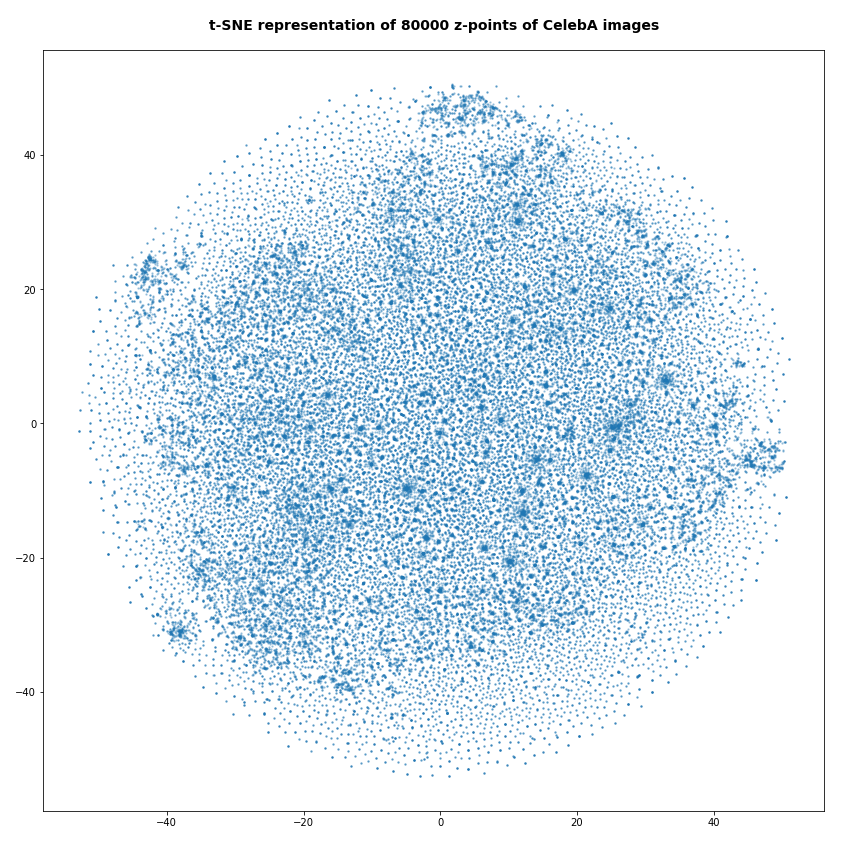
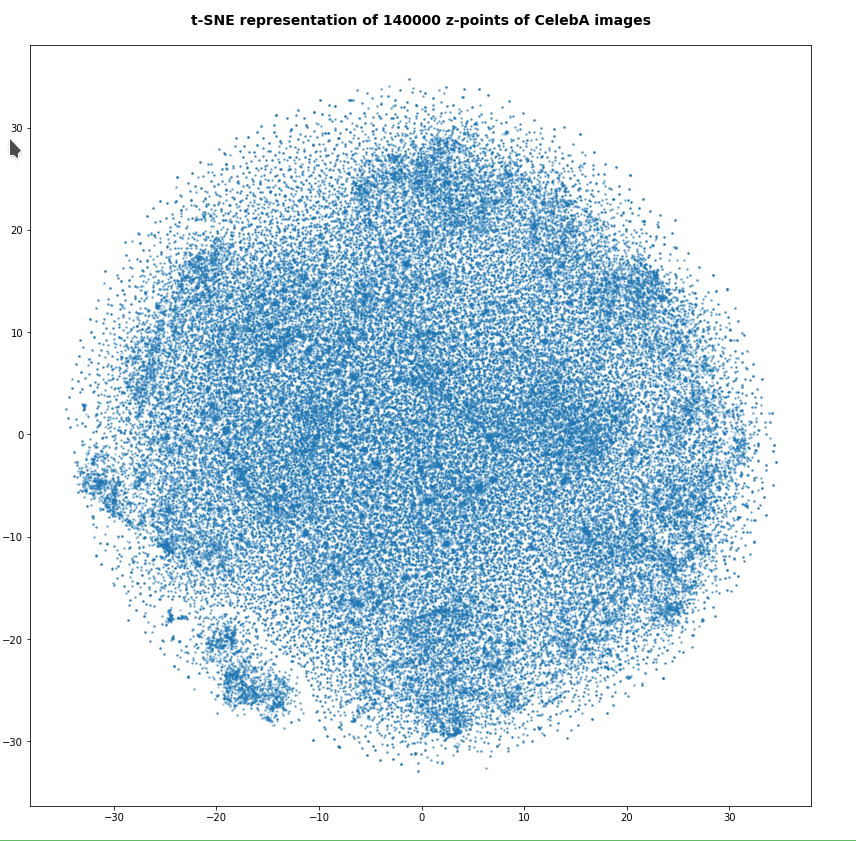
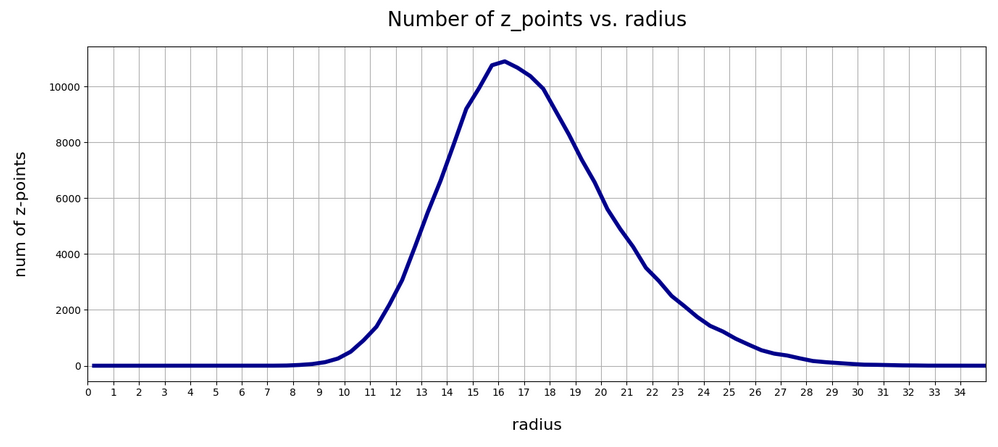
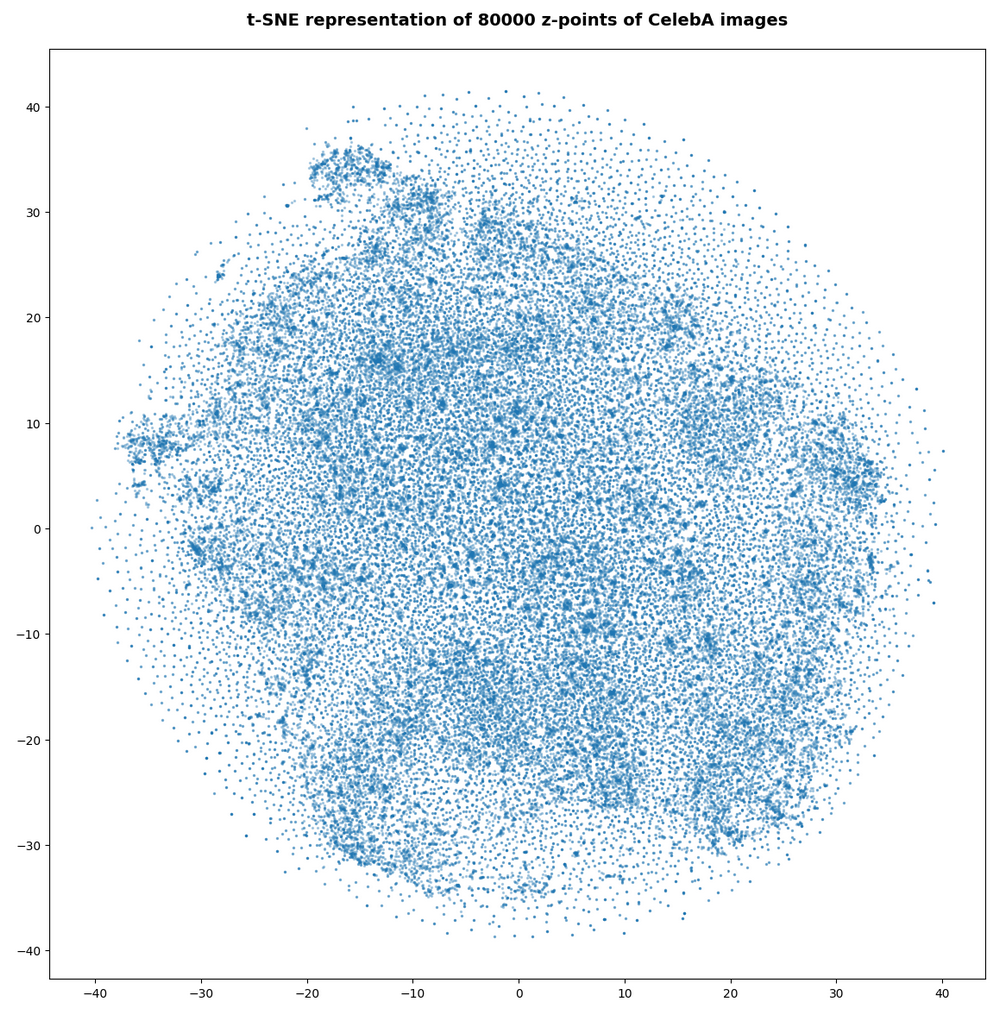
In my last post of this series I compared a Variational Autoencoder [VAE] with only a tiny amount of KL-loss to a standard Autoencoder [AE]. See the links at
for more information and preparational posts.
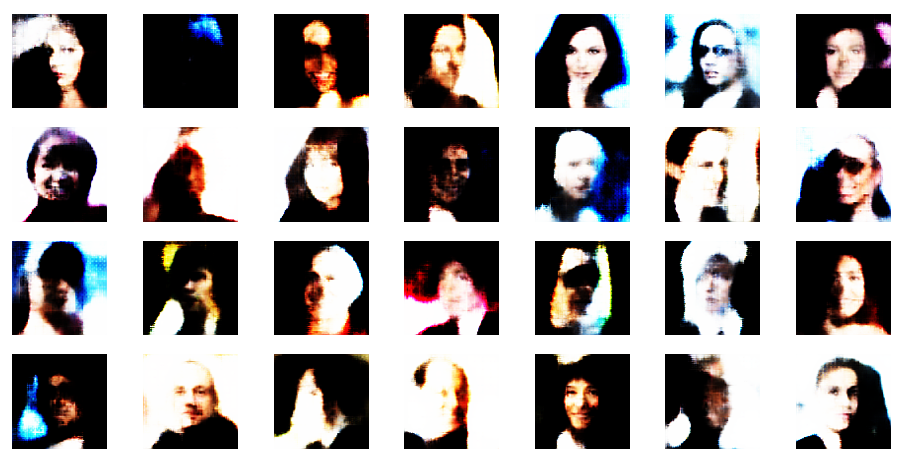
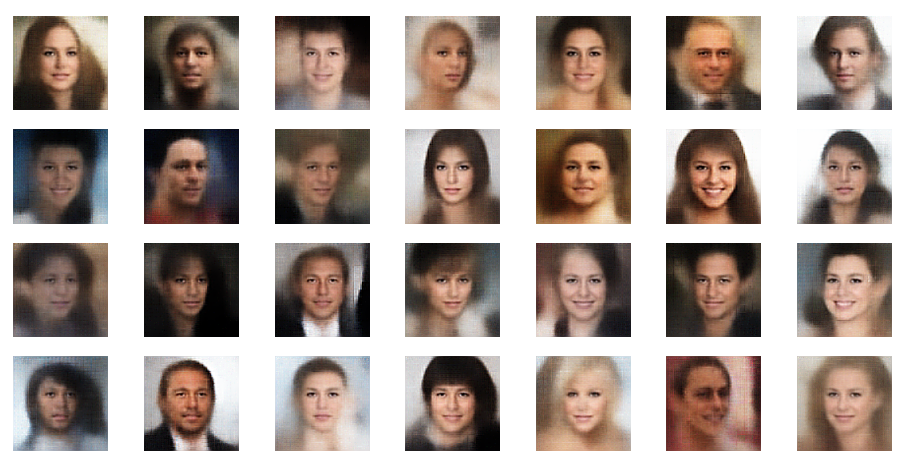
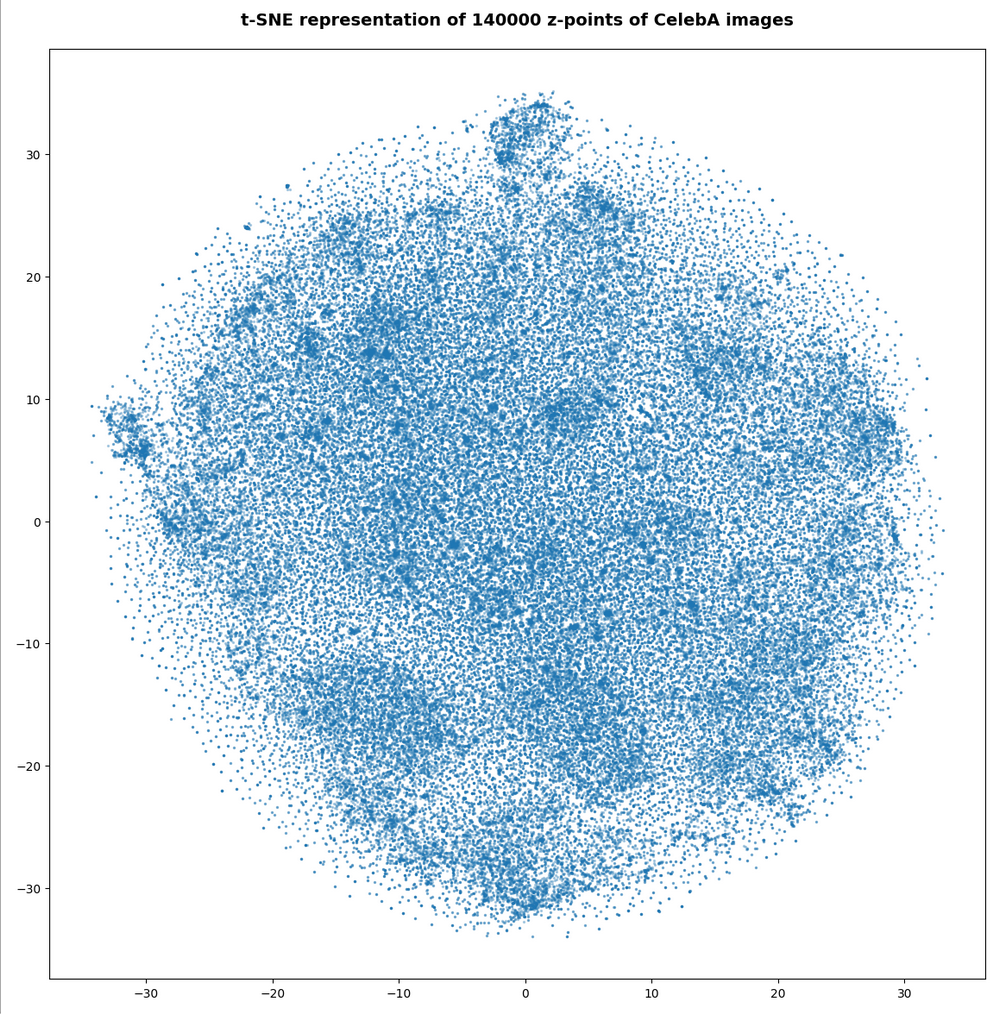
Both the Keras models for the VAE and the AE were trained on the CelebA dataset of images of human heads. We found a tight similarity regarding the clustering of predicted data points for the training and test data in the latent space. In addition the VAE with tiny KL loss failed to reconstruct reasonable human face images from arbitrarily chosen points in the latent space. Just as a standard AE does. In forthcoming posts we will continue to study the relation between VAEs and AEs.
But in this post I want to briefly point out an interesting technical problem which may arise when you start to tests predictions for certain data samples after a training phase. Your Encoder or Decoder models may include parameters which you want to experiment with when predicting results for interesting input data. This raises the question whether we can vary such parameters at inference time. Actually, this was not quite as easy as it seemed to be when I started with respective experiments. To perform them I had to learn two aspects of Keras models I had not been aware of before.
How to switch of the z-point variation at inference time?
In my particular case the starting point was the following consideration:
At inference time there is no real need for using the logvar-based variation around mu-values predicted by the Encoder.
The variation in z-point values in VAEs is done by adding a statistical value to mu-values. The added term is based on a log_var value multiplied by a statistically fluctuating factor “epsilon”, which comes from a normal Gaussian distribution around zero. mu, therefore, is the center of a distribution a specific input tensor is mapped to in latent space for consecutive predictions. The mu- and log_var-values depend on weights of two dense layers of the Encoder and thus indirectly on the optimization during training.
But while the variation is essential during training one may NOT regard it necessary for predictions. During inference we may in some experiments have good reasons to only refer to the central mu value when predicting a z-point in latent space. For test and analysis purposes it could be interesting to omit the log_var contribution.
The question then is: How we can switch off the log_var component for the Encoder’s predictions, i.e. predictions of our Keras based encoder model?
One idea is to include variables of a Python class hosting the Keras models for the Encoder, Decoder and the composed VAE in the function for the calculation of z-point vectors.
The mu-, logvar and sampling layers of the VAE’s Encoder model encapsulated in a Python class
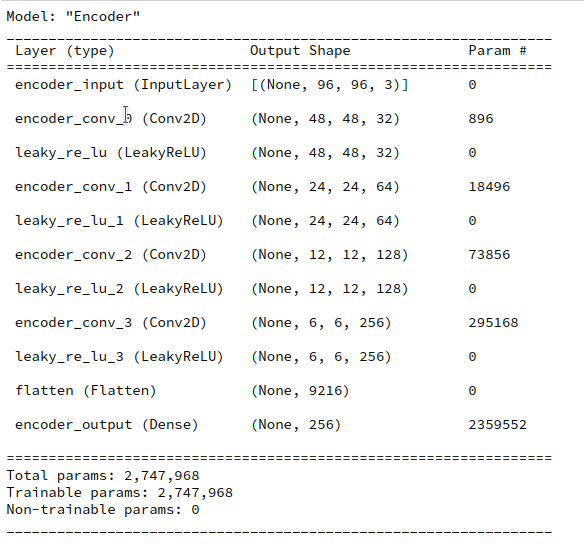
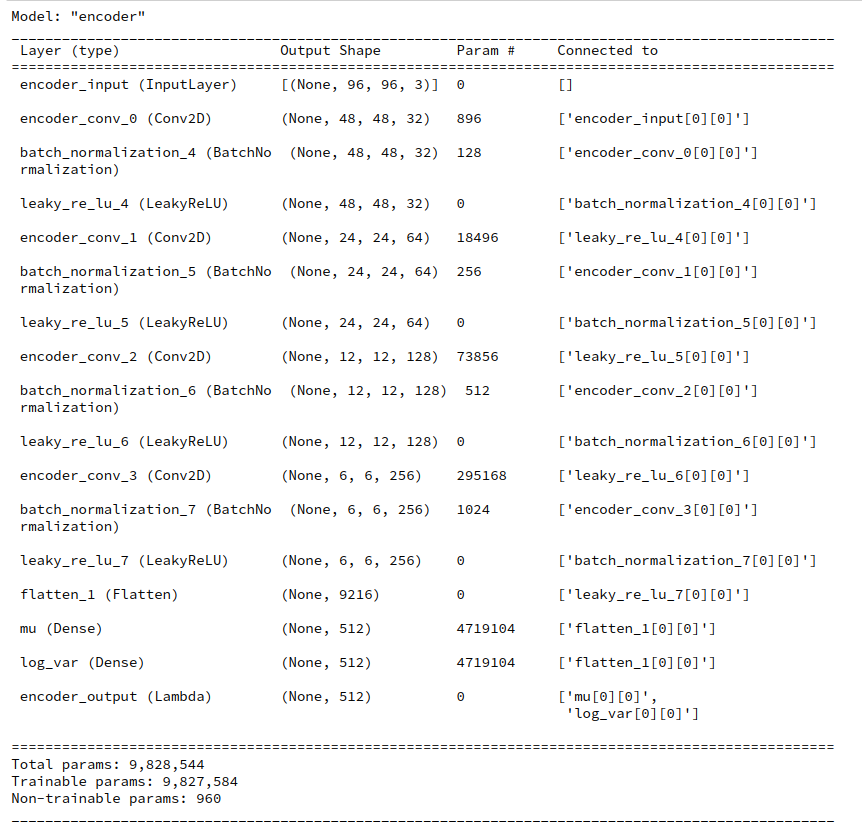
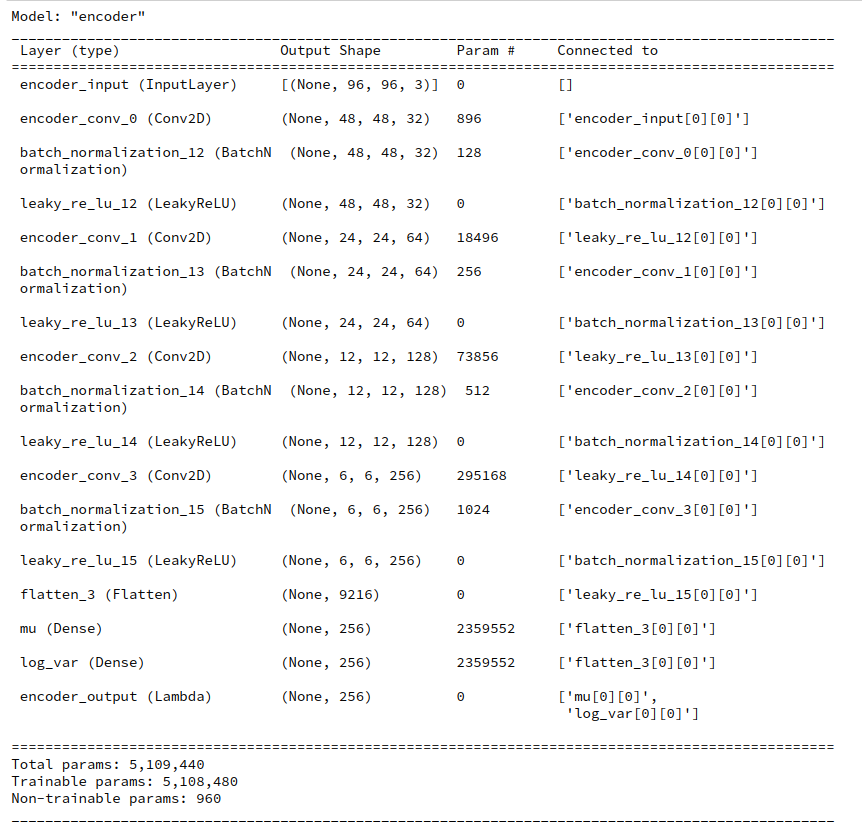
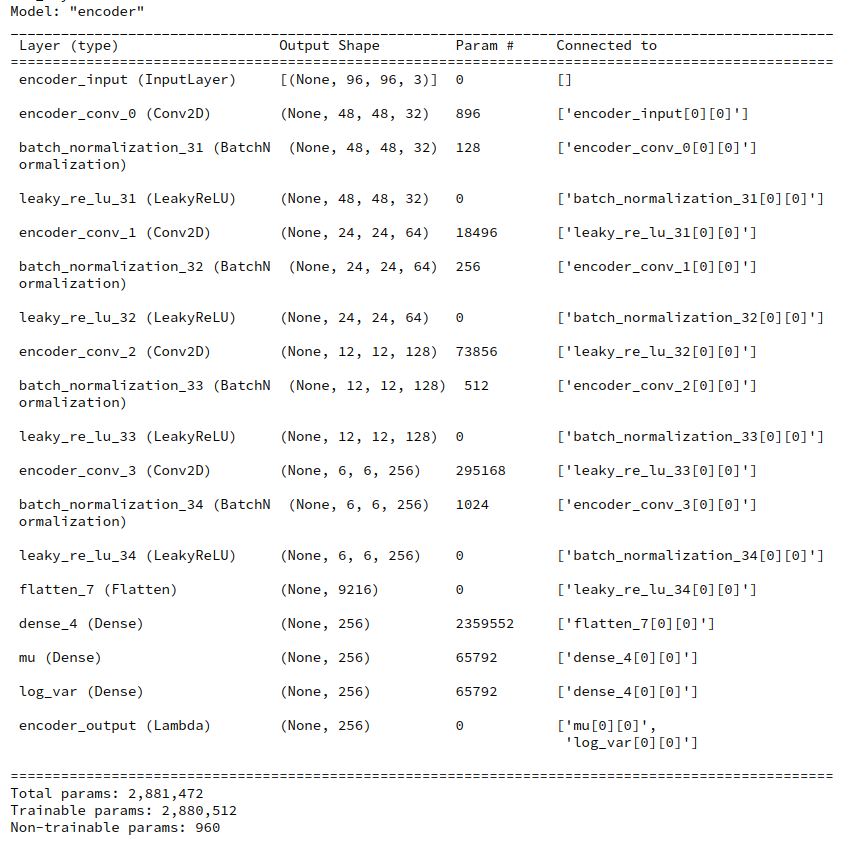
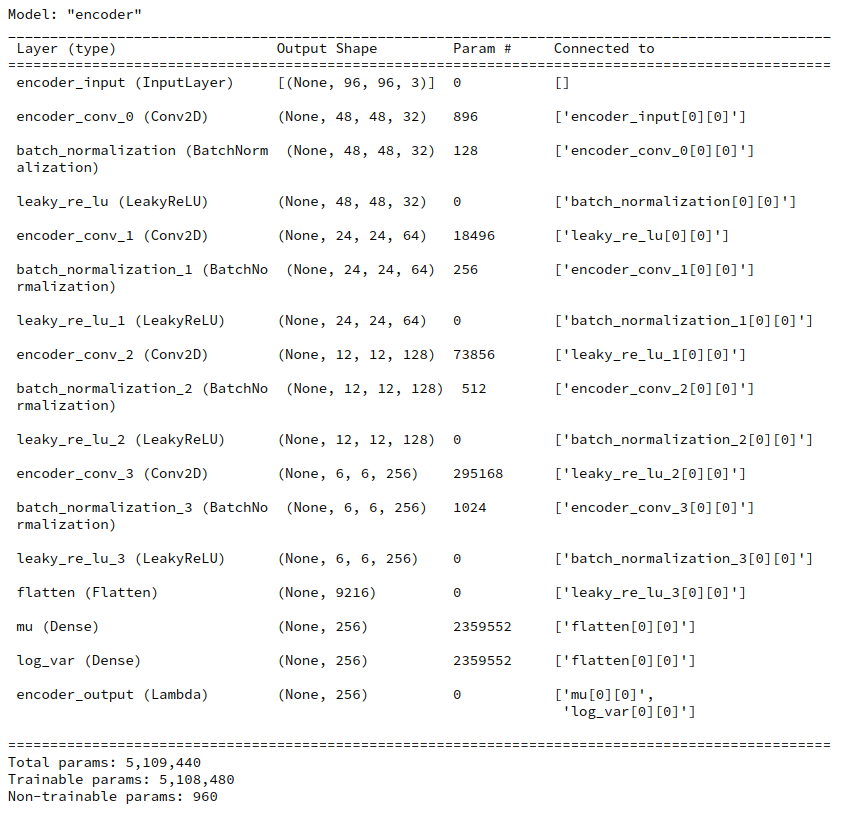
During this post series we have encapsulated the code for Encoder, Decoder and resulting VAE models in a Python class. Remember that the Encoder produced its output, namely z-points in the latent space via two “dense” Keras layers and a sampling layer (based on a Keras Lambda-layer). The dense layers followed a series of convolutional layers and gave us mu and log_var values. The Lambda-layer produced the eventual z-point-vector including the variation. In my case the code fragments for the layers look similar to the following ones:
# .... Layer model of the Encoder part
...
... # Definition of an input layer and multiple Conv2D layers
...
# "Variational" part - 2 Dense layers for a statistical distribution of z-points
self.mu = Dense(self.z_dim, name='mu')(x)
self.log_var = Dense(self.z_dim, name='log_var')(x)
# Layer to provide a z_point in the Latent Space for each sample of the batch
self._encoder_output = Lambda(z_point_sampling, name='encoder_output')([self.mu, self.log_var])
...
# The Encoder model
self.encoder = Model(inputs=self._encoder_input, outputs=[self._encoder_output, self.mu, self.log_var], name="encoder")
...
The “self” refers to a class “MyVariationalAutoencoder” comprising the Encoder’s, Decoder’s and the VAE’s model and layer structures. See for details and explained code fragments of the class e.g. the posts around Variational Autoencoder with Tensorflow 2.8 – X – VAE application to CelebA images.
The sampling is in my case done by a function “z_point_sampling”:
def z_point_sampling(args):
'''
A point in the latent space is calculated statistically
around an optimized mu for each sample
'''
mu, log_var = args # Note: These are 1D tensors !
epsilon = B.random_normal(shape=B.shape(mu), mean=0., stddev=1.)
return mu + B.exp(log_var / 2.) * epsilon * self.enc_v_eps_factor
You see that this function uses a class member variable “self.enc_v_eps_factor”.
Switch the variation with log_var on and off for predictions?
Our objective is to switch the log_var contribution on or off for input certain images or batches of such images fed into the Encoder. For this purpose we could in principle use the variable “self.enc_v_eps_factor” as a kind of boolean switch with values of either 0.0 or 1.0. To set the variable I had defined two class methods:
def set_enc_to_predict(self):
self.enc_v_eps_factor = 0.0
def set_enc_to_train(self):
self.enc_v_eps_factor = 1.0
The basic idea was that the sampling function would pick the value of enc_v_eps_factor given at the runtime of a prediction, i.e. at inference time. This assumption was, however, wrong. At least in a certain sense.
Is a class variable change impacting a layer output noted during consecutive predictions of a Keras model?
Let us assume that we have instantiated our class and assigned it to a Python variable MyVae. Let us further assume the the comprised Keras models are referenced by variables
- MyVae.encoder (for the Encoder part),
- MyVae.decoder (for the Decoder part)
- and MyVae.model (for the full VAE-model).
We do not care about further details of the VAE (consisting of the Encoder, the Decoder and GradientTape based cost control). But we should not forget that all the models’ layers and their weights determine cost functions derivatives and are therefore targets of the optimization performed during training. All factors determining gradients and value calculation with given weights are encoded with the compilation of a Keras model – for training purposes [using model.fit() with a Keras model], but as well for predictions! Without a compiled Keras model we cannot use model.predict().
This means: As soon as you have a compiled Keras model almost and load the weight-values saved after a a sufficient number of training epochs everything is settled for inference and predictions. Including the present value of self.enc_v_eps_factor. At compile time?
Well, thinking about it a bit more in detail from a developer perspective tells us:
The compilation would in principle not prevent the use of a changed variable at the run-time of a prediction. But on the other hand side we also have the feeling that Keras must do something to make training (which also requires predictions in the sense of a forward pass) and later raw predictions for batches at inference time pretty fast. Intermediate functionality changes would hamper performance – if only for the reason that you have to watch out for such changes.
So, it is natural to assume that Keras would keep any factors in the Lambda-layer taken from a class variable constant after compilation and during training or inference, i.e. predictions.
If this assumption were true then chain of actions AFTER a training of a VAE model (!) like
Define a Keras based VAE-model with a sampling layer and a factor enc_v_eps_factor = 1 => compile it (including the sub-models for the Encoder and Decoder) => load weight parameters which had been saved after training => switch the value of the class variable enc_v_eps_factor to 0.0 => Load an image or image batch for prediction
would probably NOT work as expected. To be honest: This is “wisdom” derived after experiments which did not give me the naively expected results.
Indeed, some first simple experiments showed: The value of enc_v_eps_factor (e.g. enc_v_eps_factor = 1) which was given at compile-time was used during all following prediction calculations, e.g. for a particular image in tensorial form. So, a command sequence like
MyVae.set_enc_to_train() MyVae.encoder.compile() ... # Load weights from a set path MyVae.model.load_weights(path_weights) ... z_point, mu, log_var = MyVae.encoder.predict(img1) print(z_point, mu) MyVae.set_enc_to_predict(self) z_point, mu, log_var = MyVae.encoder.predict(img1)
did not give different results. Note, that I did not change the value of enc_v_eps_factor between compile time and the first call for a prediction.
Let us look at the example in more detail.
A concrete example
After a full training of my VAE on CelebA for 24 epochs I checked for a maximum log_var-value. Non-negligible values may indeed occur for certain images and special z-vector components for some images despite the tiny KL-loss. And indeed such a value occurred for a very special (singular) image with two diagonal black border stripes on the left and right of the photographed person’s head. I do not show the image due to digital rights concerns. But let us look at the predicted z, mu and log_var values (for a specific z-point vector component) of this image. Before I compiled the VAE model I had set enc_v_eps_factor = 1.0 :
# Set the learning rate and COMPILE the model # ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~ learning_rate = 0.0005 # The following is only required for compatibility reasons b_old_optimizer = True # Set enc_v_eps_factor to 1.0 MyVae.set_enc_to_train() # Separate Encoder compilation # - does not harm the compilation of the full VAE-model, but is useful to avoid later trace warnings after changes MyVae.encoder.compile() # Compilation of the full VAE model (Encoder, Decoder, cost functions used by GradientTape) MyVae.compile_myVAE(learning_rate=learning_rate, b_old_optimizer = b_old_optimizer ) ... # Load weights from a set path MyVae.model.load_weights(path_weights) ... # start predictions ...
with
def compile_myVAE(self, learning_rate, b_old_optimizer = False):
# Version 1.1 of 221212
# Forced to special handling of the optimizer for data resulting from training before TF 11.2 , due to warnings:
# ValueError: You are trying to restore a checkpoint from a legacy Keras optimizer into a v2.11+ Optimizer,
# which can cause errors. Please update the optimizer referenced in your code to be an instance
# of `tf.keras.optimizers.legacy.Optimizer`, e.g.: `tf.keras.optimizers.legacy.Adam`.
# Optimizer handling
# ~~~~~~~~~
if b_old_optimizer:
optimizer = tf.keras.optimizers.legacy.Adam(learning_rate=learning_rate)
else:
optimizer = Adam(learning_rate=learning_rate)
....
# Solution type with train_step() and GradientTape()
if self.solution_type == 3:
self.model.compile(optimizer=optimizer)
Details of the compilation are not important – though you may be interested in the fact that training data saved after a training based on Python module version < 11.2 of TF 2 requires a legacy version of the optimizer when later using a module version ≥ 11.2 (corresponding to TF V2.11 and above).
However, the really important point is that the compilation is done given a certain value of enc_v_eps_factor = 1.
Then we load the image in form a prepared training batch with just one element set and provide it to the predict() function of the Keras model for the Encoder. We perform two predictions and ahead of the second one we change the value of enc_v_eps_factor to 0.0:
# Choose an image j = 123223 img = x_train[j] # this has already a tensor compatible format img_list = [] img_list.append(img) tf_img = tf.convert_to_tensor(img_list) # Encoder prediction z_points, mu, logvar = MyVae.encoder.predict(tf_img) print(z_points[0][230]) print(mu[0][230]) print(logvar[0][230]) # !!!! Set enc_v_eps_factor to 0.0 !!!! MyVae.set_enc_to_predict() # New Encoder prediction z_points, mu, logvar = MyVae.encoder.predict(tf_img) print() print() print(z_points[0][230]) print(mu[0][230]) print(logvar[0][230])
The result is:
... 2.637196 -0.141142 3.3761873 ... -0.2085279 -0.141142 3.3761873
The result depends on statistical variations for the factor epsilon (Gaussian statistics; see the sampling function above).
But the central point is not the deviation for the two different prediction calls. The real point is that we have used MyVae.set_enc_to_predict() ahead of the second prediction and, yet, the values for mu and log_var for the special z_point-component (230; out of 256 components) were NOT identical. I.e. the variable value enc_v_eps_factor = 1.0, which we set before the compilation, was used during both of our prediction calculations!
Can we just recompile between different calls to model.predict() ?
The experiment described above seems to indicate that the value for the class variable enc_v_eps_factor given at compile time is used during all consecutive predictions. We could, of course, enforce a zero variation for all predictions by using MyVae.set_enc_to_predict() ahead of the compilation of the Encoder model. But this would not give us no flexibility to switch the log_var contribution off ahead of predictions for some special images and then turn it on again for other images.
But the is simple – if we need not do this switching permanently: We just recompile the Encoder model!
Compilation does not take much time for Encoder models with only a few (convolutional and dense) layers. Let us test this by modifying the code above:
# Choose an image j = 123223 img = x_train[j] # this has already a tensor compatible format img_list = [] img_list.append(img) tf_img = tf.convert_to_tensor(img_list) # Set enc_v_eps_factor to 0.0 MyVae.set_enc_to_predict() # !!!! MyVae.encoder.compile() # Encoder prediction z_points, mu, logvar = MyVae.encoder.predict(tf_img) # Decoder prediction - just for fun reco_list = MyVae.decoder.predict(z_points) # just for fun print(z_points[0][230]) print(mu[0][230]) print(logvar[0][230]) print() print() # Set enc_v_eps_factor back to 1.0 MyVae.set_enc_to_train() MyVae.encoder.compile() # New Encoder prediction z_points, mu, logvar = MyVae.encoder.predict(tf_img) print(z_points[0][230]) print(mu[0][230]) print(logvar[0][230])
We get
Shape of img_list = (1, 96, 96, 3) eps_fact = 0.0 1/1 [==============================] - 0s 280ms/step 1/1 [==============================] - 0s 18ms/step Shape of reco_list = (1, 96, 96, 3) -0.141142 -0.141142 3.3761873 eps_fact = 1.0 1/1 [==============================] - 0s 288ms/step -0.63676023 -0.141142 3.3761873
This is exactly what we want!
The function for the prediction step of a Keras model is cached at inference time …
The example above gave us the impression that it could be the compilation of a model which “settles” all of the functionality used during predictions, i.e. at inference time. Actually, this is no quite true.
The documentation on a Keras model helped me to get a better understanding. Near the section on the method “predict()” we find some other interesting functions. A look at the remarks on “predict_step()“, reveals (quotation)
The logic for one inference step. This method can be overridden to support custom inference logic. This method is called by Model.make_predict_function. This method should contain the mathematical logic for one step of inference. This typically includes the forward pass.
This leads us to the function “make_predict_function()” for Keras models. And there we find the following interesting remarks – I quote:
This method can be overridden to support custom inference logic. This method is called by Model.predict and Model.predict_on_batch. Typically, this method directly controls tf.function and tf.distribute.Strategy settings, and delegates the actual evaluation logic to Model.predict_step. This function is cached the first time Model.predict or Model.predict_on_batch is called. The cache is cleared whenever Model.compile is called. You can skip the cache and generate again the function with force=True.
Ah! The function predict_step() normally covers the forward pass through the network and “make_predict_function()” caches the resulting (function) object at the first invocation of model.predict(). And the respective cache is not cleared automatically.
So, what really may have hindered my changes of the sampling functionality at inference time is a cache filled at the first call to encoder.predict()!
Let us test this!
Changing the sampling parameters after compilation, but before the first call of encoder.predict()
If our suspicion is right we should be able to set up the model from scratch again, compile it, use MyVae.set_enc_to_predict() and afterward call MyVae.encoder.predict() – and get the same values for mu and z_point.
So we do something like
# Build encoder according to layer parameters
MyVae._build_enc()
# Build decoder according to layer parameters
MyVae._build_dec()
# Build the VAE-model
MyVae._build_VAE()
...
# Set variable to 1.0
MyVae.set_enc_to_train()
# Compile
learning_rate = 0.0005
b_old_optimizer = True
MyVae.compile_myVAE(learning_rate=learning_rate, b_old_optimizer = b_old_optimizer )
MyVae.encoder.compile() # used to prevent retracing delays - when later changing encoder variables
...
# Load weights from a set path
MyVae.model.load_weights(path_weights)
...
# preparation if the selected img.
...
...
MyVae.set_enc_to_predict()
print("eps_fact = ", MyVae.enc_v_eps_factor)
# Note: NO recompilation is done !
# First prediction
z_points, mu, logvar = MyVae.encoder.predict(tf_img)
print()
print(z_points[0][230])
print(mu[0][230])
print(logvar[0][230])
..
Note that the change of enc_v_eps_factor ahead of the first call of predict(). And, indeed:
Shape of img_list = (1, 96, 96, 3) eps_fact = 0.0 ... -0.141142 -0.141142 3.3761873
Use make_predict_function(force=True) to clear and refill the cache for predict_step() and its forward pass functionality
The other option the documentation indicates is to use the function make_predict_function(force=True).
This leads to yet another experiment:
# img preparation
....
# Set enc_v_eps_factor to 1.0
MyVae.set_enc_to_train()
MyVae.encoder.compile()
print("eps_fact = ", MyVae.enc_v_eps_factor)
# Encoder prediction
z_points, mu, logvar = MyVae.encoder.predict(tf_img)
print(z_points[0][230])
print(mu[0][230])
print(logvar[0][230])
print()
print()
# Set enc_v_eps_factor to 0.0
MyVae.set_enc_to_predict()
# !!!!
MyVae.encoder.make_predict_function(
force=True
)
print("eps_fact = ", MyVae.enc_v_eps_factor)
# Encoder prediction
z_points, mu, logvar = MyVae.encoder.predict(tf_img)
print(z_points[0][230])
print(mu[0][230])
print(logvar[0][230])
We get
... eps_fact = 1.0 1/1 [==============================] - 0s 287ms/step -5.5451365 -0.141142 3.3761873 eps_fact = 0.0 1/1 [==============================] - 0s 271ms/step -0.141142 -0.141142 3.3761873
Yes, exactly as expected. This again shows us that it is the cache which counts after the first call of model.predict() – and not the compilation of the Keras model (for the Encoder) !
Other approaches?
The general question of changing parameters at inference time also triggers the question whether we may be able to deliver parameters to the function model.predict() and transfer them further to customized variants of predict_step(). I found a similar question at stack overflow
Passing parameters to model.predict in tf.keras.Model
However, the example there was rather special – and I did not apply the lines of thought explained there to my own case. But the information given in the answer may still be useful for other readers.
Conclusion
In this post we have seen that we can change parameters influencing the forward pass of a Keras model at inference time. We saw, however, that we have to clear and fill a cache to make the changes effective. This can be achieved by
- either applying a recompilation of the model
- or enforcing a clearance and refilling of the cache for the model’s function predict_step().
In the special case of a VAE this allows for deactivating and re-activating the logvar-dependent statistical variation of the z-points a specific image is mapped to by the Encoder model during predictions. This gives us the option to focus on the central mu-dependent position of certain images in the latent space during experiments at inference time.
In the next post of this series we shall have a closer look at the filamental structure of the latent space of a VAE with tiny KL loss in comparison to the z-space structure of a VAE with sufficiently high KL loss.
Ceterum censeo: The worst fascist, war criminal and killer living today is the Putler. He must be isolated at all levels, be denazified and sooner than later be imprisoned. Somebody who orders the systematic destruction of civilian infrastructure must be fought and defeated because he is a permanent danger to basic principles of humanity – not only in Europe. Long live a free and democratic Ukraine!