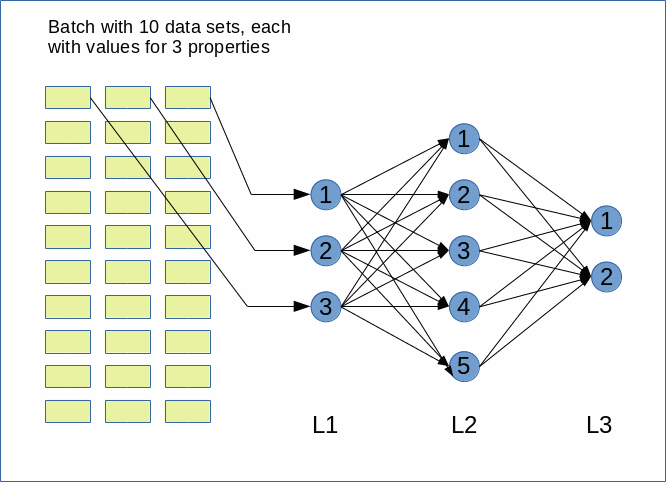
In this article series we are going to build a relatively simple Python class for the simulation of a “Multilayer Perceptron” [MLP]. A MLP is a simple form of an “artificial neural network” [ANN] with multiple layers of neurons. It has three characteristic properties: (1) Only connections between nodes of neighboring layers are allowed. (2) Information transport occurs in one forward direction. We speak of a “forward propagation of input information” through the ANN. (3) Neighboring layers are densely connected; a node of layer L_n is connected to all nodes of layer L_(n+1).
The firsts two points mean simplifications: According to (1) we do not consider so called cascaded networks. According to (2) no loops occur in the information transport. The third point, however, implies a lot of mathematical operations – not only during forward propagation of information through the ANN, but also – as we shall see – during the training and optimization of the network where we will back propagate “errors” from the output to the input layers. But for the next articles we need to care about simpler things first.
We shall use our MLP for classification tasks. Our first objective is to apply it to the MNIST dataset. In my last article
A simple program for an ANN to cover the Mnist dataset – I – a starting point
I presented already some code for the “__init__”-function on some other methods of our Python class. It enabled us to import MNIST data and split them in to a set of training and a set of test samples. Note that this is a standard approach in Machine Learning: You train on one set of data samples, but you test the classification or regression abilities of your ANN on a separate disjunctive data set.
In the present article we shall extend the functionality of our class: First we shall equip our network layers with a defined number of nodes. Then we shall provide statistical initial values for the “weights” describing the forward transport of information along the connections between the layers of our network.
Present status of our function “__init__”
The present status of our __init__function is:
def __init__(self,
my_data_set = "mnist",
n_hidden_layers = 1,
ay_nodes_layers = [0, 100, 0], # array which should have as much elements as n_hidden + 2
n_nodes_layer_out = 10, # expected number of nodes in output layer
my_activation_function = "sigmoid",
my_out_function = "sigmoid",
n_mini_batch = 1000, # number of data elements in a mini-batch
vect_mode = 'cols',
figs_x1=12.0, figs_x2=8.0,
legend_loc='upper right',
b_print_test_data = True
):
'''
Initialization of MyANN
Input:
data_set: type of dataset; so far only the "mnist", "mnist_784" datsets are known
We use this information to prepare the input data and learn about the feature dimension.
This info is used in preparing the size of the input layer.
n_hidden_layers = number of hidden layers => between input layer 0 and output layer n
ay_nodes_layers = [0, 100, 0 ] : We set the number of nodes in input layer_0 and the output_layer to zero
Will be set to real number afterwards by infos from the input dataset.
All other numbers are used for the node numbers of the hidden layers.
n_nodes_out_layer = expected number of nodes in the output layer (is checked);
this number corresponds to the number of categories NC = number of labels to be distinguished
my_activation_function : name of the activation function to use
my_out_function : name of the "activation" function of the last layer which produces the output values
n_mini_batch : Number of elements/samples in a mini-batch of training dtaa
vect_mode: Are 1-dim data arrays (vctors) ordered by columns or rows ?
figs_x1=12.0, figs_x2=8.0 : Standard sizing of plots ,
legend_loc='upper right': Position of legends in the plots
b_print_test_data: Boolean variable to control the print out of some tests data
'''
# Array (Python list) of known input data sets
self._input_data_sets = ["mnist", "mnist_784", "mnist_keras"]
self._my_data_set = my_data_set
# X, y, X_train, y_train, X_test, y_test
# will be set by analyze_input_data
# X: Input array (2D) - at present status of MNIST image data, only.
# y: result (=classification data) [digits represent categories in the case of Mnist]
self._X = None
self._X_train = None
self._X_test = None
self._y = None
self._y_train = None
self._y_test = None
# relevant dimensions
# from input data information; will be set in handle_input_data()
self._dim_sets = 0
self._dim_features = 0
self._n_labels = 0 # number of unique labels - will be extracted from y-data
# Img sizes
self._dim_img = 0 # should be sqrt(dim_features) - we assume square like images
self._img_h = 0
self._img_w = 0
# Layers
# ------
# number of hidden layers
self._n_hidden_layers = n_hidden_layers
# Number of total layers
self._n_total_layers = 2 + self._n_hidden_layers
# Nodes for hidden layers
self._ay_nodes_layers = np.array(ay_nodes_layers)
# Number of nodes in output layer - will be checked against information from target arrays
self._n_nodes_layer_out = n_nodes_layer_out
# Weights
# --------
# empty List for all weight-matrices for all layer-connections
# Numbering :
# w[0] contains the weight matrix which connects layer 0 (input layer ) to hidden layer 1
# w[1] contains the weight matrix which connects layer 1 (input layer ) to (hidden?) layer 2
self._ay_w = []
# Known Randomizer methods ( 0: np.random.randint, 1: np.random.uniform )
# ------------------
self.__ay_known_randomizers = [0, 1]
# Types of activation functions and output functions
# ------------------
self.__ay_activation_functions = ["sigmoid"] # later also relu
self.__ay_output_functions = ["sigmoid"] # later also softmax
# the following dictionaries will be used for indirect function calls
self.__d_activation_funcs = {
'sigmoid': self._sigmoid,
'relu': self._relu
}
self.__d_output_funcs = {
'sigmoid': self._sigmoid,
'softmax': self._softmax
}
# The following variables will later be set by _check_and set_activation_and_out_functions()
self._my_act_func = my_activation_function
self._my_out_func = my_out_function
self._act_func = None
self._out_func = None
# number of data samples in a mini-batch
self._n_mini_batch = n_mini_batch
# print some test data
self._b_print_test_data = b_print_test_data
# Plot handling
# --------------
# Alternatives to resize plots
# 1: just resize figure 2: resize plus create subplots() [figure + axes]
self._plot_resize_alternative = 1
# Plot-sizing
self._figs_x1 = figs_x1
self._figs_x2 = figs_x2
self._fig = None
self._ax = None
# alternative 2 does resizing and (!) subplots()
self.initiate_and_resize_plot(self._plot_resize_alternative)
# ***********
# operations
# ***********
# check and handle input data
self._handle_input_data()
print("\nStopping program regularily")
sys.exit()
The kind reader may have noticed that this is not exactly what was presented in the last article. I have introduced two additional parameters and corresponding class attributes:
“b_print_test_data” and “n_mini_batch”.
The first of these parameters controls whether we print out some test data or not. The second parameter controls the number of test samples dealt with in parallel during propagation and optimization via the so called “mini-batch-approach” mentioned in the last article.
Setting node numbers of the layers
We said in the last article that we would provide the numbers of nodes of the ANN layers via a parameter list “ay_nodes_layers”. We set the number of nodes for the input and the output layer, i.e. the first number and the last number in the list, to “0” in this array because these numbers are determined by properties of the input data set – here the MNIST dataset.
All other numbers in the array determine the amount of nodes of the hidden layers in consecutive order between the input and the output layer. So, the number at ay_nodes_layers[1] is the number of nodes in the first hidden layer, i.e. the layer which follows after the input layer.
In the last article we have understood already that the number of nodes in the input layer should be equal to the full number of “features” of our input data set – 784 in our case. The number of nodes of the output layer must instead be determined from the number of categories in our data set. This is equivalent to the number of distinct labels in the set of training data represented by an array “_y_train” (in case of MNIST: 10).
We provide three methods to check the node numbers defined by the user, set the node numbers for the input and output layers and print the numbers.
# Method which checks the number of nodes given for hidden layers
def _check_layer_and_node_numbers(self):
try:
if (self._n_total_layers != (self._n_hidden_layers + 2)):
raise ValueError
except ValueError:
print("The assumed total number of layers does not fit the number of hidden layers + 2")
sys.exit()
try:
if (len(self._ay_nodes_layers) != (self._n_hidden_layers + 2)):
raise ValueError
except ValueError:
print("The number of elements in the array for layer-nodes does not fit the number of hidden layers + 2")
sys.exit(1)
# Method which sets the number of nodes of the input and the layer
def _set_nodes_for_input_output_layers(self):
# Input layer: for the input layer we do
NOT take into account a bias node
self._ay_nodes_layers[0] = self._dim_features
# Output layer: for the output layer we check the number of unique values in y_train
try:
if ( self._n_labels != (self._n_nodes_layer_out) ):
raise ValueError
except ValueError:
print("The unique elements in target-array do not fit number of nodes in the output layer")
sys.exit(1)
self._ay_nodes_layers[self._n_total_layers - 1] = self._n_labels
# Method which prints the number of nodes of all layers
def _show_node_numbers(self):
print("\nThe node numbers for the " + str(self._n_total_layers) + " layers are : ")
print(self._ay_nodes_layers)
The code should be easy to understand. self._dim_features was set in the method “_common_handling_of mnist()” discussed in the last article. It was derived from the shape of the input data array _X_train. The number of unique labels was evaluated by the method “_get_num_labels()” – also discussed in the last article.
Note: The initial node numbers DO NOT include a bias node, yet.
If we extend the final commands in the “__init__”-function by :
# ***********
# operations
# ***********
# check and handle input data
self._handle_input_data()
# check consistency of the node-number list with the number of hidden layers (n_hidden)
self._check_layer_and_node_numbers()
# set node numbers for the input layer and the output layer
self._set_nodes_for_input_output_layers()
self._show_node_numbers()
print("\nStopping program regularily")
sys.exit()
By testing our additional code in a Jupyter notebook we get a corresponding output for
ANN = myann.MyANN(my_data_set="mnist_keras", n_hidden_layers = 2,
ay_nodes_layers = [0, 100, 50, 0],
n_nodes_layer_out = 10,
vect_mode = 'cols',
figs_x1=12.0, figs_x2=8.0,
legend_loc='upper right',
b_print_test_data = False
)
of:
The node numbers for the 4 defined layers are: [784 100 50 10]
Good!
Setting initial random numbers for the weights
Initial values for the ANN weights have to be given as matrices, i.e. 2-dim arrays. However, the randomizer functions provided by Numpy give you vectors as output. So, we need to reshape such vectors into the required form.
First we define a method to provide random floating point and integer numbers:
# ---
# method to create an array of randomized values
def _create_vector_with_random_values(self, r_low=None, r_high=None, r_size=None, randomizer=0 ):
'''
Method to create a vector of length "r_size" with "random values" in [r_low, r_high]
generated by method "randomizer"
Input:
ramdonizer : integer which sets randomizer method; presently only
0: np.random.uniform
1: np.randint
[r_low, r_high]: range of the random numbers to be created
r_size: Size of output array
Output: A 1-dim numpy array of length rand-size - produced as a class member
'''
# check parameters
try:
if (r_low==None or r_high == None or r_size == None ):
raise ValueError
except ValueError:
print("One of
the required parameters r_low, r_high, r_size has not been set")
sys.exit(1)
rmizer = int(randomizer)
try:
if (rmizer not in self.__ay_known_randomizers):
raise ValueError
except ValueError:
print("randomizer not known")
sys.exit(1)
# 2 randomizers (so far)
if (rmizer == 0):
ay_r_out = np.random.randint(int(r_low), int(r_high), int(r_size))
if (rmizer == 1):
ay_r_out = np.random.uniform(r_low, r_high, size=int(r_size))
return ay_r_out
Presently, we can only use two randomizer functions to be used – numpy.random.randint and numpy.random.uniform. The first one provides random integer values, the other one floating point values – both within a defined interval. The parameter “r_size” defines how many random numbers shall be created and put into an array. The code requires no further explanation.
Now, we define two methods to create the weight matrices for the connections
- between the input layer L0 to the first hidden layer,
- between further hidden layers and eventually between the last hidden layer and the output layer
As we allow for all possible connections between nodes the dimensions of the matrices are determined by the numbers of nodes in the connected neighboring layers. Each node of a layer L_n cam be connected to each node of layer L_(n+1). Our methods are:
# Method to create the weight matrix between L0/L1
# ------
def _create_WM_Input(self):
'''
Method to create the input layer
The dimension will be taken from the structure of the input data
We need to fill self._w[0] with a matrix for conections of all nodes in L0 with all nodes in L1
We fill the matrix with random numbers between [-1, 1]
'''
# the num_nodes of layer 0 should already include the bias node
num_nodes_layer_0 = self._ay_nodes_layers[0]
num_nodes_with_bias_layer_0 = num_nodes_layer_0 + 1
num_nodes_layer_1 = self._ay_nodes_layers[1]
# fill the matrix with random values
rand_low = -1.0
rand_high = 1.0
rand_size = num_nodes_layer_1 * (num_nodes_with_bias_layer_0)
randomizer = 1 # method np.random.uniform
w0 = self._create_vector_with_random_values(rand_low, rand_high, rand_size, randomizer)
w0 = w0.reshape(num_nodes_layer_1, num_nodes_with_bias_layer_0)
# put the weight matrix into array of matrices
self._ay_w.append(w0.copy())
print("\nShape of weight matrix between layers 0 and 1 " + str(self._ay_w[0].shape))
# Method to create the weight-matrices for hidden layers
def _create_WM_Hidden(self):
'''
Method to create the weights of the hidden layers, i.e. between [L1, L2] and so on ... [L_n, L_out]
We fill the matrix with random numbers between [-1, 1]
'''
# The "+1" is required due to range properties !
rg_hidden_layers = range(1, self._n_hidden_layers + 1, 1)
# for random operation
rand_low = -1.0
rand_high = 1.0
for i in rg_hidden_layers:
print ("Creating weight matrix for layer " + str(i) + " to layer " + str(i+1) )
num_nodes_layer = self._ay_nodes_layers[i]
num_nodes_with_bias_layer = num_nodes_layer + 1
# the number of the next layer is taken without the bias node!
num_nodes_layer_
next = self._ay_nodes_layers[i+1]
# assign random values
rand_size = num_nodes_layer_next * num_nodes_with_bias_layer
randomizer = 1 # np.random.uniform
w_i_next = self._create_vector_with_random_values(rand_low, rand_high, rand_size, randomizer)
w_i_next = w_i_next.reshape(num_nodes_layer_next, num_nodes_with_bias_layer)
# put the weight matrix into our array of matrices
self._ay_w.append(w_i_next.copy())
print("Shape of weight matrix between layers " + str(i) + " and " + str(i+1) + " = " + str(self._ay_w[i].shape))
Three things may need explanation:
- The first thing is the inclusion of a bias-node in all layers. A bias node only provides input to the next layer but does not receive input from a preceding layer. It gives an additional bias to the input a layer provides to a neighbor layer. (Actually, it provides an additional degree of freedom for the optimization process.) So, we include connections from a bias-node of a layer L_n to all nodes (without the bias-node) in layer L_(n+1).
- The second thing is the reshaping of the vector of random numbers into a matrix. Of course, the dimension of the vector of random numbers must fit the product of the 2 matrix dimensions. Note that the first dimension always corresponds to number of nodes in layer L_(n+1) and the second dimension to the number of nodes in layer L_n (including the bias-node)!
We need this special form to support the vectorized propagation properly later on. - The third thing is that we save our (Numpy) weight matrices as elements of a Python list.
In my opinion this makes the access to these matrices flexible and easy in the case of multiple hidden layers.
Setting the activation and output functions
We must set the activation and the output function. This is handled by a method “_check_and_set_activation_and_out_functions()”.
def _check_and_set_activation_and_out_functions(self):
# check for known activation function
try:
if (self._my_act_func not in self.__d_activation_funcs ):
raise ValueError
except ValueError:
print("The requested activation function " + self._my_act_func + " is not known!" )
sys.exit()
# check for known output function
try:
if (self._my_out_func not in self.__d_output_funcs ):
raise ValueError
except ValueError:
print("The requested output function " + self._my_out_func + " is not known!" )
sys.exit()
# set the function to variables for indirect addressing
self._act_func = self.__d_activation_funcs[self._my_act_func]
self._out_func = self.__d_output_funcs[self._my_out_func]
if self._b_print_test_data:
z = 7.0
print("\nThe activation function of the standard neurons was defined as \"" + self._my_act_func + '"')
print("The activation function gives for z=7.0: " + str(self._act_func(z)))
print("\nThe output function of the neurons in the output layer was defined as \"" + self._my_out_func + "\"")
print("The output function gives for z=7.0: " + str(self._out_func(z)))
It requires not much of an explanation. For the time being we just rely on the given definition of the “__init__”-interface, which sets both to “sigmoid()”. The internal dictionaries __d_activation_funcs[] and __d_output_funcs[] provide the functions to
internal variables (indirect addressing).
Some test output
A test output for the enhanced code
# ***********
# operations
# ***********
# check and handle input data
self._handle_input_data()
# check consistency of the node-number list with the number of hidden layers (n_hidden)
self._check_layer_and_node_numbers()
# set node numbers for the input layer and the output layer
self._set_nodes_for_input_output_layers()
self._show_node_numbers()
# create the weight matrix between input and first hidden layer
self._create_WM_Input()
# create weight matrices between the hidden layers and between tha last hidden and the output layer
self._create_WM_Hidden()
# check and set activation functions
self._check_and_set_activation_and_out_functions()
print("\nStopping program regularily")
sys.exit()
and a Jupyter cell
gives :
The shapes of the weight matrices correspond correctly to the numbers of nodes in the 4 layers defined. (Do not forget about the bias-nodes!).
Conclusion
We have reached a status where our ANN class can read in the MNIST dataset and set initial random values for the weights. This means that we can start to do some more interesting things. In the next article
A simple program for an ANN to cover the Mnist dataset – III – forward propagation
we shall program the “forward propagation”. We shall perform the propagation for a mini-batch of many data samples in one step. We shall see that this is a very simple task, which only requires a few lines of code.