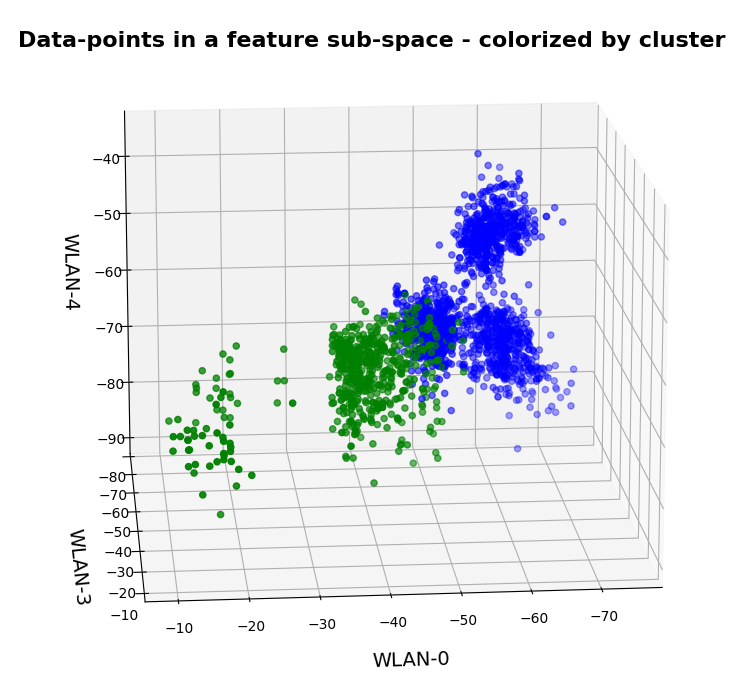
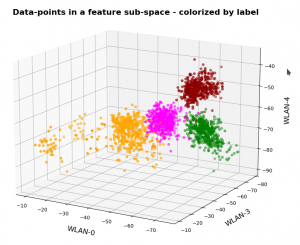
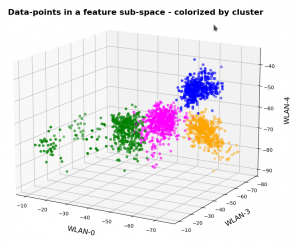
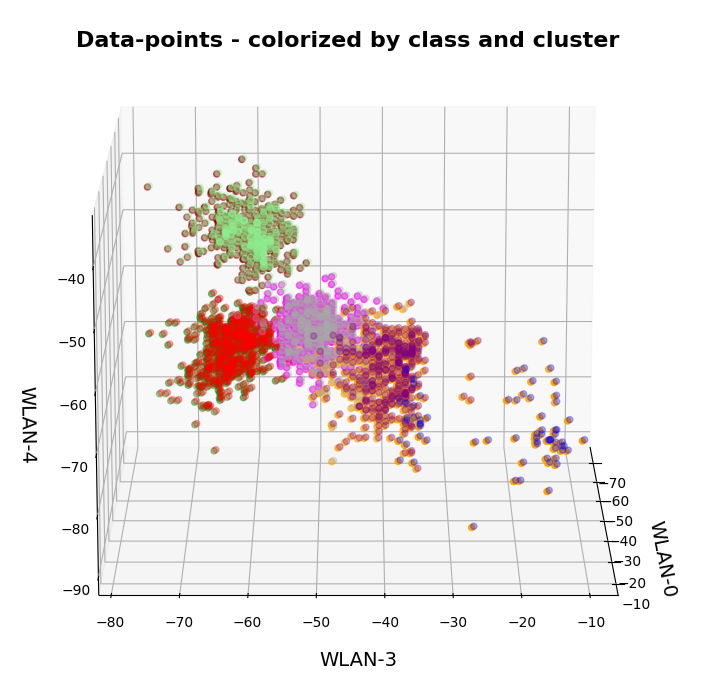
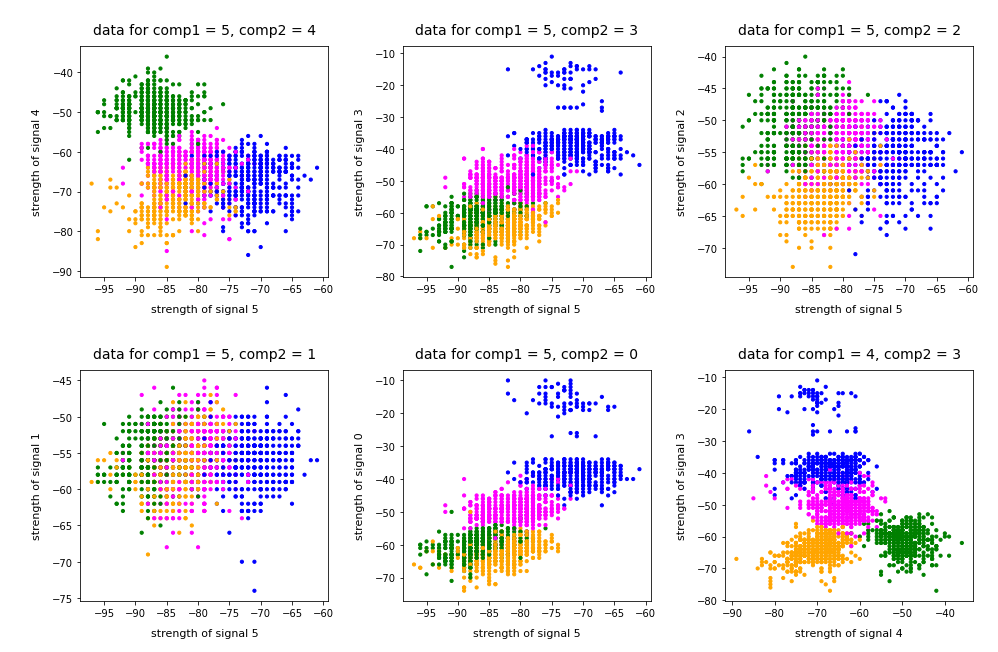
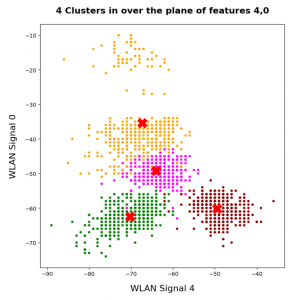
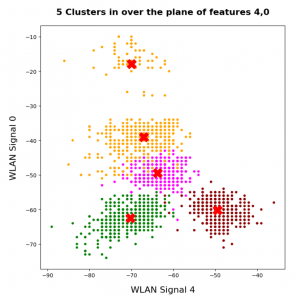
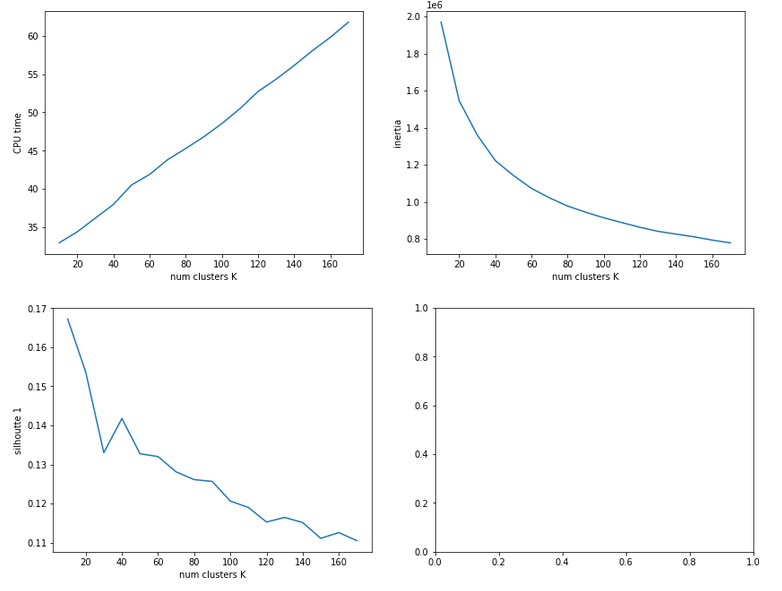
In this series about KMeans
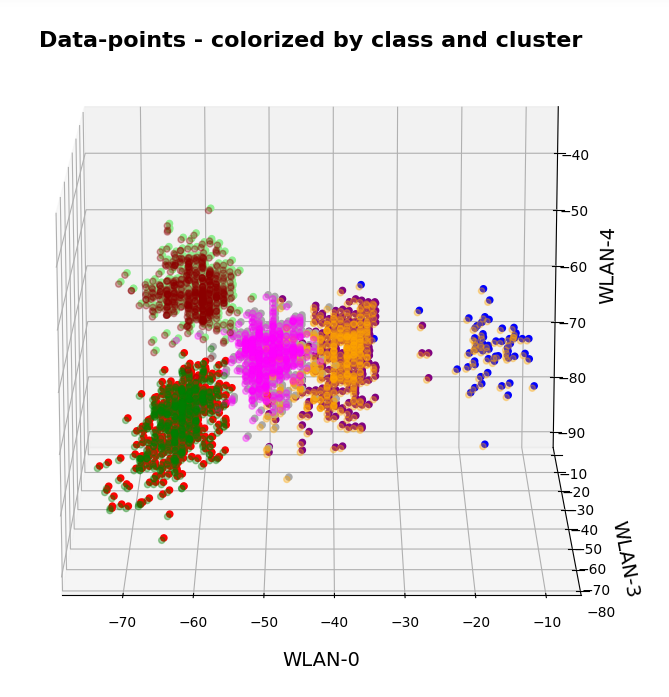
KMeans as a classifier for the WIFI and MNIST datasets – I – Cluster analysis of the WIFI example
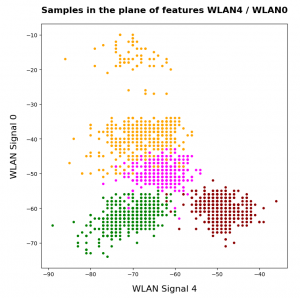
KMeans as a classifier for the WIFI and MNIST datasets – II – PCA in combination with KMeans for the WIFI-example
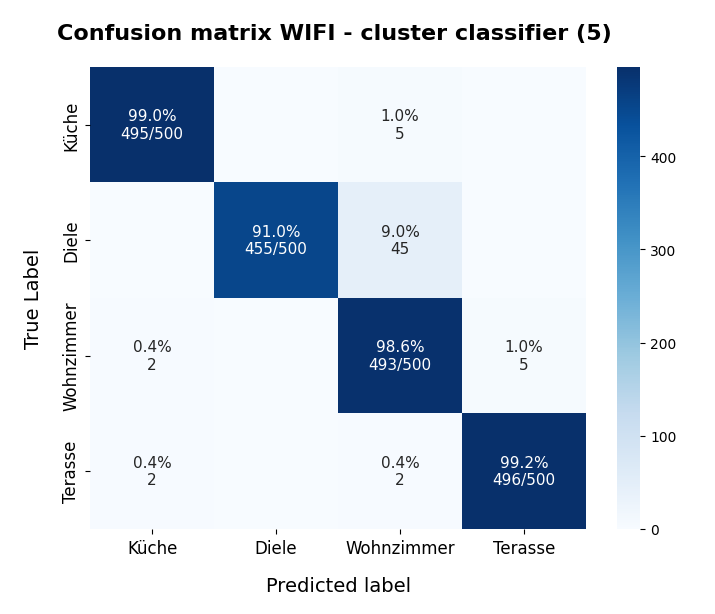
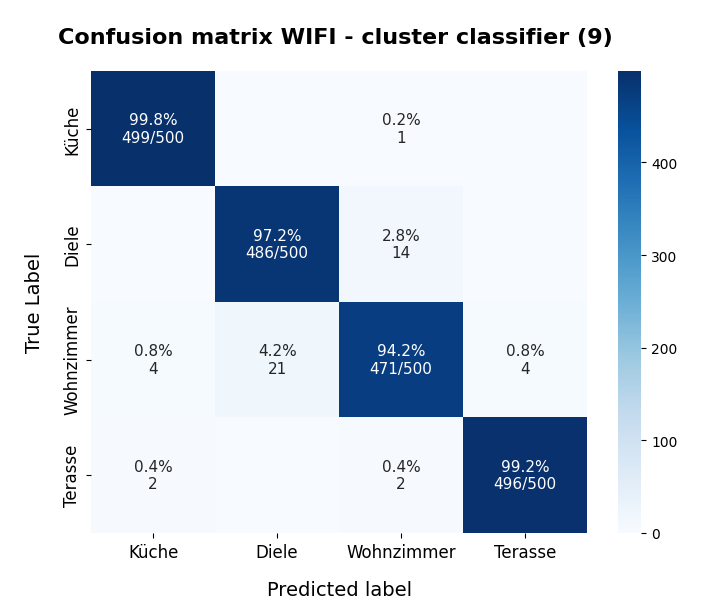
KMeans as a classifier for the WIFI and MNIST datasets – III – KMeans as a classifier for the WIFI-example
KMeans as a classifier for the WIFI and MNIST datasets – IV – KMeans on PCA transformed data
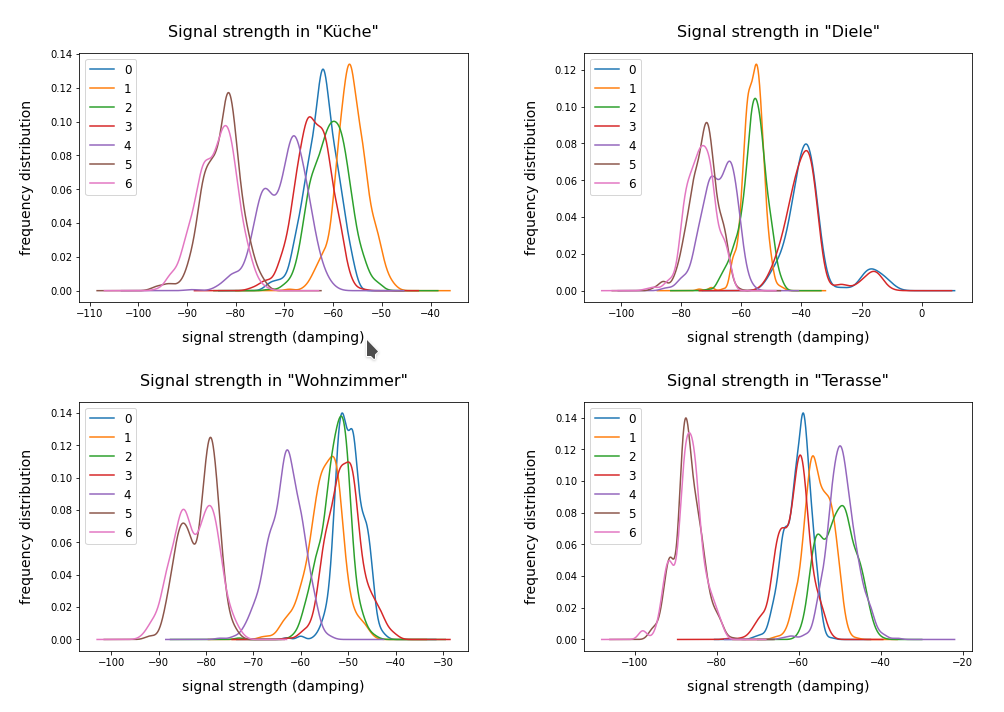
we have so far studied the application of KMeans to the WIFI dataset of the UCI Irvine. We now apply the Kmeans clustering algorithm to the MNIST dataset – also in an extended form, namely as a classifier. The MNIST dataset – a collection of 28x28px images of handwritten numbers – has already been discussed in other sections of this blog and is well documented on the Internet. I, therefore, do not describe its basic properties in this post. A typical image of the collection is
Load MNIST – dimensionality of the feature space and scaling of the data
Due to the ease of use, I loaded the MNIST data samples via TF2 and the included Keras interface. Otherwise TF2 was not used for the following experiments. Instead the clustering algorithms were taken from “sklearn”.
Each MNIST image can be transformed into a one-dimensional array with dimension 784 (= 28 * 28). This means the MNIST feature space has a dimension of 784 – which is much more than the seven dimensions we dealt with when analyzing the WIFI data in the last post. All MNIST samples were shuffled for individual runs.
Scaling of MNIST data for clustering?
A good question is whether we should scale or normalize the sample data for clustering – and if so by what formula. I could not answer this question directly; instead I tested multiple methods. Previous experience with PCA and MNIST indicated that Sklearn’s “Normalizer” would be helpful, but I did not take this as granted.
A simple scaling method is to just divide the pixel values by 255. This brings all 784 data array elements of each image into the value range [0,1]. Note that this scaling does not change relative length differences of the sample vectors in the feature space. Neither does it shift or change the width of the data distribution around its mean value. Other methods would be to standardize the data or to normalize them, e.g. by using respective algorithms from Scikit-Learn. Using either method in combination with a cluster analysis corresponds to a theory about the cluster distribution in the feature space. Normalization would mean that we assume that the clusters do not so much depend on the vector length in the feature space but mainly on the angle of the sample vectors. We shall later see what kind of scaling helps when we classify the MNIST data based on clusters.
In a first approach we leave the data as they are, i.e. unscaled.
Parameters for clustering
All the following cluster calculations were done on 3 out of 8 available (hyperthreaded) CPU cores. For Kmeans and MiniBatchKMeans we used
n_init = 100 # number of initial cluster configurations to test max_iter = 100 # maximum number of iterations tol = 1.e-4 # final deviation of subsequent results (= stop condition) random_state = 2 # a random state nmber for repeatable runs mb_size = 200 # size of minibatches (for MiniBatchKMeans)
The number of clusters “num_clus” was defined individually for each run.
Analysis by KMeans? Too expensive …
A naive approach to perform an elbow analysis, as we did for the WIFI-data, would be to apply KMeans of Sklearn directly to the MNIST data. But a test run on the CPU shows that such an endeavor would cost too much time. With 3 CPU cores and only a very limited number of clusters and iterations
n_init = 10 # only a few initial configurations max_iter = 50 tol = 1.e-3 num_clus = 25 # only a few clusters
a KMeans fit() run applied to 60,000 training samples [len(X_train) => 60,000]
kmeans.fit(X_train)
requires around 42 secs. For 200 clusters the cluster analysis requires around 214 secs. Doing an elbow analysis would therefore require many hours of computational time.
To overcome this problem I had to use MiniBatchKMeans. It is by factors > 80 faster.
Elbow analysis with the help of MiniBatchKmeans
When we use the following setting for MiniBatchKMeans
n_init = 50 # only a few initial configurations max_iter = 100 tol = 1.e-4 mb_size = 200
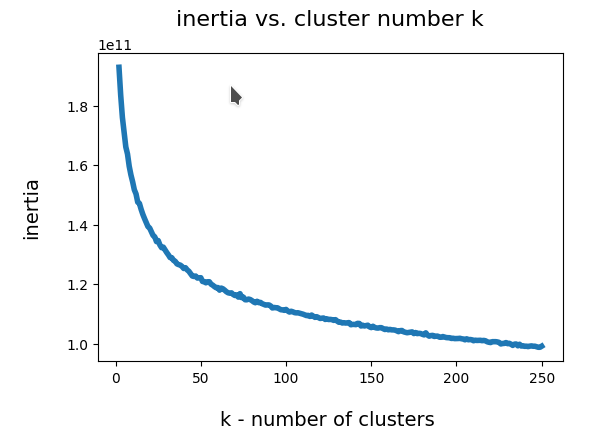
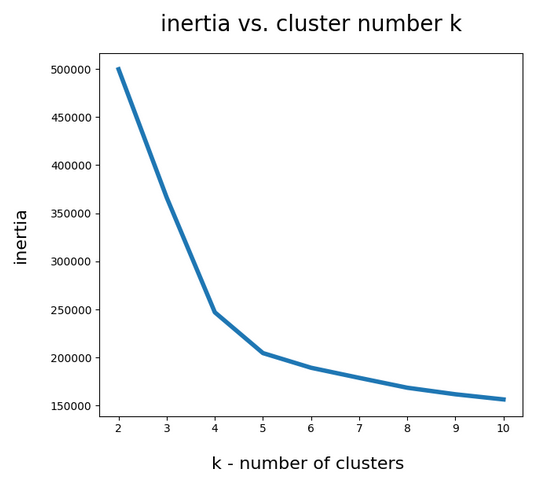
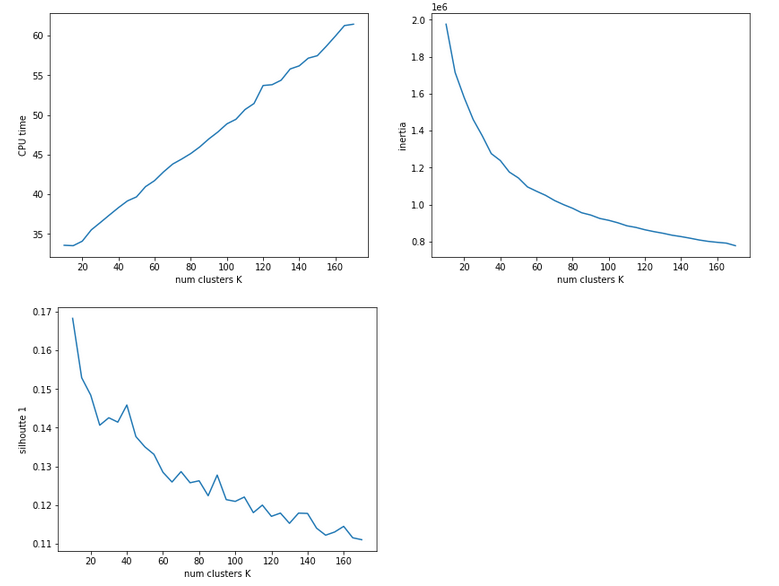
I could perform an elbow analysis for all cluster-numbers 1 < k <= 250 in less than 20 minutes. The following graphics shows the resulting intertia curve vs. cluster number:
The “elbow” is not very pronounced. But I would say that by using a cluster number around 200 we are on the safe side. By the way: The shape of the curve does not change very much when we apply Sklearn’s Normalizer to the MNIST data ahead of the cluster analysis.
Classifying unscaled data with the help of clusters
We now perform a prediction of our adapted cluster algorithm regarding the cluster membership for the training data and for k=225 clusters:
n_clu = 225 mb_size = 200 max_iter = 120 n_init = 100 tol = 1.e-4
Based on the resulting data we afterward apply the same type of algorithm which we used for the WIFI data to construct a “classifier” based on clusters and a respective predictor function (see the last post of this series).
The data distribution for the 10 different digits of the training set was:
class 0 : 5905 class 1 : 6721 class 2 : 6031 class 3 : 6082 class 4 : 5845 class 5 : 5412 class 6 : 5917 class 7 : 6266 class 8 : 5860 class 9 : 5961
How good is the cluster membership of a sample for a digit class defined?
Well, out of 225 clusters there were only around 15 for which I got an “error” above 40%, i.e. for which the relative fraction of data samples deviating from the dominant class of the cluster was above 40%. For the vast majority of clusters, however, samples of one specific digit class dominated the clusters members by more than 90%.
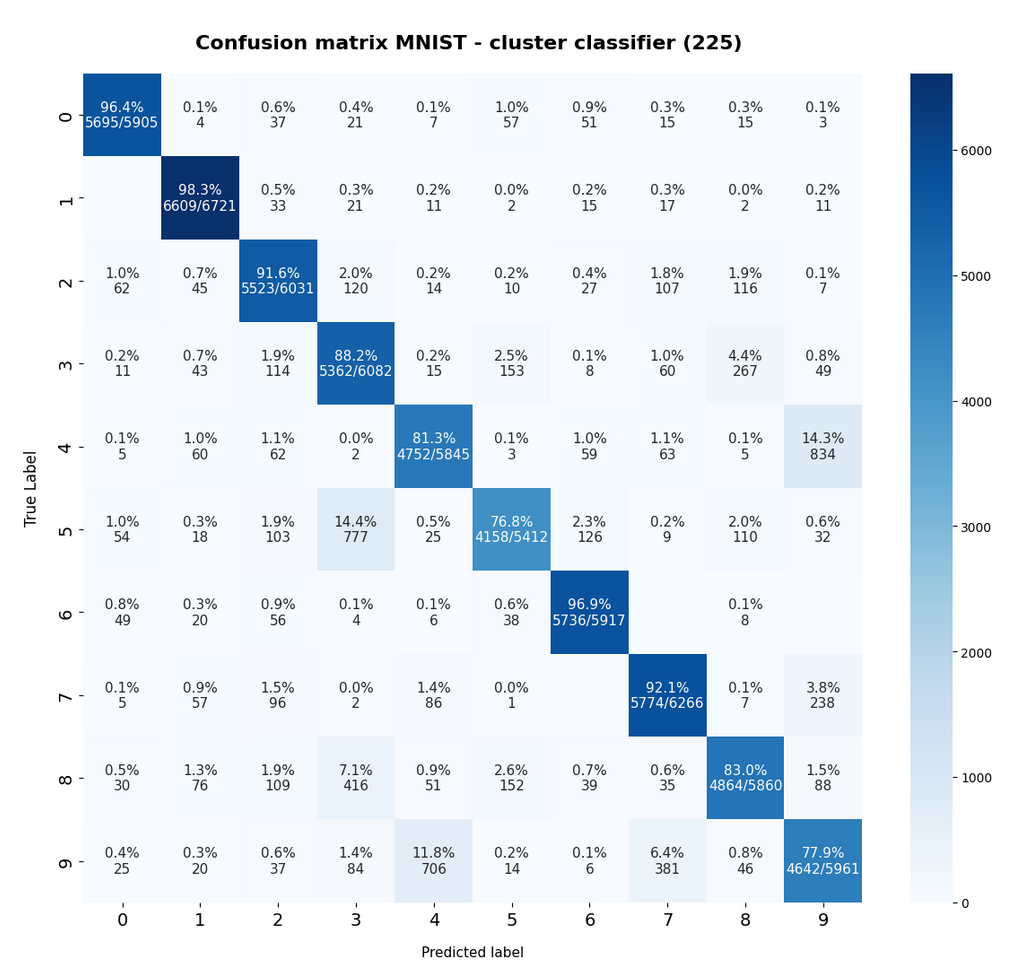
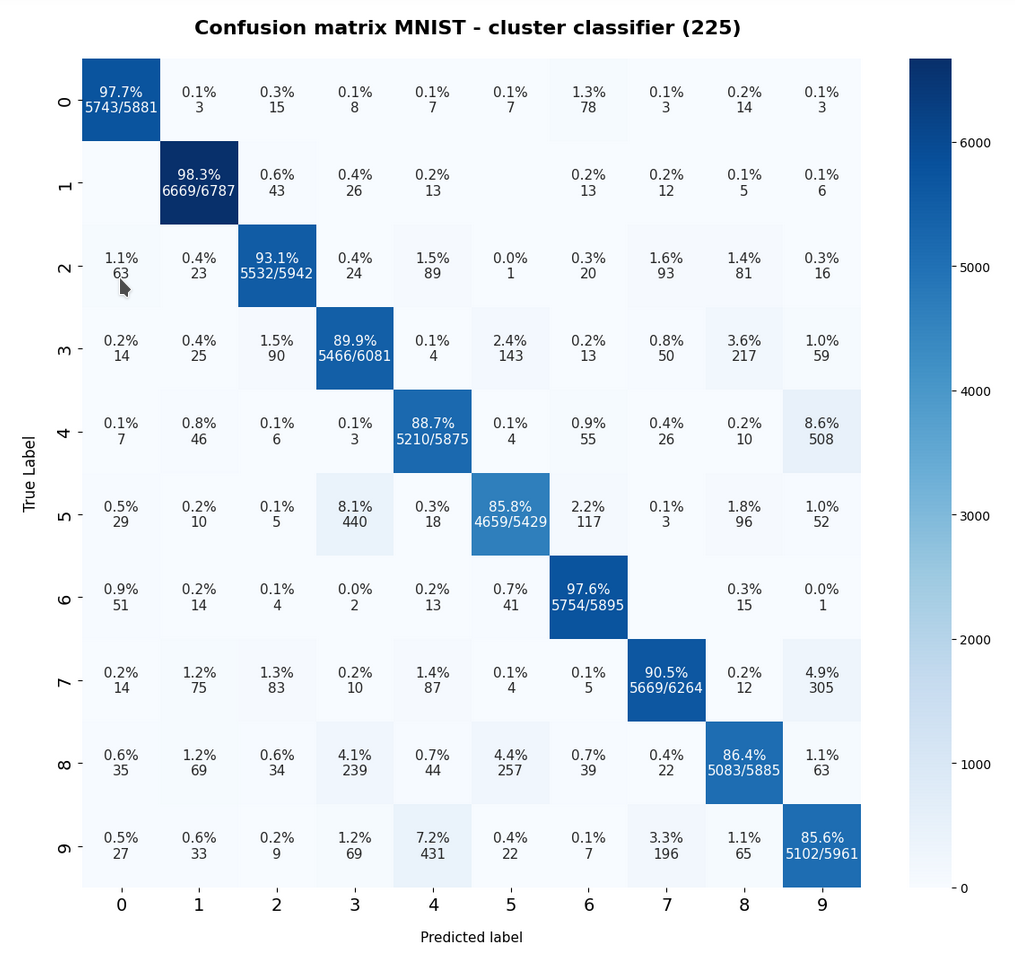
The resulting confusion matrix of our new “cluster classifier” for the (unscaled) MNIST data looks like
[[5695 4 37 21 7 57 51 15 15 3] [ 0 6609 33 21 11 2 15 17 2 11] [ 62 45 5523 120 14 10 27 107 116 7] [ 11 43 114 5362 15 153 8 60 267 49] [ 5 60 62 2 4752 3 59 63 5 834] [ 54 18 103 777 25 4158 126 9 110 32] [ 49 20 56 4 6 38 5736 0 8 0] [ 5 57 96 2 86 1 0 5774 7 238] [ 30 76 109 416 51 152 39 35 4864 88] [ 25 20 37 84 706 14 6 381 46 4642]]
This confusion matrix comes at no surprise: The digits “4”, “5”, “8”, “9” are somewhat error prone. Actually, everybody familiar with MNIST images knows that sometimes “4”s and “9”s can be mixed up even by the human eye. The same is true for handwritten “5”s, “8”s and “3”s.
Another representation of the confusion matrix is:
The calculation for the matrix elements was done in a standard way – the sum over percentages in a row gives 100% (the slight deviation in the matrix is due to rounding). I.e. we look at erors of the type TN (True Negatives).
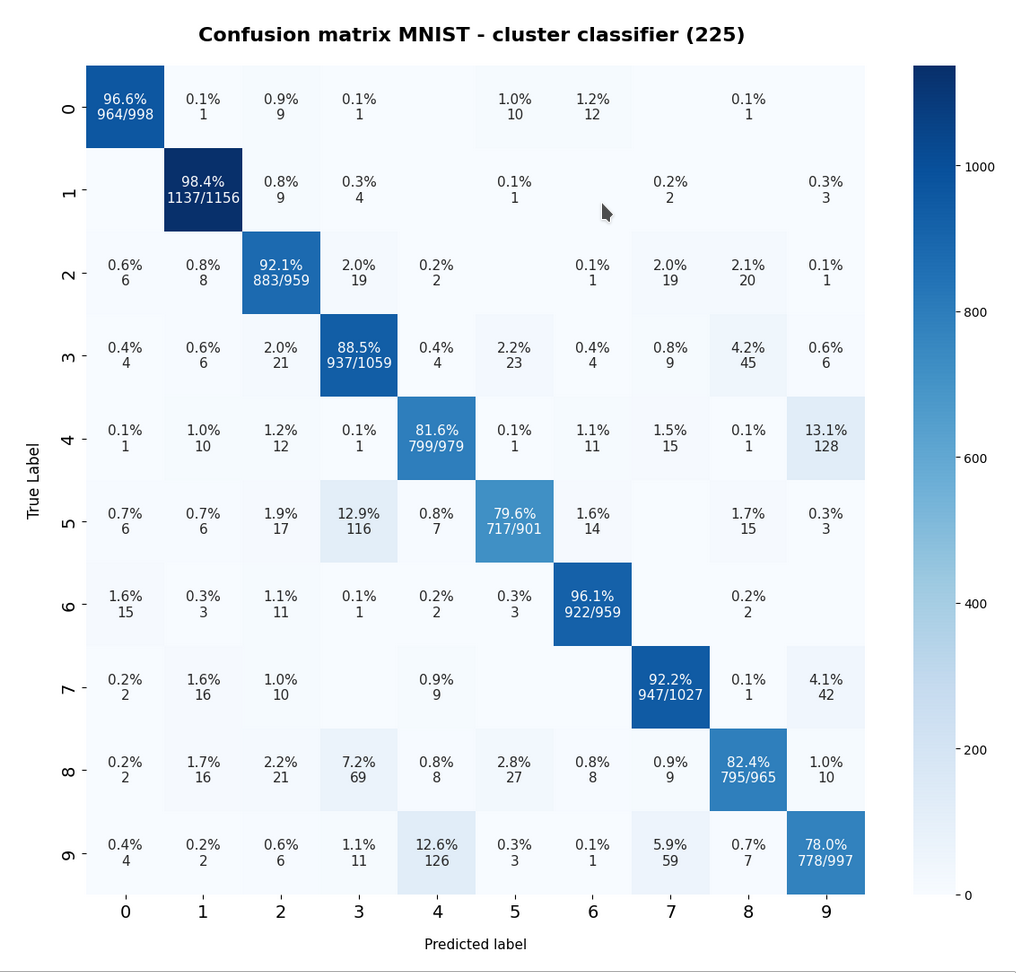
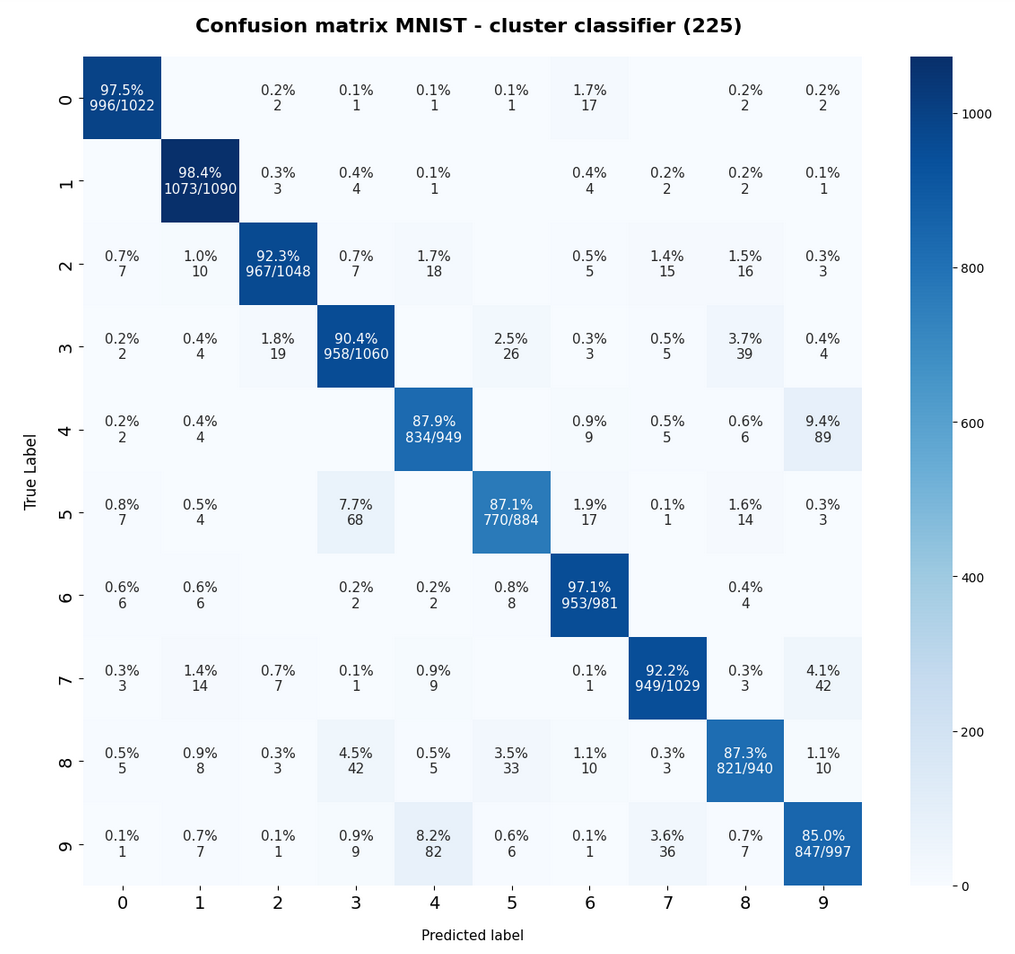
The confusion matrix for the remaining 10,000 test data samples is:
The relative errors we get for our classifier when applied to the train and test data is
rel_err_train = 0.115 ,
rel_err_test = 0.112
All for unscaled MNIST data. Taking into account the crudeness of the whole approach this is a rather convincing result. It proves that it is worth the effort to perform a cluster analysis on high dimensional data:
- It provides a first impression whether the data are structured in the feature space such that we can find relatively good separable clusters with dominant members belonging to just one class.
- It also shows that a cluster based classification for many datasets cannot reach accuracy levels of CNNs, but that it may still deliver good results. Without any supervised training …
The second point also proves that the distance of the data points to the various cluster centers contains valuable information about the class membership. So, a MLP or CNN based classification could be performed on transformed MNIST data, namely distance vectors of sample datapoints to the different cluster centers. This corresponds to a dimension reduction of the classification problem. Actually, in a different part of this blog, I have already shown that such an approach delivers accuracy values beyond 98%.
For MNIST we can say that the samples define a relatively well separable cluster structure in the feature space. The granularity required to resolve classes sufficiently well means a clsuter number of around 200 < k < 250. Then we get an accuracy close to 90% for cluster based classification.
t-SNE representation of the MNIST data
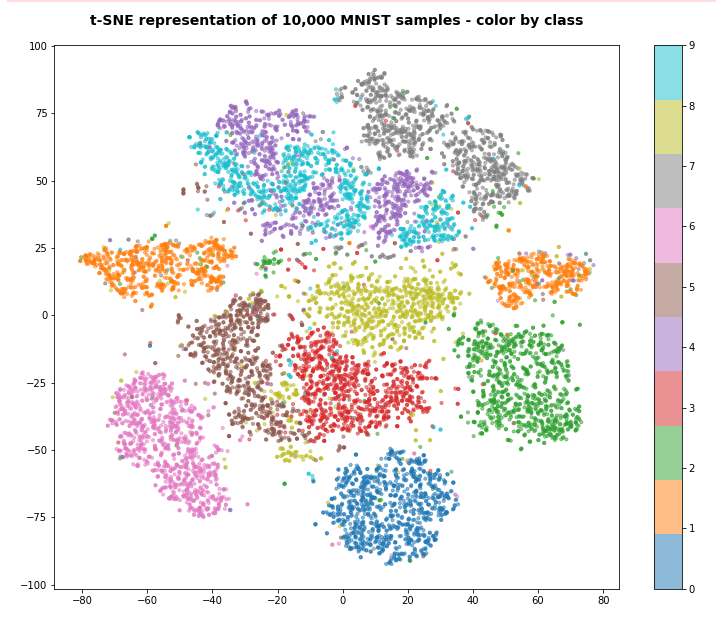
Can we somehow confirm this finding about a good cluster-class-relation independently? Well, in a limited way. The t-SNE algorithm, which can be used to “project” multidimensional data onto a 2-dimensional plane, respects the vicinity of vectors in the original feature space whilst deriving a 2-dim representation. So, a rather well structured t-SNE diagram is an indication of clustering in the feature space. And indeed for 10,000 randomly selected samples of the (shufffled) training data we get:
The colorization was done by classes, i.e. digits. We see a relatively good separation of major “clusters” with data points belonging to a specific class. But we also can identify multiple problem zones, where data points belonging to different classes are intermixed. This explains the confusion matrix. It also explains why we need so many fine-grained clusters to get a reasonable resolution regarding a reliable class-cluster-relation.
Classifying scaled and normalized MNIST data with the help of clusters
Can we improve the accuracy of our cluster based classification a bit? This would, e.g., require some transformation which leads to a better cluster separation. To see the effect of two different scalers I tried the “Normalizer” and then also the “StandardScaler” of Sklearn. Actually, they work in opposite direction regarding accuracy:
The “Normalizer” improves accuracy by more than 1.5%, while the “Standardizer” reduces it by almost the same amount.
I only discuss results for “Normalization” below. The confusion matrix for the training data becomes:
and for the test data:
The relative error for the test data is
Error for trainings data:
avg_err_train = 0.085 :: num_err_train = 5113
Error for test data:
avg_err_test = 0.083 :: num_err_test = 832
So, the relative accuracy is now around 91.5%.
The result depends a bit on the composition of the training and the test dataset after an initial shuffling. But the value remains consistently above 90%.
Data compression by Autoencoders and clustering
Just for interest I also had a look at a very different approach to invoke clustering:
I first applied a simple CNN-based AutoEncoder [AE] to compress the MNIST data into a 25-dimensional space and applied our clustering methods afterwards.
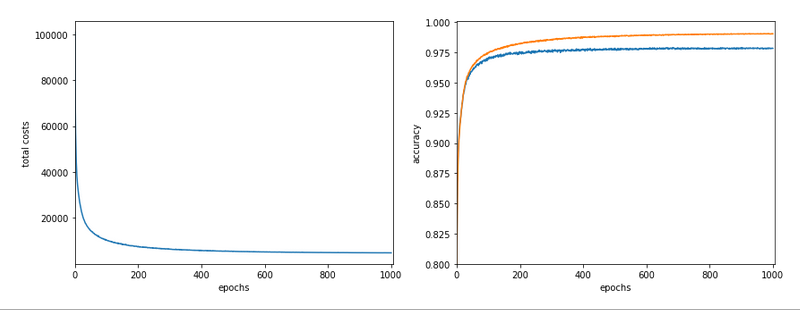
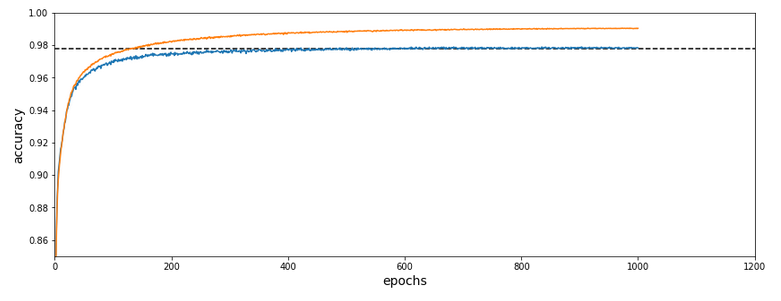
I shall not discuss the technology of autoenconders in this post. The only relevant point in our context is that an autoencoder provides an efficient non-linear way of data compression and dimensionality reduction. Among many other useful properties and abilities … . Note: I did not use a “Variational Autoencoder” which would have allowed for even better results. The loss function for the AE was a simple quadratic loss. The autoencoder was trained on 50,000 training samples and for 40 epochs.
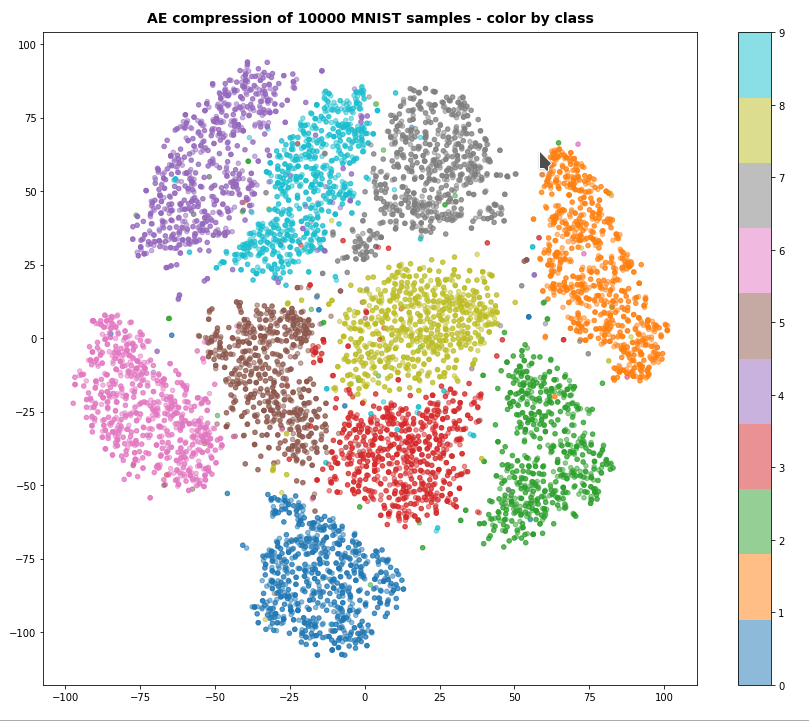
A t-SNE based plot of the “clusters” for test data in the 25-dimensional space looks like:
We see that the separation of the data belonging to different classes is somewhat better than before. Therefore, we expect a slightly better classification based on clusters, too. Without any scaling we get the following confusion data:
[[5817 7 10 3 1 14 15 2 27 1] [ 3 6726 29 2 0 1 10 5 12 10] [ 49 35 5704 35 14 4 10 61 87 7] [ 8 78 48 5580 22 148 2 40 111 29] [ 47 27 18 0 4967 0 44 38 3 673] [ 32 20 10 150 8 5039 73 4 43 28] [ 31 11 23 2 2 47 5746 0 15 1] [ 6 35 35 6 32 0 1 5977 7 163] [ 17 67 22 86 16 217 24 22 5365 52] [ 35 32 11 92 184 15 1 172 33 5406]] Error averaged over (all) clusters : 6.74
The resulting relative error for the test data was:
avg_err_test = 0.0574 :: num_err_test = 574
With Normalization:
Error for test data:
avg_err_test = 0.054 :: num_err_test = 832
So, after performing the autoencoder training on normalized data we consistently get
an accuracy of around 94%.
This is not too much of a gain. But remember:
We performed a cluster analysis on a feature space with only 25 dimensions – which of course is much cheaper. However, we paid a prize, namely the Autoencoder training which lasted about 150 secs on my old Nvidia 960 GTX.
And note: Even with only 100 clusters we get above 92% on the AE-compressed data.
Conclusion
We have shown that using a non-supervised cluster analysis of the MNIST data with around 225 clusters allows for classifying images with an accuracy around 90.5%. In combination with an Autoencoder compression we even reaches values around 94%. This is comparable with other non-optimized standard algorithms aside of neural networks.
This means that the MNIST data samples are organized in a well separable cluster structure in their feature space. A test run with normalized data showed that the clusters (and their centers) differ mostly by their direction relative to the origin of the feature space and not so much by their distance from the origin. With a relatively fine grained resolution we could establish a simple cluster-class-relation which allowed for cluster based classification.
The accuracy is, of course, below the values reachable with optimized MLPS (98%) and CNNs (above 99%). But, clustering is a fast, reliable and non-supervised method. In addition in combination with t-SNE we can create plots which can easily be understood by the customers. So, even for more complex data I would always recommend to try a cluster based classification approach if you need to provide plots and quick results. Sometimes the accuracy may even be sufficient for your customer’s purposes.