I move a small step forward on my trials as a beginner in machine learning. This is once again going to be an article from which experts on “Artificial Neural networks” [ANNs] will learn nothing new. But some of my readers may also be newbies both to Python, to numpy and to ANNs. For them this article may be helpful.
I shall have a brief look at some matrix (or array) operations which constitute important parts of the information propagation on ANNs. How would such operations look like in Python with Numpy? One should have a clear understanding of the so called “dot”-operation between multidimensional arrays in this context. Such operations can be performed highly optimized on GPUs – and whole sets of data samples can be handled in form of vectorized code instructions. I will try to explain this in form of matrix terms required to simulate information transport between some layers of a very simple network.
Propagation operations on ANNs
Basic Artificial Neural Networks consist of layers with neurons. Each layer has a defined number of neurons. Each neuron is connected with all neurons of the next layer (receiving neurons). The output “O(k, L)” of a neuron N(k,L) of Layer “L” to a neuron N(m,L+1) is multiplied by some “weight” W[(m, L+1),(k, L),].
The contributions of all neurons on layer “L” to N(m, L+1) are summed up. The result is then transformed by some local “activation” function and afterwards presented by N(m, L+1) as output which shall be transported to the neurons of layer L+2. The succession of such operations is called “propagation“.
At the core of propagation we, therefore, find operations which we can express in the following form :
Input vector at N(m, L+1) : I(m, L+1) = SUM(over_all_k) [ W[(m, L+1), (k, L)] * O(k, L) ]
Output vector at N(m, L+1) : O(m, L+1) = f_act( I(m, L+1) )
To make things simpler: Let us set f_act = 1. Then we get
O(m, L+1) = SUM(over all k) [ W[(m, L+1), (k,N)] * O(k, L) ]
The SUM is basically a matrix operations on vector O(k, L). If output vectors were always given by singular values in “k” rows (!) then we could write
O(L+1) = W(L, L+1) * O(L) )
Batches and numerical optimization of the matrix operations for propagation
In the past I have programmed gas flow simulations in exploding stars in Fortran/C and simulated cascaded production networks for Supply Chains in PHP. There, you work with deep and complicated layer structures, too. Therefore I felt well prepared for writing a small program to set up a simple artificial neural network with some hidden layers.
Something, you will find in literature (e.g. in the book of S. Rashka, “Python machine Learning”, 2016, PACKT Publishing] are remarks on batch based training. Rashka e.g. rightfully claims in his discussion of an example code that the use of mini-batches of input data sets is advantageous (see chapter 12, page 379 in the book mentioned above):
“Mini-batch learning has the advantage over online learning that we can make use of our vectorized implementations to improve computational efficiency.”
I swallowed this remark first without much thinking. As a physicist I had used highly optimized libraries for Linear Algebra operation on supercomputers with vector registers already 35 years ago. Ok, numpy can perform a matrix operation on some vector very fast, especially on GPUs – so what?
But when I tried to follow the basic line of programming of
Rashka for an ANN in Python, I suddenly found that I did not fully understand what the code was doing whilst handling batches of input data sets. The reason was that I had not at all grasped the full implications of the quoted remark. It indicated not just the acceleration of a matrix operation on one input vector; no, instead, it indicated that we could and should perform matrix operations in parallel on all input vectors given by a full series of many different data sets in a mini-batch collection.
So, we then want to perform an optimized matrix operation on another matrix of at least 2 dimensions. When you think about it, you will quickly understand that any kind of operation of one n-dim matrix on another matrix with more and different dimensions must be defined very precisely to avoid misunderstandings and miscalculations.
To make things simple:
Let us take a (3,5)-matrix A and a (10,3)-matrix B. What should a “dot“-like operation A * B mean in this case? Would it be suitable at all, regarding the matrix element arrangements in rows and columns?
I studied the numpy documentation on “numpy.dot(A, B)”. See e.g. here. Ooops, not fully trivial: “If A is an N-D array and B is an M-D array (where M>=2), it is a sum product over the last axis of A and the second-to-last axis of B”. The matrices must obviously be arranged in a suitable way ….
So, I decided to performing some simple matrix experiments to get a clear understanding. I assumed a fictitious ANN of three layers and some direct propagation between the layers. What does propagation mean in such a case, how is it expressed in matrix operation terms in Python and what does batch-handling mean in this context?
Three simple layers and a batch of input data sets
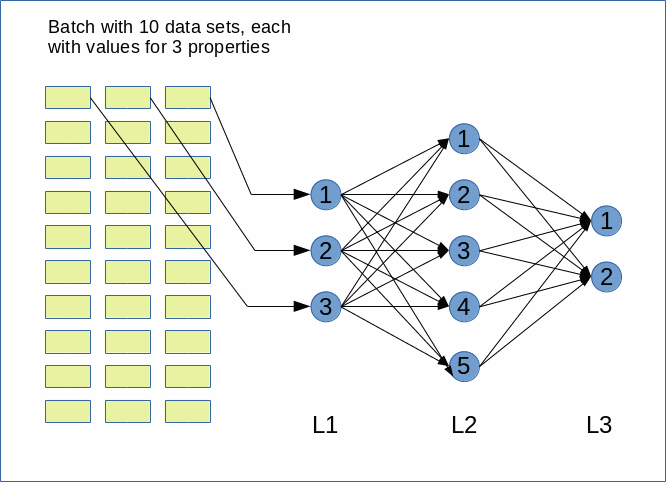
The following graphics shows my simple network:
Our batch has 10 data sets of test data for our ANN. Each data set describes 3 properties; thus the input layer “L1” has 3 nodes. Our “hidden” layer “L2” shall have 5 nodes and our output layer “L3” only 2 nodes.
Weight matrices and propagation operations
Now, let us define some simple matrices with input vectors for transportation and weight-matrices for the layer-connections with Python. All activation functions shall in our simple example just perform a multiplication by 1.
The input to layer “L1” – i.e. a batch of 10 data sets – shall be given by a matrix of 10 lines (dimension 1) and 3 columns (dimension 2)
import numpy as np
import scipy
import time
A1 = np.ones((10,3))
A1[:,1] *= 2
A1[:,2] *= 4
print("\nMatrix of input vectors to layer L1:\n")
print(A1)
print("\nMatrix of input vectors to layer 1:\n")
print(A1)
Matrix of input vectors to layer 1:
Matrix of input vectors to layer L1:
[[1. 2. 4.]
[1. 2. 4.]
[1. 2. 4.]
[1. 2. 4.]
[1. 2. 4.]
[1. 2. 4.]
[1. 2. 4.]
[1. 2. 4.]
[1. 2. 4.]
[1. 2. 4.]]
We now define the weight matrix “W1” for transport between layer L1 and layer L2 as:
W1= np.random.randint(1, 10, 5*3)
W1 = w1.reshape(5,3)
print("\nMatrix of weights W1(L1, L2) :\n")
print(W1)
Matrix of weights W1(L1, L2) :
[[3 8 5]
[4 9 7]
[4 1 3]
[8 8 4]
[3 1 9]]
We set the first dimension to the number of nodes in the target layer (L2) and the second dimension to
the number of nodes in the lower preceding layer (here: L1). Expressed in the shape notation for an array “(dimension 1, dimension 2, …) “:
Shape of W : (dim 1, dim 2) = (node number of target layer, node number of preceding layer)
If we now tried a numpy “dot”-operation for our matrices
A2 = np.dot(W1, A1)
we would get an error:
ValueError: shapes (5,3) and (10,3) not aligned: 3 (dim 1) != 10 (dim 0)
Well, this is actually consistent with the numpy documentation: The last dimension of “W1” should be consistent with the second to last (=first) dimension of “strong>A1“.
What we need to do here is to transpose the matrix of our input vectors for our multiple data sets:
A1 = np.transpose(A1)
print("\nMatrix of output vectors of layer 1 to layer 2:\n")
print(A1)
Matrix of output vectors of layer 1 to layer 2:
[[1. 1. 1. 1. 1. 1. 1. 1. 1. 1.]
[2. 2. 2. 2. 2. 2. 2. 2. 2. 2.]
[4. 4. 4. 4. 4. 4. 4. 4. 4. 4.]]
Then
A2 = np.dot(W1, A1)
print("\n\nA2:\n")
print(A2)
A2:
[[39. 39. 39. 39. 39. 39. 39. 39. 39. 39.]
[50. 50. 50. 50. 50. 50. 50. 50. 50. 50.]
[18. 18. 18. 18. 18. 18. 18. 18. 18. 18.]
[40. 40. 40. 40. 40. 40. 40. 40. 40. 40.]
[41. 41. 41. 41. 41. 41. 41. 41. 41. 41.]]
Now, we need a weight matrix W2 between layer L1 and layer L2:
W2= np.random.randint(1, 3, 2*5)
W2 = W2.reshape(2,5)
print("\n\nW2:\n")
print(W2)
W2:
[[2 2 1 2 2]
[2 2 2 2 1]]
We followed the same policy – and used the number of nodes in target layer L2 as the first dimension of the weight array.
Now, do we have to to anything with array A2, if we want to use it as an input for the next matrix operation ?
A3 = np.dot(W2, A2)
No, we do not need to change anything! “A2” has the required form, already. The second to last dimension in “A2” is 5 in our example – as is the last dimension in “W2“!
So, the following will work directly :
A3 = np.dot(W2, A2)
print("\n\nA3:\n")
print(A3)
A3:
[[358. 358. 358. 358. 358. 358. 358. 358. 358. 358.]
[335. 335. 335. 335. 335. 335. 335. 335. 335. 335.]]
You can try out our complete “propagation” code in a Jupyter notebook cell.
import numpy as np
import scipy
import time
A1 = np.ones((10,3))
A1[:,1] *= 2
A1[:,2] *= 4
print("\nMatrix of input vectors to layer L1:\n")
print(A1)
W1= np.random.randint(1, 10, 5*3)
W1 = W1.reshape(5,3)
print("\nMatrix of weights W1(L1, L2) :\n")
print(W1)
A1 = np.transpose(A1)
print("\nMatrix of output vectors of layer L1 to layer L2:\n")
print(A1)
A2 = np.dot(W1, A1)
print("\n\nA2:\n")
print(A2)
W2= np.random.randint(1, 3, 2*5)
W2 = W2.reshape(2,5)
print("\n\nW2:\n")
print(W2)
A3 = np.dot(W2, A2)
print("\n\nA3:\n")
print(A3)
Conclusion
The first thing we learned is that matrix operations on simple vectors can be extended to a set of vectors, i.e. a matrix. In the case of ANNs one dimension of such an (numpy) array would cover the number of data sets of a typical input mini-batch. The other dimension would cover the “properties” of the input data.
The second thing we saw is that the input matrix to an ANN often must be transposed to work together with weight matrices, if such matrices follow the policy that the first dimension is given by the number of cells in the target layer and the second dimension by the number of nodes of the
preceding layer.
The weighted input to the first target layer is then given by a matrix “dot”-operation on the transposed input matrix. The outcome is a matrix where the first dimension is defined by the number of nodes of the target layer and the second dimension the number of data sets. This has already the correct form for a further propagation to the next layer. Note that the application of an activation function to each of the matrix elements would not change the required arrangement of matrix data!
After the input layer we, therefore, can just continue to directly apply weight matrices to output matrices by a numpy-“dot”-operation. The weight matrix structure just should reflect our shape policy of
(dim1, dim2) = (node number of target layer, node number of preceding layer)
.
In the next articles we shall use these insights to build a Python class for the setup and training of a simple ANN with one or two hidden layers for the analysis of the famous MNIST dataset.