I continue with my series about a Python program to build simple MLPs:
A simple Python program for an ANN to cover the MNIST dataset – VI – the math behind the „error back-propagation“
A simple program for an ANN to cover the Mnist dataset – V – coding the loss function
A simple program for an ANN to cover the Mnist dataset – IV – the concept of a cost or loss function
A simple program for an ANN to cover the Mnist dataset – III – forward propagation
A simple program for an ANN to cover the Mnist dataset – II – initial random weight values
A simple program for an ANN to cover the Mnist dataset – I – a starting point
On our tour we have already learned a lot about multiple aspects of MLP usage. I name forward propagation, matrix operations, loss or cost functions. In the last article of this series
A simple program for an ANN to cover the Mnist dataset – VI – the math behind the „error back-propagation“
I tried to explain some of the math which governs “Error Back Propagation” [EBP]. See the PDF attached to the last article.
EBP is an algorithm which applies the “Gradient Descent” method for the optimization of the weights of a Multilayer Perceptron [MLP]. “Gradient Descent” itself is a method where we step-wise follow short tracks perpendicular to contour lines of a hyperplane in a multidimensional parameter space to hopefully approach a global minimum. A step means a change of of parameter values – in our context of weights. In our case the hyperplane is the surface formed by the cost function over the weights. If we have m weights we get a hyperplane in an (m+1) dimensional space.
To apply gradient descent we have to calculate partial derivatives of the cost function with respect to the weights. We have discussed this in detail in the last article. If you read the PDF you certainly have noted: Most of the time we shall execute matrix operations to provide the components of the weight gradient. Of course, we must guarantee that the matrices’ dimensions fit each other such that the required operations – as an element-wise multiplication and the numpy.dot(X,Y)-operation – become executable.
Unfortunately, there are some challenges regarding this point which we have not covered, yet. One objective of this article is to get prepared for these potential problems before we start coding EBP.
Another point worth discussing is: Is there really just one cost function when we use mini-batches in combination with gradient descent? Regarding the descriptions and the formulas in the PDF of the last article this was and is not fully clear. We only built sums there over cost contributions of all the records in a mini-batch. We did NOT use a loss function which assigned to costs to deviations of the predicted result (after forward propagation) from known values for all training data records.
This triggers the question what our
code in the end really does if and when it works with mini-batches during weight optimization … We start with this point.
In the following I try to keep the writing close to the quantity notations in the PDF. Sorry for a bad display of the δs in HTML.
Gradient descent and mini-batches – one or multiple cost functions?
Regarding the formulas given so far, we obviously handle costs and gradient descent batch-wise. I.e. each mini-batch has its own cost function – with fewer contributions than a cost function for all records would have. Each cost function has (hopefully) a defined position of a global minimum in the weights’ parameter space. Taking this into consideration the whole mini-batch approach is obviously based on some conceptually important assumptions:
- The basic idea is that the positions of the global minima of all the cost-functions for the different batches do not deviate too much from each other in the basic parameter space.
- If we additionally defined a cost function for all test data records (over all batches) then this cost function should display a global minimum positioned in between the ones of the batches’ cost functions.
- This also means that there should be enough records in each batch with a really statistical distribution and no specialties associated with them.
- Contour lines and gradients on the hyperplanes defined by the loss functions will differ from each other. On average over all mini-batches this should not hinder convergence into a common optimum.
To understand the last point let us assume that we have a batch for MNIST dataset where all records of handwritten digits show a tendency to be shifted to the left border of the basic 28×28 pixel frames. Then this batch would probably give us other weights than other batches.
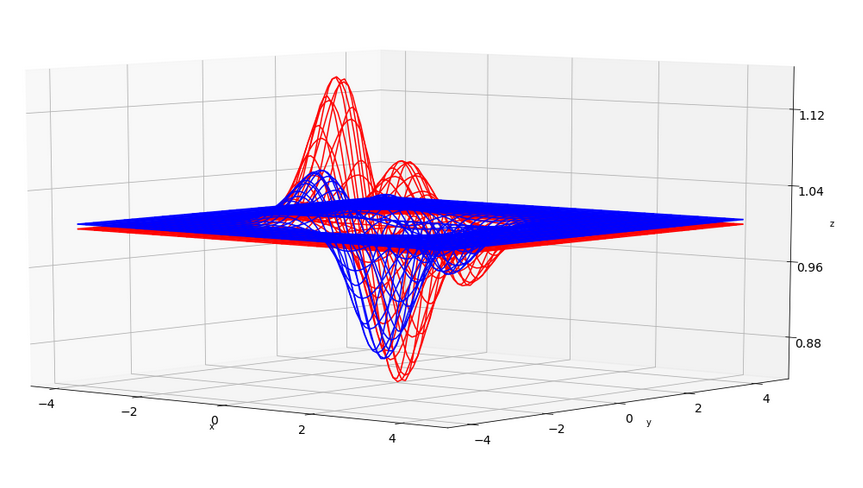
To get a deeper understanding, let us take only two batches. By chance their cost functions may deviate a bit. In the plots below I have just simulated this by two assumed “cost” functions – each forming a hyperplane in 3 dimensions over only two parameter (=weight) dimensions x and y. You see that the “global” minima of the blue and the red curve deviate a bit in their position.
The next graph shows the sum, i.e. the full “cost function”, in green in comparison to the (vertically shifted and scaled) original functions.
Also here you clearly see the differences in the minimas’ positions. What does this mean for gradient descent?
Firstly, the contour lines on the total cost function would deviate from the ones on the cost function hyperplanes of our 2 batches. So would the directions of the different gradients at the point presently reached in the parameter space during optimization! Working with batches therefore means jumping around on the surface of the total cost function a bit erratically and not precisely along the direction of steepest descent there. By the way: This behavior can be quite helpful to overcome local minima.
Secondly, in our simplified example we would in the end not converge completely, but jump or circle around the minimum of the total cost function. Reason: Each batch forces the weight corrections for x,y into different directions namely those of
its own minimum. So, a weight correction induced by one bath would be countered by corrections imposed by the optimization for the other batch. (Regarding MNIST it would e.g. be interesting to run a batch with handwritten digits of Europeans against a batch with digits written by Americans and see how the weights differ after gradient descent has converged for each batch.)
This makes us understand multiple things:
- Mini-batches should be built with a statistical distribution of records and their composition should be changed statistically from epoch to epoch.
- We need a criterion to stop iterating over too many epochs senselessly.
- We should investigate whether the number and thus the size of mini-batches influences the results of EBP.
- At the end of an optimization run we could invest in some more iterations not for the batches, but for the full cost function of all training records and see if we can get a little deeper into the minimum of this total cost function.
- We should analyze our batches – if we keep them up and do not create them statistically anew at the beginning of each epoch – for special data records whose properties are off of the normal – and maybe eliminate those data records.
Repetition: Why Back-propagation of 2 dimensional matrices and not vectors?
The step wise matrix operations of EBP are to be performed according to a scheme with the following structure:
- On a given layer N apply a layer specific matrix “NW.T” (depending on the weights there) by some operational rule on some matrix “(N+1)δS“, which contains some data already calculated for layer (N+1).
- Take the results and modify it properly by multiplying it element-wise with some other matrix ND (containing derivative expressions for the activation function) until you get a new NδS.
- Get partial derivatives of the cost function with respect to the weights on layer (N-1) by a further matrix operation of NδS on a matrix with output values (N-1)A.TS on layer (N-1).
- Proceed to the next layer in backward direction.
The input into this process is a matrix of error-dependent quantities, which are defined at the output layer. These values are then back-propagated in parallel to the inner layers of our MLP.
Now, why do we propagate data matrices and not just data vectors? Why are we allowed to combine so many different multiplications and summations described in the last article when we deal with partial derivatives with respect to variables deep inside the network?
The answer to the first question is numerical efficiency. We operate on all data records of a mini-batch in parallel; see the PDF. The answer to the second question is 2-fold:
- We are allowed to perform so many independent operations because of the linear structure of our cost-functions with respect to contributions coming from the records of a mini-batch and the fact that we just apply linear operations between layers during forward propagation. All contributions – however non-linear each may be in itself – are just summed up. And propagation itself between layers is defined to be linear.
- The only non-linearity occurring – namely in the form of non-linear activation functions – is to be applied just on layers. And there it works only node-wise! We do not
couple values for nodes on one and the same layer.
In this sense MLPs are very simple by definition – although they may look complex! (By the way and if you wonder why MLPs are nevertheless so powerful: One reason has to do with the “Universal Approximation Theorem”; see the literature hint at the end.)
Consequence of the simplicity: We can deal with δ-values (see the PDF) for both all nodes of a layer and all records of a mini-batch in parallel.
Results derived in the last article would change dramatically if we had rules that coupled the Z- or A-values of different nodes! E.g. if the squared value at node 7 in layer X must always be the sum of squared values at nodes 5 an 6. Believe me: There are real networks in this world where such a type of node coupling occurs – not only in physics.
Note: As we have explained in the PDF, the nodes of a layer define one dimension of the NδS“-matrices,
the number of mini-batch records the other. The latter remains constant. So, during the process the δ-matrices change only one of their 2 dimensions.
Some possible pitfalls to tackle before EBP-coding
Now, my friends, we can happily start coding … Nope, there are actually some minor pitfalls, which we have to explain first.
Special cost-, activation- and output-functions
I refer to the PDF mentioned above and its formulas. The example explained there referred to the “Log Loss” function, which we took as an example cost function. In this case the outδS and the 3δS-terms at the nodes of the outermost layer turned out to be quite simple. See formula (21), (22), (26) and (27) in the PDF.
However, there may be other cost functions for which the derivative with respect to the output vector “a” at the outermost nodes is more complicated.
In addition we may have other output or activation functions than the sigmoid function discussed in the PDF’s example. Further, the output function may differ from the activation function at inner layers. Thus, we find that the partial derivatives of these functions with respect to their variables “z” must be calculated explicitly and as needed for each layer during back propagation; i.e., we have to provide separate and specific functions for the provision of the required derivatives.
At the outermost layer we apply the general formulas (84) to (88) with matrix ED containing derivatives of the output-function Eφ(z) with respect to the input z to find EδS with E marking the outermost layer. Afterwards, however, we apply formula (92) – but this time with D-elements referring to derivatives of the standard activation-function φ used at nodes of inner layers.
The special case of the Log Loss function and other loss functions with critical denominators in their derivative
Formula (21) shows something interesting for the quantity outδS, which is a starting point for backward propagation: a denominator depending on critical factors, which directly involve output “a” at the outer nodes or “a” in a difference term. But in our one-hot-approach “a” may become zero or come close to it – during training by accident or by convergence! This is a dangerous thing; numerically we absolutely want to avoid any division by zero or by small numbers close to the numerical accuracy of a programming language.
What mathematically saves us in the special case of Log Loss are formulas (26) and (27), where due to some “magic” the dangerous denominator is cancelled by a corresponding factor in the numerator when we evaluate EδS.
In the general case, however,
we must investigate what numerical dangers the functional form of the derivative of the loss function may bring with it. In the end there are two things we should do:
- Build a function to directly calculate EδS and put as much mathematical knowledge about the involved functions and operations into it as possible, before employing an explicit calculation of values of the cost function’s derivative.
- Check the involved matrices, whose elements may appear in denominators, for elements which are either zero or close to it in the sense of the achievable accuracy.
For our program this means: Whether we calculate the derivative of a cost function to get values for “outδS” will depend on the mathematical nature of the cost function. In case of Log Loss we shall avoid it. In case of MSE we shall perform the numerical operation.
Handling of bias nodes
A further complication of our aspired coding has its origin in the existence of bias nodes on every inner layer of the MLP. A bias node of a layer adds an additional degree of freedom whilst adjusting the layer’s weights; a bias node has no input, it produces only a constant output – but is connected with weights to all normal nodes of the next layer.
Some readers who are not so familiar with “artificial neural networks” may ask: Why do we need bias nodes at all?
Well, think about a simple matrix operation on a 2 dim-vector; it changes its direction and length. But if we want to approximate a function for regression or a separation hyperplanes for classification by a linear operation then we need another element which corresponds to a constant translation part in a linear transformation: z = w1*x1 + w2*x2 + const.. Take a simple function y=w*x + c. The “c” controls where the line crosses the y axis. We need such a parameter if our line should separate clusters of points separably distributed somewhere in the (x,y)-plane; the w is not sufficient to orientate and position the hyperplane in the (x,y)-plane.
This very basically what bias neurons are good for regarding the basically linear operation between two MLP-layers. They add a constant to an otherwise linear transformation.
Do we need a bias node on all layers? Definitely on the input layer. However, on the hidden layers a trained network could by learning evolve weights in such a way that a bias neuron comes about – with almost zero weights on one side. At least in principle; however, we make it easier for the MLP to converge by providing explicit “bias” neurons.
What did we do to account for bias nodes in our Python code so far? We extended the matrices describing the output arrays ay_A_out of the activation function (for input ay_Z_in) on the input and all hidden layers by elements of an additional row. This was done by the method “add_bias_neuron_to_layer()” – see the codes given in article III.
The important point is that our weight matrices already got a corresponding dimension when we built them; i.e. we defined weights for the bias nodes, too. Of course, during optimization we must calculate partial derivatives of the cost function with respect to these weights.
The problem is:
We need to back propagate a delta-matrix Nδ for layer N via
( (NW.T).dot(Nδ) ). But then we can not apply a simple element-wise matrix multiplication with the (N-1)D(z)-matrix at layer N-1. Reason: The dimensions do not fit, if we calculate the elements of D only for the existing Z-Values at layer N-1.
There are two solutions for coding:
- We can add a row artificially and intermediately to the Z-matrix to calculate the D-matrix, then calculate NδS as
( (NW.T).dot(Nδ) ) * (N_1)D
and eliminate the first artificial row appearing in NδS afterwards. - The other option is to reduce the weight-matrix (NW) by a row intermediately and restore it again afterwards.
What we do is a matter of efficiency; in our coding we shall follow the first way and test the difference to the second way afterwards.
Check the matrix dimensions
As all steps to back-propagate and to circumvent the pitfalls require a bit of matrix wizardry we should at least check at every step during EBP backward-propagation that the dimensions of the involved matrices fit each other.
Outlook
Guys, after having explained some of the matrix math in the previous article of this series and the problems we have to tackle whilst programming the EBP-algorithm we are eventually well prepared to add EBP-methods to our Python class for MLP simulation. We are going to to this in the next article:
A simple program for an ANN to cover the Mnist dataset – VIII – coding Error Backward Propagation
Literature
“Machine Learning – An Applied Mathematics Introduction”, Paul Wilmott, 2019, Panda Ohana Publishing