followed by a call of self._derive_OIP_for_Prec_Img()
@requires: Large scale fluctuation data saved in self._li_of_flucts[]
@requires: Additional enrichment information in self._li_of_fluct_enrichments[]
@param prec_index: This is an index ([0, 7[) of a large scale fluctuation pattern which was saved in self._li_of_flucts[]
during the run of the method "_precursor()". The image tensor is reconstructed from the fluctuation pattern.
@warning: We support a maximum of 8 selected fluctuation patterns for which a map reacts
@warning: However, less precursor patterns may be found - so you should watch for the output of _precursor() before you run this method
@param li_facts: A list of factors which the control the relative strength of the precursor image vs.
4 additional fluctuation patterns
@warning: Normally, it makes no sense to set li_facts[1] > 0 - because this will destroy the original large scale pattern
@note: For other input parameters see _build_initial_img_data()
'''
# Get input parameters
self._oip_strategy
= strategy
self._ay_facts = np.array(li_facts)
self._ay_epochs = np.array(li_epochs)
self._li_dim_steps = li_dim_steps
fluct_data = None
# Reconstruct an precursor image from a saved large scale fluctuation pattern (result of _precursor() )
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
length_li_cov = len(self._li_of_flucts)
if prec_index > -1:
if length_li_cov < prec_index+1:
print("index too large for number of detected patterns (", length_li_cov, ")")
sys.exit()
cov_p = self._li_of_flucts[prec_index][1][1]
fluct0_scale_p = tf.image.resize(cov_p, [28,28], method="bicubic", antialias=True)
# Add fluctuation enrichments - if saved [ len(self._li_fluct_enrichments) > 0 ]
if len(self._li_fluct_enrichments) > 0:
# Scaling enrichment flucts with bicubic interpolation to the required image size
fluct1_p = self._li_fluct_enrichments[2][0]
fluct2_p = self._li_fluct_enrichments[2][1]
fluct3_p = self._li_fluct_enrichments[2][2]
fluct4_p = self._li_fluct_enrichments[2][3]
fact0_p = self._li_fluct_enrichments[0][0]
fact1_p = self._li_fluct_enrichments[0][1]
fact2_p = self._li_fluct_enrichments[0][2]
fact3_p = self._li_fluct_enrichments[0][3]
fact4_p = self._li_fluct_enrichments[0][4]
fluct1_scale_p = tf.image.resize(fluct1_p, [28,28], method="bicubic", antialias=True)
fluct2_scale_p = tf.image.resize(fluct2_p, [28,28], method="bicubic", antialias=True)
fluct3_scale_p = tf.image.resize(fluct3_p, [28,28], method="bicubic", antialias=True)
fluct4_scale_p = fluct4_p
fluct_scale_p = fact0_p*fluct0_scale_p \
+ fact1_p*fluct1_scale_p + fact2_p*fluct2_scale_p \
+ fact3_p*fluct3_scale_p + fact4_p*fluct4_scale_p
else:
fluct_scale_p = fluct0_scale_p
# get the img-data
fluct_data_p = tf.image.per_image_standardization(fluct_scale_p)
fluct_data_p = tf.where(fluct_data_p > 5.e-6, fluct_data_p, tf.zeros_like(fluct_data_p))
self._initial_inp_img_data = fluct_data_p
self._inp_img_data = fluct_data_p
# Strategy 0: Simple superposition of the precursor image AND additional patterns at 4 different wave-length
# ~~~~~~~~~~
if self._oip_strategy == 0:
dim_1_1 = self._li_dim_steps[0][0]
dim_1_2 = self._li_dim_steps[0][1]
dim_2_1 = self._li_dim_steps[1][0]
dim_2_2 = self._li_dim_steps[1][1]
dim_3_1 = self._li_dim_steps[2][0]
dim_3_2 = self._li_dim_steps[2][1]
dim_4_1 = self._li_dim_steps[3][0]
dim_4_2 = self._li_dim_steps[3][1]
fact0 = self._ay_facts[0]
fact1 = self._ay_facts[1]
fact2 = self._ay_facts[2]
fact3 = self._ay_facts[3]
fact4 = self._ay_facts[4]
# print some parameter information
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
if b_print:
print("\nInitial image composition - strategy 0:\n Superposition of 4 different wavelength patterns")
print("Parameters:\n",
fact0, " => precursor image \n",
fact1, " => (" + str(dim_1_1) +", " + str(dim_1_2) + ") :: ",
fact2, " => (" + str(dim_2_1) +", " + str(dim_2_2) + ") :: ",
fact3, " => (" + str(dim_3_1) +", " + str(dim_3_2) + ") :: ",
fact4, " => (" + str(dim_4_1) +", " + str(dim_4_2) + ")"
)
# fluctuations
fluct1 = 2.0 * ( np.random.random((1, dim_1_1, dim_1_2, 1)) - 0.5 )
fluct2 = 2.0 * ( np.random.random((1, dim_2_1, dim_2_2, 1)) - 0.5 )
fluct3 = 2.0 * ( np.random.random((1, dim_3_1, dim_3_2, 1)) - 0.5 )
fluct4 = 2.0 * ( np.random.random((1, dim_4_1, dim_4_2, 1)) - 0.5 )
# Scaling with bicubic interpolation to the required image size
fluct1_scale = tf.image.resize(fluct1, [28,28], method="bicubic", antialias=True)
fluct2_scale = tf.image.resize(fluct2, [28,28], method="bicubic", antialias=True)
fluct3_scale = tf.image.resize(fluct3, [28,28], method="bicubic", antialias=True)
fluct4_scale = fluct4
# superposition with the already calculated image
fluct_data = fact0 * self._initial_inp_img_data \
+ fact1*fluct1_scale + fact2*fluct2_scale \
+ fact3*fluct3_scale + fact4*fluct4_scale
# get the standardized plus smoothed and unsmoothed image
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
# TF2 provides a function performing standardization of image data function
fluct_data_unsmoothed = fluct_data
fluct_data_smoothed = tf.image.per_image_standardization(
tf.image.resize( fluct_data, [28,28],
method="bicubic", antialias=True) )
if b_smoothing:
self._initial_inp_img_data = fluct_data_smoothed
else:
self._initial_inp_img_data = fluct_data_unsmoothed
# There should be no difference
img_init_unsmoothed = fluct_data_unsmoothed[0,:,:,0].numpy()
img_init_smoothed = fluct_data_smoothed[0,:,:,0].numpy()
if b_display:
ax1_1.imshow(img_init_unsmoothed, cmap=plt.cm.get_cmap('viridis'))
ax1_2.imshow(img_init_smoothed, cmap=plt.cm.get_cmap('viridis'))
print("Initial images plotted")
return self._initial_inp_img_data
#
# Method to prepare a Precursor run which checks a variety of large scale fluctuations for optimum actvation
# ***********************************
def _prepare_precursor(self, map_index = 120, b_print = False):
'''
@summary: A method to prepare a Precursor run by setting up GradientTape and the _ierate() function for an optimization loop
@version: 0.2, 30.09.2020
@changes: Minor - adaption to _setup_gradient_tape_and_iterate_function() instead of the obsolete _setup_gradient_tape()
@requires: A loaded CNN-Model and an already built OIP-model
@requires: A valid map-index as input parameter
@note: This method sets up the GradientTape and _iterate only once
- it will not be done again during the investigation of thousands of different input images (with large scale fluctuations)
which we investigate during the _precursor()-run.
@param map_index: Index selecting a map for the CNN layer defined previously by _build_oip_model()
@param b_print: Boolean - decides about intemediate output
'''
# Get input parameters
# ~~~~~~~~~~~~~~~~~~~~~~
self._map_index = map_index
# Rest some variables
self._val_oip_grads = None
self._val_oip_loss = None
self._iterate = None
# Setup the TF2 GradientTape watch
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
self._setup_gradient_tape_and_iterate_function(b_print = b_print)
if b_print:
print("GradientTape watch activated and _iterate()-function defined")
# Method to prepare a Precursor run which checks a variety of large scale fluctuations for optimum actvation
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
def _precursor(self,
li_pre_val=[0.2, 0.5, 0.8],
num_epochs=10,
loss_limit = 0.5,
b_print = True,
b_with_enriched_fluct = False,
li_facts = (1.0, 0.0, 0.0, 0.0, 0.1),
li_dim_steps = ( (3,3), (7,7), (14,14), (28,28) ),
b_check = True,
fig_a = None,
li_axa = None,
# the next parameters are new in V0.6
b_show_test_input_images = False,
fig_test_img = None,
ax_b = None,
interval_test_img = 1):
'''
@summary: Method to investigate thousands of input images with large scale fluctuations for the reaction of a specified map (filters)
and a given number of epochs in pattern creation
@version: 0.6, 12.12.2020
@changes: V0.5: Minor - documentation, skipped some superfluous statements
@changes: V0.6: Added intermediate plot output of constructed input images (a fig_a-reference must have been provided!)
@note: We select the 8 most dominant images - or less, if there are fewer input images which trigger the map
@requires: Previous run of _prepare_cursor() with a definition of the map-index
@note: We vary only 3 given pixel values on (3x3) grids (19683 possibilities)
@note: The optimization loop is completely done within this method - due to performance reasons
@param li_pre_val: A list of three scaled pixel values between ]0, 1[ which shall be combined in (3x3)-fluctuation patterns
@param num_epochs: The number of epochs used in the optimization loop for pattern creation
@note: It is worthwile to experiment with the number of epochs - the (3x3)-pattern selection may change !!!
@param loss_limit: Threshold of loss for which we register a fluctuation image as relevant
@param b_print: Boolean - controls printout for intermediate results - useful to see map response
@param b_check: Check the response of map for the first saved pattern - check the image reconstruction at the same time
@note: Parameters to enrich the (3x3)-large scale fluctuation with a small scale pattern
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
@param b_with_enriched_fluct: Boolean - controls whether we enrich the long-range pattern with other additional patterns
@param li_facts: A list of factors which the control the relative strength of the precursor image vs.
4 additional fluctuation patterns
@param li_dim_steps: A list of square dimensions for setting the length scale of the fluctuations
@note: Parameters for plotting
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
@param fig_a: reference to the 8 axes-frame
@param li_axa: A Python list of references to external (Jupyter-) axes-frames for plotting
Plots for intermediate test images
Note that for inernediate image plotting the cell must have a command line with metacommand
%matplotlib notebook
plt.ion()
@param b_show_test_input_images = Boolean; show intermediate
fluctuation patterns
@param fig_test_img: reference to external image plot frame.
@param ax_b: Rference to axis-frame for the intermediate display of test-images
@note: Preparations for plotting:
---------------------------------
fig_b : a simple figure for an image
fig_a: We need 8 axis-frames which must be provided externally
With Jupyter this can externally be done by statements like
# figure
# -----------
fig_b_pre, ax_b_pre = plt.subplots(1, figsize=(3,3))
fig_b_pre.canvas.draw()
fig_a_pre = plt.figure(2, figsize=(10,5))
axa_1 = fig_a_pre.add_subplot(241)
axa_2 = fig_a_pre.add_subplot(242)
axa_3 = fig_a_pre.add_subplot(243)
axa_4 = fig_a_pre.add_subplot(244)
axa_5 = fig_a_pre.add_subplot(245)
axa_6 = fig_a_pre.add_subplot(246)
axa_7 = fig_a_pre.add_subplot(247)
axa_8 = fig_a_pre.add_subplot(248)
li_axa = [axa_1, axa_2, axa_3, axa_4, aab_5, axa_6, axa_7, axa_8]
fig_a_pre.canvas.draw()
'''
# Internal parameter for number of selected input patterns
num_selected = 8
# check if length of li_axa is sufficient
if li_axa == None or len(li_axa) < num_selected:
print("Error: The length of the provided list with axes-frames for plotting must be at least ", num_selected )
sys.exit()
# get required exernal params
# ~~~~~~~~~~~~~~~~~~~~~~~~
self._n_steps = 2 # only a dummy
self._epsilon = 0.01 # only a dummy
# number of epochs for optimization strategy 0
if num_epochs == None:
len_epochs = len(self._ay_epochs)
self._n_epochs = self._ay_epochs[len_epochs-1]
else:
self._n_epochs = num_epochs
# Create a fixed random flutution pattern which we later can overlay to the long range fluctuation patterns
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
self._li_dim_steps = li_dim_steps # dimensions for fluctuations
self._ay_facts = np.array(li_facts)
dim_1_1 = self._li_dim_steps[0][0]
dim_1_2 = self._li_dim_steps[0][1]
dim_2_1 = self._li_dim_steps[1][0]
dim_2_2 = self._li_dim_steps[1][1]
dim_3_1 = self._li_dim_steps[2][0]
dim_3_2 = self._li_dim_steps[2][1]
dim_4_1 = self._li_dim_steps[3][0]
dim_4_2 = self._li_dim_steps[3][1]
fact0 = self._ay_facts[0]
fact1 = self._ay_facts[1]
fact2 = self._ay_facts[2]
fact3 = self._ay_facts[3]
fact4 = self._ay_facts[4]
# Create fluctuation patterns for enrichment
fluct1 = 2.0 * ( np.random.random((1, dim_1_1, dim_1_2, 1)) - 0.5 )
fluct2 = 2.0 * ( np.random.random((1, dim_2_1, dim_2_2, 1)) - 0.5 )
fluct3 = 2.0 * ( np.random.random((1, dim_3_1, dim_3_2, 1)) - 0.5 )
fluct4 = 2.0 * ( np.random.random((1, dim_4_1, dim_4_2, 1)) - 0.5 )
li_fluct = [fluct1, fluct2, fluct3, fluct4]
# Scaling with bicubic interpolation to the required image size
fluct1_scale = tf.image.resize(fluct1, [28,28], method="bicubic", antialias=True)
fluct2_scale = tf.image.resize(fluct2, [28,28], method="bicubic", antialias=True)
fluct3_scale = tf.image.resize(fluct3, [28,28], method="bicubic", antialias=True)
fluct4_scale = fluct4
# Create cartesian product of combinatoric possibilities for a (3x3)-grid of long range fluctuations
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
cp = list(product(li_pre_val, repeat=9))
num = len(cp)
print ("We test ", num, " possibilities for a (3x3) fluctuations ")
# Prepare lists to save parameter data for the fluctuation pattern
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
# Intermediate list to save relevant long scale fluctuations
d_cov = {}
# list to save parameters for an enrichments of the large scale pattern with small fluctuations
if b_with_enriched_fluct:
self._li_fluct_enrichments = [li_facts, li_dim_steps, li_fluct]
else:
self._li_fluct_enrichments = []
# Loop to check for relevant fluctuations => Loop over all combinations
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
for i_cp in range(num):
# create the value distribution
cov = np.array(cp[i_cp])
cov = cov.reshape(1,3,3,1)
# create basic image to investigate
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
fluct_scale = tf.image.resize(cov, [28,28], method="bicubic", antialias=True)
# enrich with additional smallscale fluctuations
if b_with_enriched_fluct:
# superposition with the already calculated image
fluct_scale = fact0 * fluct_scale \
+ fact1*fluct1_scale + fact2*fluct2_scale \
+ fact3*fluct3_scale + fact4*fluct4_scale
#standardization of the image data
fluct_data = tf.image.per_image_standardization(fluct_scale)
# eliminatng very small values - prove to be helpful in many cases
fluct_data = tf.where(fluct_data > 5.e-6, fluct_data, tf.zeros_like(fluct_data))
# save image data
self._initial_inp_img_data = fluct_data
self._inp_img_data = fluct_data
# Display the present input image
if b_show_test_input_images and i_cp%interval_test_img == 0:
img_b = fluct_data[0, :,:, 0].numpy()
ax_b.imshow(img_b, cmap=plt.cm.get_cmap('viridis'))
fig_test_img.canvas.draw()
# optimization loop
# ~~~~~~~~~~~~~~~~~
for j in range(self._n_epochs):
# Get output values of our Keras iteration function
self._val_oip_loss, self._val_oip_grads = self._iterate([self._inp_img_data])
# gradient ascent => Correction of the input image data
self._inp_img_data += self._val_oip_grads * self._epsilon
# Standardize the corrected image - we won't get a convergence otherwise
self._inp_img_data = tf.image.per_image_standardization(self._inp_img_data)
# Check if we have a loss value > 0.x and save the (3x3)-data
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
if self._val_oip_loss > loss_limit:
d_cov[i_cp] = [self._val_oip_loss, cov]
if b_print:
tf.print("i = ", i_cp, " loss = ", self._val_oip_loss)
# We restrict the number to those 8 with highest loss
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
if len(d_cov) > 0:
self._li_of_flucts = sorted(d_cov.items() , reverse = True, key=lambda x: x[1][0])
# print("num of relevant covs = ", len(self._li_of_flucts), len(d_cov))
print("\nnum of relevant covs = ", len(self._li_of_flucts))
self._li_of_flucts = self._li_of_flucts[:num_selected].copy()
#save( 'li_of_flucts.npy', np.array(self._li_of_flucts, dtype=np.float32) )
save( 'li_of_flucts.npy', self._li_of_flucts)
# save the enrichment-setting
save('li_of_cov_enrichments.npy',self._li_fluct_enrichments)
# Check of map really reacted - and check the reconstruction mechanism
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~
if b_check:
print("check of map reaction to first selected image")
cov_t = self._li_of_flucts[0][1][1]
#print("cov_t-shape = ", cov_t.shape)
#cov_del = cov_t - cov
#print("cov_del =\n", cov_del)
fluct0_scale_t = tf.image.resize(cov_t, [28,28], method="bicubic", antialias=True)
# Add fluctuation enrichments - if saved
if b_with_enriched_fluct:
# Scaling enrichment flucts with bicubic interpolation to the required image size
fluct1_t = self._li_fluct_enrichments[2][0]
fluct2_t = self._li_fluct_enrichments[2][1]
fluct3_t = self._li_fluct_enrichments[2][2]
fluct4_t = self._li_fluct_enrichments[2][3]
fact0_t = self._li_fluct_enrichments[0][0]
fact1_t = self._li_fluct_enrichments[0][1]
fact2_t = self._li_fluct_enrichments[0][2]
fact3_t = self._li_fluct_enrichments[0][3]
fact4_t = self._li_fluct_enrichments[0][4]
fluct1_scale_t = tf.image.resize(fluct1_t, [28,28], method="bicubic", antialias=True)
fluct2_scale_t = tf.image.resize(fluct2_t, [28,28], method="bicubic", antialias=True)
fluct3_scale_t = tf.image.resize(fluct3_t, [28,28], method="bicubic", antialias=True)
fluct4_scale_t = fluct4_t
fluct_scale_t = fact0_t*fluct0_scale_t \
+ fact1_t*fluct1_scale_t + fact2_t*fluct2_scale_t \
+ fact3_t*fluct3_scale_t + fact4_t*fluct4_scale_t
else:
fluct_scale_t = fluct0_scale_t
#standardization
fluct_data_t = tf.image.per_image_standardization(fluct_scale_t)
fluct_data_t = tf.where(fluct_data_t > 5.e-6, fluct_data_t, tf.zeros_like(fluct_data_t))
self._initial_inp_img_data = fluct_data_t
self._inp_img_data = fluct_data_t
self._precursor_img = fluct_data_t
# optimization loop
for j in range(self._n_epochs):
self._val_oip_loss, self._val_oip_grads = self._iterate([self._inp_img_data])
self._inp_img_data += self._val_oip_grads * self._epsilon
self._inp_img_data = tf.image.per_image_standardization(self._inp_img_data)
print("loss for 1st selected img = ", self._val_oip_loss )
# show the imgs
# ~~~~~~~~~~~~~~
self._display_precursor_imgs(li_axa = li_axa, fig_a = fig_a)
# list contains no patterns
else:
print("No image found !")
# Method to display initial fluctuation images identified as objects for OIP creation
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
def _display_precursor_imgs(self, li_axa = None, fig_a = None):
'''
@summary: Method to display up to 8 selected precursor images
@version: 0.2, 02.10.2020
@change: Only some documentation
@note: We first
reconstruct the image from saved data of the large scale fluctuations
@note: We then display the images in externally delivered axes-frames of matplotlib
@requires: A filled set of valid fluctuation patterns in self._li_of_flucts[][][] - determined by a run of _precursor()
@requires: Information on fluctuation enrichments in self._li_fluct_enrichments[][] - determined by a _precursor() run
@requires: A set of axes-frames for plotting - preferably defined in a Jupyter cell calling thi smethod
@note: Parameters for plotting
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
@param _li_axa: A Python list of references to external (Jupyter-) axes-frames for plotting
'''
# length of _li_of_flucts[] vs. length of li_axa
len_cov = len(self._li_of_flucts)
if li_axa == None or len(li_axa) < len_cov:
print("Error: The length of the provided list with axes-frames for plotting must be at least ", len_cov )
sys.exit()
# Loop to reconstruct and display the found precursor images
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
for j in range(len(self._li_of_flucts)):
print(j, "loss = ", self._li_of_flucts[j][1][0])
# reconstruct the image from the data of the precursor run
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
cov = self._li_of_flucts[j][1][1]
fluct0_scale_t = tf.image.resize(cov, [28,28], method="bicubic", antialias=True)
# add enrichments if defined
if len(self._li_fluct_enrichments) > 0:
# Scaling enrichment flucts with bicubic interpolation to the required image size
fluct1_t = self._li_fluct_enrichments[2][0]
fluct2_t = self._li_fluct_enrichments[2][1]
fluct3_t = self._li_fluct_enrichments[2][2]
fluct4_t = self._li_fluct_enrichments[2][3]
fact0_t = self._li_fluct_enrichments[0][0]
fact1_t = self._li_fluct_enrichments[0][1]
fact2_t = self._li_fluct_enrichments[0][2]
fact3_t = self._li_fluct_enrichments[0][3]
fact4_t = self._li_fluct_enrichments[0][4]
fluct1_scale_t = tf.image.resize(fluct1_t, [28,28], method="bicubic", antialias=True)
fluct2_scale_t = tf.image.resize(fluct2_t, [28,28], method="bicubic", antialias=True)
fluct3_scale_t = tf.image.resize(fluct3_t, [28,28], method="bicubic", antialias=True)
fluct4_scale_t = fluct4_t
fluct_scale_t = fact0_t*fluct0_scale_t \
+ fact1_t*fluct1_scale_t + fact2_t*fluct2_scale_t \
+ fact3_t*fluct3_scale_t + fact4_t*fluct4_scale_t
else:
fluct_scale_t = fluct0_scale_t
fluct_datx = tf.image.per_image_standardization(fluct_scale_t)
fluct_dat = tf.where(fluct_datx > 5.e-6, fluct_datx, tf.zeros_like(fluct_datx))
img = fluct_dat[0, :,:, 0].numpy()
li_axa[j].imshow(img, cmap=plt.cm.get_cmap('viridis'))
fig_a.canvas.draw()
return
# Method to transform an img tensor into a standard image - used for contrast enhancemant
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
def _transform_tensor_to_img(self, T_img = None, centre_move = 0.33, fact = 1.0):
'''
@summary: Method to transform (standardized) tensor img data into standard img array data
and to apply contrast enhancement(> V0.2)
@change: V0.2: Additional
parameers to control contrast enhancement.
We scale the the value distribution, move its center and apply some clipping
@note: Clipping is used to remove pixel values outside [0, 255]
@version: 0.2, 12.10.2020
@requires: A defined TF2/Keras backend
@param T_img: The TF2 or keras tensor for image data
@param centre_move: A decimal to move the centre of the pixel value distribution
'''
ay_x = T_img.numpy() # floating point array
maxi_o = np.max(T_img)
avg_o = np.mean(T_img)
mini_o = np.min(T_img)
std_dev_o = np.std(T_img)
print("\nInfos on pixel value distribution during contrast enhancement: ")
print("\max_orig = ", maxi_o, " :: avg_orig = ", avg_o, " :: min_orig: ", mini_o)
print("std_dev_orig = ", std_dev_o)
# the following operation should have no effect on standardized images
ay_x -= ay_x.mean()
ay_x /= (ay_x.std() + B.epsilon())
maxi = np.max(ay_x)
avg = np.mean(ay_x)
mini = np.min(ay_x)
std_dev = np.std(ay_x)
print("max_ay = ", maxi, " :: avg_ay = ", avg, " :: min_ay: ", mini)
print("std_dev_ay = ", std_dev)
div = fact * 0.5 * ( abs(maxi_o) + abs(mini_o) )
print("div = ", div)
ay_x /= div # scaling
ay_x += centre_move # moving the data center - 0.5 would move of the distribution into [0,1]
# # smaller values emphasize light/dark contratss
maxi = np.max(ay_x)
avg = np.mean(ay_x)
mini = np.min(ay_x)
std_dev = np.std(ay_x)
print("max_fin = ", maxi, " :: avg_fin = ", avg, " :: min_fin: ", mini)
print("std_dev_fin = ", std_dev)
ay_x = np.clip(ay_x, 0, 1)
ay_x *= 255
ay_x_img = np.clip(ay_x, 0, 255).astype('uint8')
maxi = np.max(ay_x_img)
avg = np.mean(ay_x_img)
mini = np.min(ay_x_img)
std_dev = np.std(ay_x_img)
print("max_img = ", maxi, " :: avg_img = ", avg, " :: min_img: ", mini)
print("std_dev_img = ", std_dev, "\n")
return ay_x_img
Links
A simple CNN for the MNIST dataset – XI – Python code for filter visualization and OIP detection