Welcome back, my friends of MLP coding. In the last article we gave the code developed in the article series
A simple Python program for an ANN to cover the MNIST dataset – I – a starting point
a first performance boost by two simple measures:
- We set all major arrays to the “numpy.float32” data type instead of the default “float64”.
- In addition we eliminated superfluous parts of the backward [BW] propagation between the first hidden layer and the input layer.
This brought us already down to around
11 secs for 35 epochs on the MNIST dataset, a batch-size of 500 and an accuracy around 99 % on the training set
This was close to what Keras (and TF2) delivered for the same batch size. It marks the baseline for further performance improvements of our MLP code.
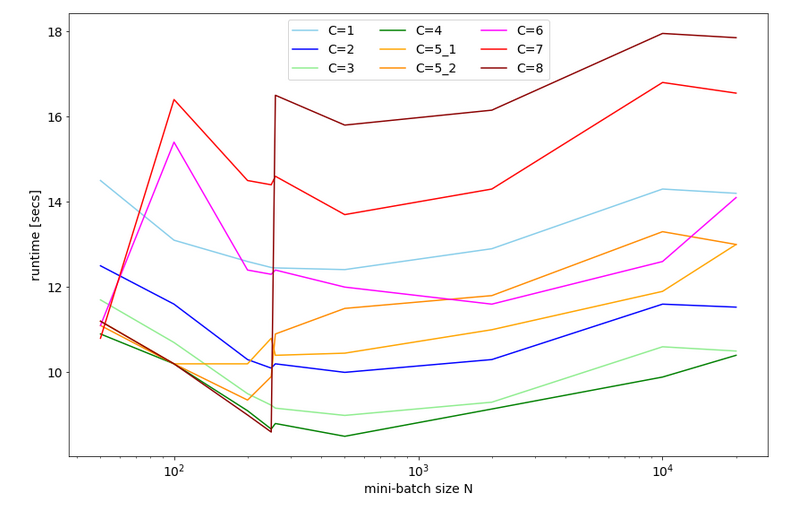
Can we get better than 11 secs for 35 epochs? The answer is: Yes, we can – but only in small steps. So, do not expect any gigantic performance leaps for the training loop itself. But, there was and is also our observation that there is no significant acceleration with growing batch sizes over 1000 – but with Keras we saw such an acceleration.
In this article I shall shortly discuss why we should care about big batch sizes – at least in combination with FW-propagation. Afterwards I want to draw your attention to a specific code segment of our MLP. We shall see that an astonishingly simple array operation dominates the CPU time of our straight forward coded FW propagation. Especially for big batch sizes!
Actually, it is an operation I would never have guessed to be such an an obstacle to efficiency if somebody had asked me. As a naive Python beginner I had to learn that the arrangement of arrays in the computer’s memory sometimes have an impact – especially when big arrays are involved. To get to this generally useful insight we will have to invest some effort into performance tests of some specific Numpy operations on arrays. The results give us some options for possible performance improvements; but in the end we shall circumvent the impediment all together.
The discussion will indicate that we should change our treatment of bias neurons fundamentally. We shall only go a preliminary step in this direction. This step will give us already a 15% improvement regarding the training time. But even more important, it will reward us with a significant improvement – by a factor > 2.5 – with respect to the FW-propagation of the complete training and test data sets, i.e. for the FW-propagation of “batches” with big sizes (60000 or 10000 samples).
“np.” abbreviates the “Numpy” library below. I shall sometimes speak of our 2-dimensional Numpy “arrays” as “matrices” in an almost synonymous way. See, however, one of the links at the bottom of the article for the subtle differences of related data types. For the time being we can safely ignore the mathematical differences between matrices, stacks of matrices and tensors. But we should have a clear understanding of the profound difference between the “*“-operation and the “np.dot()“-operation on 2-dimensional arrays.
Why are big batch sizes relevant?
There are several reasons why we should care about an efficient treatment of big batches. I name a few, only:
- Numpy operations on bigger matrices may become more efficient on systems with multiple CPUs, CPU cores or multiple GPUs.
- Big batch sizes together with a relatively small learning rate will lead to a smoother descent path on the cost hyperplane. Could become important in some intricate real life scenarios beyond MNIST.
- We should test the achieved accuracy on evaluation and test datasets during training. This data sets may have a much bigger size than the training batches.
The last point addresses the problem of overfitting: We may approach a minimum of the loss function of the training data set, but may leave the minimum of the cost function (and of related errors) of the test data set at some point. Therefore, we should check the accuracy on evaluation and test data sets already during the training phase. This requires the FW-propagation of such sets – preferably in one sweep. I.e. we talk about the propagation of really big batches with 10000 samples or more.
How do we measure the accuracy? Regarding the training set we gather averaged errors of batches during the training run and determine the related accuracy at the end of every printout period via an average over all batches: The average is taken over the absolute values of the difference between the sigmoidal output and the one-hot encoded target values of the batch samples. Note that this will give us slightly different values than tests where Numpy.argmax() is applied to the output first.
We can verify the accuracy also on the complete training and test data sets. Often we will do so after each and every epoch. Then we involve argmax(), by the way to get numbers in terms of correctly classified samples.
We saw that the forward [FW] propagation of the complete training data set “X_train” in one sweep requires a substantial (!) amount of CPU time in the present state of our code. When we perform such a test at each and every epoch on the training set the pure training time is prolonged by roughly a factor 1.75. As said: In real live scenarios we would rather or in addition perform full accuracy tests on prepared evaluation and test data sets – but they are big “batches” as well.
So, one relevant question is: Can we reduce the time required for a forward [FW] propagation of complete training and test data sets in one vectorized sweep?
Which operation dominates the CPU time of our present MLP forward propagation?
The present code for the FW-propagation of a mini-batch through my MLP comprises the following statements – enriched below by some lines to measure the required CPU-time:
''' -- Method to handle FW propagation for a mini-batch --'''
def _fw_propagation(self, li_Z_in, li_A_out):
'''
Parameter:
li_Z_in : list of input values at all layers - li_Z_in[0] is already filled -
other elemens to to be filled during FW-propagation
li_A_out: list of output values at all layers - to be filled during FW-propagation
'''
# index range for all layers
# Note that we count from 0 (0=>L0) to E L(=>E) /
# Careful: during BW-propagation we need a clear indexing of the lists filled during FW-propagation
ilayer = range(0, self._n_total_layers-1)
# propagation loop
# ***************
for il in ilayer:
# Step 1: Take input of last layer and apply activation function
# ******
ts=time.perf_counter()
if il == 0:
A_out_il = li_Z_in[il] # L0: activation function is identity !!!
else:
A_out_il = self._act_func( li_Z_in[il] ) # use defined activation function (e.g. sigmoid)
te=time.perf_counter(); ta = te - ts; print("\nta = ", ta, " shape = ", A_out_il.shape, " type = ", A_out_il.dtype, " A_out flags = ", A_out_il.flags)
# Step 2: Add bias node
# ******
ts=time.perf_counter()
A_out_il = self._
add_bias_neuron_to_layer(A_out_il, 'row')
li_A_out[il] = A_out_il
te=time.perf_counter(); tb = te - ts; print("tb = ", tb, " shape = ", A_out_il.shape, " type = ", A_out_il.dtype)
# Step 3: Propagate by matrix operation
# ******
ts=time.perf_counter()
Z_in_ilp1 = np.dot(self._li_w[il], A_out_il)
li_Z_in[il+1] = Z_in_ilp1
te=time.perf_counter(); tc = te - ts; print("tc = ", tc, " shape = ", li_Z_in[il+1].shape, " type = ", li_Z_in[il+1].dtype)
# treatment of the last layer
# ***************************
ts=time.perf_counter()
il = il + 1
A_out_il = self._out_func( li_Z_in[il] ) # use the defined output function (e.g. sigmoid)
li_A_out[il] = A_out_il
te=time.perf_counter(); tf = te - ts; print("\ntf = ", tf)
return None
The attentive reader notices that I also included statements to print out information about the shape and so called “flags” of the involved arrays.
I give you some typical CPU times for the MNIST dataset first. Characteristics of the test runs were:
- data were taken during the first two epochs;
- the batch-size was 10000; i.e. we processed 6 batches per epoch;
- “ta, tb, tc, tf” are representative data for a single batch comprising 10000 MNIST samples.
Averaged timing results for such batches are:
Layer L0 ta = 2.6999987312592566e-07 tb = 0.013209896002081223 tc = 0.004847299001994543 Layer L1 ta = 0.005858420001459308 tb = 0.0005839099976583384 tc = 0.00040631899901200086 Layer L2 ta = 0.0025550600003043655 tb = 0.00026626299950294197 tc = 0.00022965300013311207 Layer3 tf = 0.0008438359982392285
Such CPU time data vary of course a bit (2%) with the background activity on my machine and with the present batch, but the basic message remains the same. When I first saw it I could not believe it:
Adding a bias-neuron to the input layer obviously dominated the CPU-consumption during forward propagation. Not the matrix multiplication at the input layer L0!
I should add at this point that the problem increases with growing batch size! (We shall see this later in elementary test, too). This means that propagating the complete training or test dataset for accuracy check at each epoch will cost us an enormous amount of CPU time – as we have indeed seen in the last article. Performing a full propagation for an accuracy test at the end of each and every epoch increased the total CPU time roughly by a factor of 1.68 (19 sec vs. 11.33 secs for 35 epochs; see the last article).
Adding a row of constant input values of bias neurons
I first wanted to know, of course, whether my specific method of adding a bias neuron to the A-output matrix at each layer really was so expensive. My naive approach – following a suggestion in a book of S. Rashka, by the way – was:
def add_bias_neuron_to_layer(A, how='column'):
if how == 'column':
A_new = np.ones((A.shape[0], A.shape[1]+1), dtype=np.float32)
A_new[:, 1:] = A
elif how == 'row':
A_new = np.ones((A.shape[0]+1, A.shape[1]), dtype=np.float32)
A_new[1:, :] = A
return A_new
What we do here is to create a new array which is bigger by one row and fit the original array into it. Seemed to be a clever approach at the time of coding (and actually it is faster than using np.vstack or np.hstack). The operation is different from directly adding a row to the existing input array explicitly, but it still requires a lot of row operations.
As we have seen I call this function in “_fw_
propagation()” by
A_out_il = self._add_bias_neuron_to_layer(A_out_il, 'row')
“A_out_il” is the transposition of a slice of the original X_train array. The slice in our test case for MNIST had a shape of (10000, 784).
This means that we talk about a matrix with shape (784, 10000) in the case of the MNIST dataset before adding the bias neuron and a shape of (785, 10000) after. I.e. we add a row with 10000 constant entries at the beginning of our transposed slice. Note also that the function returns a new array in memory.
Thus, our approach contains two possibly costly operations. Why did we do such a strange thing in the first place?
Well, when we coded the MLP it seemed to be a good idea to include the fact that we have bias neurons directly in the definition of the weight matrices and their shapes. So, we need(ed) to fit our input matrices at the layers to the defined shape of the weight matrices. As we see it now, this is a questionable strategy regarding performance. But, well, let us not attack something at the very center of the MLP code for all layers (except the output layer) at this point in time. We shall do this in a forthcoming article.
A factor of 3 ??
To understand my performance problem a bit better, I did the following test in a Jupyter cell:
''' Method to add values for a bias neuron to A_out all with C-cont. arrays '''
def add_bias_neuron_to_layer_C(A, how='column'):
if how == 'column':
A_new = np.ones((A.shape[0], A.shape[1]+1), dtype=np.float32)
A_new[:, 1:] = A
elif how == 'row':
A_new = np.ones((A.shape[0]+1, A.shape[1]), dtype=np.float32)
A_new[1:, :] = A
return A_new
input_shape =(784, 10000)
ay_inpC = np.array(np.random.random_sample(input_shape)*2.0, dtype=np.float32)
tx = time.perf_counter()
ay_inpCb = add_bias_neuron_to_layer_C(ay_inpC, 'row')
li_A.append(ay_inpCb)
ty = time.perf_counter(); t_biasC = ty - tx;
print("\n bias time = ", "%10.8f"%t_biasC)
print("shape_biased = ", ay_inpCb.shape)
to get:
bias time = 0.00423444
Same batch-size, but substantially faster – by roughly a factor of 3! – compared to what my MLP code delivered. Actually the timing data varied a bit between 0.038 and 0.045 (with an average at 0.0042) when repeating the run. To exclude any problems with calling the function from within a Python class I repeated the same test inside the class “MyANN” during FW-propagation – with the same result (as it should be; see the first link at the end of this article).
So: Applying one and the same function on a randomly filled array was much faster than applying it on my Numpy (input) array “A_out_il” (with the same shape). ????
C- and F-contiguous arrays
It took me a while to find the reason: “A_out_il” is the result of a matrix transposition. In Numpy this corresponds to a certain view on the original array data – but this still has major consequences for the handling of the data:
A 2 dimensional array or matrix is an ordered addressable sequence of data in the computer’s memory. Now, if you yourself had to program an array representation in memory on a basic level you would – due to performance reasons – make a choice whether you arrange data row-wise or column-wise. And you would program functions for array-operations with your chosen “order” in mind!
Actually, if you google a bit you find that the two ways of arranging array or matrix data are both well established. In connection with Numpy we speak of either a C-contiguous order or a F-contiguous order of the array data. In the first case (C) data are stored and addressed row by row and can be read efficiently this way, in the other (F) case data are arranged
column by column. By the way: The “C” refers to the C-language, the “F” to Fortran.
On a Linux system Numpy normally creates and operates with C-contiguous arrays – except when you ask Numpy explicitly to work differently. Quite many array related functions, therefore, have a parameter “order”, which you can set to either ‘C’ or ‘F’.
Now, let us assume that we have a C-contiguous array. What happens when we transpose it – or look at it in a transposed way? Well, logically it then becomes F-contiguous! Then our “A_out_il” would be seen as F-contiguous. Could this in turn have an impact on performance? Well, I create “A_out_il” in method “_handle_mini_batch()” of my MyANN-class via
# Step 0: List of indices for data records in the present mini-batch
# ******
ay_idx_batch = self._ay_mini_batches[num_batch]
# Step 1: Special preparation of the Z-input to the MLP's input Layer L0
# ******
# Layer L0: Fill in the input vector for the ANN's input layer L0
li_Z_in_layer[0] = self._X_train[ay_idx_batch] # numpy arrays can be indexed by an array of integers
li_Z_in_layer[0] = li_Z_in_layer[0].T
...
...
Hm, pretty simple. But then, what happens if we perform our rather special adding of the bias-neuron row-wise, as we logically are forced to? Remember, the array originally had a shape of (10000, 784) and after transposing a shape of (784, 10000), i.e. the columns then represent the samples of the mini-batch. Well, instead of inserting a row of 10000 data contiguously into memory in one swipe into a C-contiguous array we must hop to the end of each contiguous column of the F-contiguous array “A_out_il” in memory and add one element there. Even if you would optimize it there are many more addresses and steps involved. Can’t become efficient ….
How can we see, which order an array or view onto it follows? We just have to print its “flags“. And I indeed got:
flags li_Z_in[0] = C_CONTIGUOUS : False F_CONTIGUOUS : True OWNDATA : False WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False
Additional tests with Jupyter
Let us extend the tests of our function in the Jupyter cell in the following way to cover a variety of options related to our method of adding bias neurons:
# The bias neuron problem
# ************************
import numpy as np
import scipy
from scipy.special import expit
import time
''' Method to add values for a bias neuron to A_out - by creating a new C-cont. array '''
def add_bias_neuron_to_layer_C(A, how='column'):
if how == 'column':
A_new = np.ones((A.shape[0], A.shape[1]+1), dtype=np.float32)
A_new[:, 1:] = A
elif how == 'row':
A_new = np.ones((A.shape[0]+1, A.shape[1]), dtype=np.float32)
A_new[1:, :] = A
return A_new
''' Method to add values for a bias neuron to A_out - by creating a new F-cont. array '''
def add_bias_neuron_to_layer_F(A, how='column'):
if how == 'column':
A_new = np.ones((A.shape[0], A.shape[1]+1), order='F', dtype=np.float32)
A_new[:, 1:] = A
elif how == 'row':
A_new = np.ones((A.shape[0]+1, A.shape[1]), order='F', dtype=np.float32)
A_new[1:, :] = A
return A_new
rg_j = range(50)
li_A = []
t_1 = 0.0; t_2 = 0.0;
t_3 = 0.0; t_4 = 0.0;
t_5 = 0.0; t_6 = 0.0;
t_7 = 0.0; t_8 = 0.0;
# two types of input shapes
input_shape1 =(784, 10000)
input_shape2 =(10000, 784)
for j in rg_j:
# For test 1: C-cont. array with shape (784, 10000)
# in a MLP programm delivering X_train as (
10000, 784) we would have to (re-)create it
# explicitly with the C-order (np.copy or np.asarray)
ay_inpC = np.array(np.random.random_sample(input_shape1)*2.0, order='C', dtype=np.float32)
# For test 2: C-cont. array with shape (10000, 784) as it typically is given by a slice of the
# original X_train
ay_inpC2 = np.array(np.random.random_sample(input_shape2)*2.0, order='C', dtype=np.float32)
# For tests 3 and 4: transposition - this corresponds to the MLP code
ay_inpF = ay_inpC2.T
# For test 5: The original X_train or mini-batch data are somehow given in F-cont.form,
# then inpF3 below would hopefully be in C-cont. form
ay_inpF2 = np.array(np.random.random_sample(input_shape2)*2.0, order='F', dtype=np.float32)
# For test 6
ay_inpF3 = ay_inpF2.T
# Test 1: C-cont. input to add_bias_neuron_to_layer_C - with a shape that fits already
# ******
tx = time.perf_counter()
ay_Cb = add_bias_neuron_to_layer_C(ay_inpC, 'row')
li_A.append(ay_Cb)
ty = time.perf_counter(); t_1 += ty - tx;
# Test 2: Standard C-cont. input to add_bias_neuron_to_layer_C - but col.-operation due to shape
# ******
tx = time.perf_counter()
ay_C2b = add_bias_neuron_to_layer_C(ay_inpC2, 'column')
li_A.append(ay_C2b)
ty = time.perf_counter(); t_2 += ty - tx;
# Test 3: F-cont. input to add_bias_neuron_to_layer_C (!) - but row-operation due to shape
# ****** will give us a C-cont. output array which later is used in np.dot() on the left side
tx = time.perf_counter()
ay_C3b = add_bias_neuron_to_layer_C(ay_inpF, 'row')
li_A.append(ay_C3b)
ty = time.perf_counter(); t_3 += ty - tx;
# Test 4: F-cont. input to add_bias_neuron_to_layer_F (!) - but row-operation due to shape
# ****** will give us a F-cont. output array which later is used in np.dot() on the left side
tx = time.perf_counter()
ay_F4b = add_bias_neuron_to_layer_F(ay_inpF, 'row')
li_A.append(ay_F4b)
ty = time.perf_counter(); t_4 += ty - tx;
# Test 5: F-cont. input to add_bias_neuron_to_layer_F (!) - but col-operation due to shape
# ****** will give us a F-cont. output array with wrong shape for weight matrix
tx = time.perf_counter()
ay_F5b = add_bias_neuron_to_layer_F(ay_inpF2, 'column')
li_A.append(ay_F5b)
ty = time.perf_counter(); t_5 += ty - tx;
# Test 6: C-cont. input to add_bias_neuron_to_layer_C (!) - row-operation due to shape
# ****** will give us a C-cont. output array with wrong shape for weight matrix
tx = time.perf_counter()
ay_C6b = add_bias_neuron_to_layer_C(ay_inpF3, 'row')
li_A.append(ay_C6b)
ty = time.perf_counter(); t_6 += ty - tx;
# Test 7: C-cont. input to add_bias_neuron_to_layer_F (!) - row-operation due to shape
# ****** will give us a F-cont. output array with wrong shape for weight matrix
tx = time.perf_counter()
ay_F7b = add_bias_neuron_to_layer_F(ay_inpC2, 'column')
li_A.append(ay_F7b)
ty = time.perf_counter(); t_7 += ty - tx;
print("\nTest 1: nbias time C-cont./row with add_.._C() => ", "%10.8f"%t_1)
print("shape_ay_Cb = ", ay_Cb.shape, " flags = \n", ay_Cb.flags)
print("\nTest 2: nbias time C-cont./col with add_.._C() => ", "%10.8f"%t_2)
print("shape of ay_C2b = ", ay_C2b.shape, " flags = \n", ay_C2b.flags)
print("\nTest 3: nbias time F-cont./row with add_.._C() => ", "%10.8f"%t_3)
print("shape of ay_C3b = ", ay_C3b.shape, " flags = \n", ay_C3b.flags)
print("\nTest 4: nbias time F-cont./row with add_.._F() => ", "%10.8f"%t_4)
print("shape of ay_F4b = ", ay_F4b.shape, " flags = \n", ay_F4b.flags)
print("\nTest 5: nbias time F-cont./col
with add_.._F() => ", "%10.8f"%t_5)
print("shape of ay_F5b = ", ay_F5b.shape, " flags = \n", ay_F5b.flags)
print("\nTest 6: nbias time C-cont./row with add_.._C() => ", "%10.8f"%t_6)
print("shape of ay_C6b = ", ay_C6b.shape, " flags = \n", ay_C6b.flags)
print("\nTest 7: nbias time C-cont./col with add_.._F() => ", "%10.8f"%t_7)
print("shape of ay_F7b = ", ay_F7b.shape, " flags = \n", ay_F7b.flags)
You noticed that I defined two different ways of creating the bigger array into which we place the original one.
Results are:
Test 1: bias time C-cont./row with add_.._C() => 0.20854935 shape_ay_Cb = (785, 10000) flags = C_CONTIGUOUS : True F_CONTIGUOUS : False OWNDATA : True WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False Test 2: bias time C-cont./col with add_.._C() => 0.25661559 shape of ay_C2b = (10000, 785) flags = C_CONTIGUOUS : True F_CONTIGUOUS : False OWNDATA : True WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False Test 3: bias time F-cont./row with add_.._C() => 0.67718296 shape of ay_C3b = (785, 10000) flags = C_CONTIGUOUS : True F_CONTIGUOUS : False OWNDATA : True WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False Test 4: nbias time F-cont./row with add_.._F() => 0.25958392 shape of ay_F4b = (785, 10000) flags = C_CONTIGUOUS : False F_CONTIGUOUS : True OWNDATA : True WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False Test 5: nbias time F-cont./col with add_.._F() => 0.20990409 shape of ay_F5b = (10000, 785) flags = C_CONTIGUOUS : False F_CONTIGUOUS : True OWNDATA : True WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False Test 6: nbias time C-cont./row with add_.._C() => 0.22129941 shape of ay_C6b = (785, 10000) flags = C_CONTIGUOUS : True F_CONTIGUOUS : False OWNDATA : True WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False Test 7: nbias time C-cont./col with add_.._F() => 0.67642328 shape of ay_F7b = (10000, 785) flags = C_CONTIGUOUS : False F_CONTIGUOUS : True OWNDATA : True WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False
These results
- confirm that it is a bad idea to place a F-contiguous array or (F-contiguous view on an array) into a C-contiguous one the way we presently do it;
- confirm that we should at least create the surrounding array with the same order as the input array, which we place into it.
The best combinations are
- either to put an original C-contiguous array with fitting shape into a C-contiguous one with one more row,
- or to place an original F-contiguous array with suitable shape into a F-contiguous one with one more column.
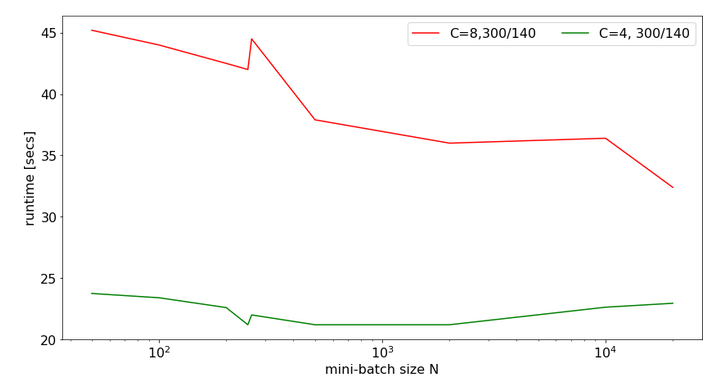
By the way: Some systematic tests also showed that the time difference between the first and the third operation grows with batch size:
bs = 60000, rep. = 30 => t1=0.70, t3=2.91, fact=4.16 bs = 50000, rep. = 30 => t1=0.58, t3=2.34, fact=4.03 bs = 40000, rep. = 50 => t1=0.78, t3=3.07, fact=3.91 bs = 30000, rep. = 50 => t1=0.60, t3=2.21, fact=3.68 bs = 20000, rep. = 60 => t1=0.49, t3=1.63, fact=3.35 bs = 10000, rep. = 60 => t1=0.26, t3=0.82, fact=3.20 bs = 5000, rep. = 60 => t1=0. 11, t3=0.35, fact=3.24 bs = 2000, rep. = 60 => t1=0.04, t3=0.10, fact=2.41 bs = 1000, rep. = 200 => t1=0.17, t3=0.38, fact=2.21 bs = 500, rep. = 1000 => t1=0.15, t3=0.32, fact=2.17 bs = 500, rep. = 200 => t1=0.03, t3=0.06, fact=2.15 bs = 100, rep. = 1500 => t1=0.04, t3=0.07, fact=1.92
“rep” is the loop range (repetition), “fact” is the factor between the fastest operation (test1: C-cont. into C-cont.) and the slowest (test3: F-cont. into C-cont). (The best results were selected among multiple runs with different repetitions for the table above).
We clearly see that our problem gets worse with batch sizes above bs=1000!
Problems with shuffling?
Okay, let us assume we wanted to go either of the 2 optimization paths indicated above. Then we would need to prepare the input array in a suitable form. But, how does such an approach fit to the present initialization of the input data and the shuffling of “X_train” at the beginning of each epoch?
If we keep up our policy of adding a bias neuron to the input layer by the mechanism we use we either have to get the transposed view into C-contiguous form or at least create the new array (including the row) in F-contiguous form. (The latter will not hamper the later np.dot()-multiplication with the weight-matrix as we shall see below.) Or we must circumvent the bias neuron problem at the input layer in a different way.
Actually, there are two fast shuffling options – and both are designed to work efficiently with rows, only. Another point is that the result is always C-contiguous. Let us look at some tests:
# Shuffling
# **********
dim1 = 60000
input_shapeX =(dim1, 784)
input_shapeY =(dim1, )
ay_X = np.array(np.random.random_sample(input_shapeX)*2.0, order='C', dtype=np.float32)
ay_Y = np.array(np.random.random_sample(input_shapeY)*2.0, order='C', dtype=np.float32)
ay_X2 = np.array(np.random.random_sample(input_shapeX)*2.0, order='C', dtype=np.float32)
ay_Y2 = np.array(np.random.random_sample(input_shapeY)*2.0, order='C', dtype=np.float32)
# Test 1: Shuffling of C-cont. array by np.random.shuffle
tx = time.perf_counter()
np.random.shuffle(ay_X)
np.random.shuffle(ay_Y)
ty = time.perf_counter(); t_1 = ty - tx;
print("\nShuffle Test 1: time C-cont. => t = ", "%10.8f"%t_1)
print("shape of ay_X = ", ay_X.shape, " flags = \n", ay_X.flags)
print("shape of ay_Y = ", ay_Y.shape, " flags = \n", ay_Y.flags)
# Test 2: Shuffling of C-cont. array by random index permutation
# as we have coded it for the beginning of each epoch
tx = time.perf_counter()
shuffled_index = np.random.permutation(dim1)
ay_X2, ay_Y2 = ay_X2[shuffled_index], ay_Y2[shuffled_index]
ty = time.perf_counter(); t_2 = ty - tx;
print("\nShuffle Test 2: time C-cont. => t = ", "%10.8f"%t_2)
print("shape of ay_X2 = ", ay_X2.shape, " flags = \n", ay_X2.flags)
print("shape of ay_Y2 = ", ay_Y2.shape, " flags = \n", ay_Y2.flags)
# Test3 : Copy Time for writing the whole X-array into 'F' ordered form
# such that slices transposed get C-order
ay_X3x = np.array(np.random.random_sample(input_shapeX)*2.0, order='C', dtype=np.float32)
tx = time.perf_counter()
ay_X3 = np.copy(ay_X3x, order='F')
ty = time.perf_counter(); t_3 = ty - tx;
print("\nTest 3: time to copy to F-cont. array => t = ", "%10.8f"%t_3)
print("shape of ay_X3 = ", ay_X3.shape, " flags = \n", ay_X3.flags)
# Test4 - shuffling of rows in F-cont. array => The result is C-contiguous!
tx = time.perf_counter()
shuffled_index = np.random.permutation(dim1)
ay_X3, ay_Y2 = ay_X3[shuffled_index], ay_Y2[shuffled_index]
ty = time.perf_counter(); t_4 = ty - tx;
print("\nTest 4: Shuffle rows of F-
cont. array => t = ", "%10.8f"%t_4)
print("shape of ay_X3 = ", ay_X3.shape, " flags = \n", ay_X3.flags)
# Test 5 - transposing and copying after => F-contiguous with changed shape
tx = time.perf_counter()
ay_X5 = np.copy(ay_X.T)
ty = time.perf_counter(); t_5 = ty - tx;
print("\nCopy Test 5: time copy to F-cont. => t = ", "%10.8f"%t_5)
print("shape of ay_X5 = ", ay_X5.shape, " flags = \n", ay_X5.flags)
# Test 6: shuffling columns in F-cont. array
tx = time.perf_counter()
shuffled_index = np.random.permutation(dim1)
ay_X6 = (ay_X5.T[shuffled_index]).T
ty = time.perf_counter(); t_6 = ty - tx;
print("\nCopy Test 6: shuffling F-cont. array in columns => t = ", "%10.8f"%t_6)
print("shape of ay_X6 = ", ay_X6.shape, " flags = \n", ay_X6.flags)
Results are:
Shuffle Test 1: time C-cont. => t = 0.08650427 shape of ay_X = (60000, 784) flags = C_CONTIGUOUS : True F_CONTIGUOUS : False OWNDATA : True WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False shape of ay_Y = (60000,) flags = C_CONTIGUOUS : True F_CONTIGUOUS : True OWNDATA : True WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False Shuffle Test 2: time C-cont. => t = 0.02296818 shape of ay_X2 = (60000, 784) flags = C_CONTIGUOUS : True F_CONTIGUOUS : False OWNDATA : True WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False shape of ay_Y2 = (60000,) flags = C_CONTIGUOUS : True F_CONTIGUOUS : True OWNDATA : True WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False Test 3: time to copy to F-cont. array => t = 0.09333340 shape of ay_X3 = (60000, 784) flags = C_CONTIGUOUS : False F_CONTIGUOUS : True OWNDATA : True WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False Test 4: Shuffle rows of F-cont. array => t = 0.25790425 shape of ay_X3 = (60000, 784) flags = C_CONTIGUOUS : True F_CONTIGUOUS : False OWNDATA : True WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False Copy Test 5: time copy to F-cont. => t = 0.02146052 shape of ay_X5 = (784, 60000) flags = C_CONTIGUOUS : False F_CONTIGUOUS : True OWNDATA : True WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False Copy Test 6: shuffling F-cont. array in columns by using the transposed view => t = 0.02402249 shape of ay_X6 = (784, 60000) flags = C_CONTIGUOUS : False F_CONTIGUOUS : True OWNDATA : False WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False
The results reveal three points:
- Applying a random permutation of an index is faster than using np.random.shuffle() on the array.
- The result is C-contiguous in both cases.
- Shuffling of columns can be done in a fast way by shuffling rows of the transposed array.
So, at the beginning of each epoch we are in any case confronted with a C-contiguous array of shape (batch_size, 784). Comparing this with the test data further above seems to leave us with three choices:
- Approach 1: At the beginning of each epoch we copy the input array into a F-contiguous one, such that the required transposed array afterwards is C-contiguous and our present version of “_add_bias_neuron_to_layer()” works fast with adding a row of bias nodes. The result
would be a C-contiguous array with shape (785, size_batch). - Approach 2: We define a new method “_add_bias_neuron_to_layer_F()” which creates an F-contiguous array with an extra row into which we fit the existing (transposed) array “A_out_il”. The result would be a F-contiguous array with shape (785, size_batch).
- Approach 3: We skip adding a row for bias neurons altogether.
The first method has the disadvantage that the copy-process requires time itself at the beginning of each epoch. But according to the test data the total gain would be bigger than the loss (6 batches!). The second approach has a small disadvantage because “_add_bias_neuron_to_layer_F()” is slightly slower than its row oriented counterpart – but this will be compensated by a slightly faster matrix dot()-multiplication. All in all the second option seems to be the better one – in case we do not find a completely different approach. Just wait a minute …
Intermezzo: Matrix multiplication np.dot() applied to C- and/or F-contiguous arrays
As we have come so far: How does np.dot() react to C- or F-contiguous arrays? The first two optimization approaches would end in different situations regarding the matrix multiplication. Let us cover all 4 possible combinations by some test:
# A simple test on np.dot() on C-contiguous and F-contiguous matrices
# *******************************************************
# Is the dot() multiplication fasterfor certain combinations of C- and F-contiguous matrices?
input_shape =(800, 20000)
ay_inpC1 = np.array(np.random.random_sample(input_shape)*2.0, dtype=np.float32 )
#print("shape of ay_inpC1 = ", ay_inpC1.shape, " flags = ", ay_inpC1.flags)
ay_inpC2 = np.array(np.random.random_sample(input_shape)*2.0, dtype=np.float32 )
#print("shape of ay_inpC2 = ", ay_inpC2.shape, " flags = ", ay_inpC2.flags)
ay_inpC3 = np.array(np.random.random_sample(input_shape)*2.0, dtype=np.float32 )
print("shape of ay_inpC3 = ", ay_inpC3.shape, " flags = ", ay_inpC3.flags)
ay_inpF1 = np.copy(ay_inpC1, order='F')
ay_inpF2 = np.copy(ay_inpC2, order='F')
ay_inpF3 = np.copy(ay_inpC3, order='F')
print("shape of ay_inpF3 = ", ay_inpF3.shape, " flags = ", ay_inpF3.flags)
weight_shape =(101, 800)
weightC = np.array(np.random.random_sample(weight_shape)*0.5, dtype=np.float32)
print("shape of weightC = ", weightC.shape, " flags = ", weightC.flags)
weightF = np.copy(weightC, order='F')
print("shape of weightF = ", weightF.shape, " flags = ", weightF.flags)
rg_j = range(300)
ts = time.perf_counter()
for j in rg_j:
resCC1 = np.dot(weightC, ay_inpC1)
resCC2 = np.dot(weightC, ay_inpC2)
resCC3 = np.dot(weightC, ay_inpC3)
resCC1 = np.dot(weightC, ay_inpC1)
resCC2 = np.dot(weightC, ay_inpC2)
resCC3 = np.dot(weightC, ay_inpC3)
te = time.perf_counter(); tcc = te - ts; print("\n dot tCC time = ", "%10.8f"%tcc)
ts = time.perf_counter()
for j in rg_j:
resCF1 = np.dot(weightC, ay_inpF1)
resCF2 = np.dot(weightC, ay_inpF2)
resCF3 = np.dot(weightC, ay_inpF3)
resCF1 = np.dot(weightC, ay_inpF1)
resCF2 = np.dot(weightC, ay_inpF2)
resCF3 = np.dot(weightC, ay_inpF3)
te = time.perf_counter(); tcf = te - ts; print("\n dot tCF time = ", "%10.8f"%tcf)
ts = time.perf_counter()
for j in rg_j:
resF1 = np.dot(weightF, ay_inpC1)
resF2 = np.dot(weightF, ay_inpC2)
resF3 = np.dot(weightF, ay_inpC3)
resF1 = np.dot(weightF, ay_inpC1)
resF2 = np.dot(weightF, ay_inpC2)
resF3 = np.dot(weightF, ay_inpC3)
te = time.perf_counter(); tfc = te - ts; print("\n dot tFC time = ", "%10.8f"%tfc)
ts = time.
perf_counter()
for j in rg_j:
resF1 = np.dot(weightF, ay_inpF1)
resF2 = np.dot(weightF, ay_inpF2)
resF3 = np.dot(weightF, ay_inpF3)
resF1 = np.dot(weightF, ay_inpF1)
resF2 = np.dot(weightF, ay_inpF2)
resF3 = np.dot(weightF, ay_inpF3)
te = time.perf_counter(); tff = te - ts; print("\n dot tFF time = ", "%10.8f"%tff)
The results show some differences – but they are relatively small:
shape of ay_inpC3 = (800, 20000) flags = C_CONTIGUOUS : True F_CONTIGUOUS : False OWNDATA : True WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False shape of ay_inpF3 = (800, 20000) flags = C_CONTIGUOUS : False F_CONTIGUOUS : True OWNDATA : True WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False shape of weightC = (101, 800) flags = C_CONTIGUOUS : True F_CONTIGUOUS : False OWNDATA : True WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False shape of weightF = (101, 800) flags = C_CONTIGUOUS : False F_CONTIGUOUS : True OWNDATA : True WRITEABLE : True ALIGNED : True WRITEBACKIFCOPY : False UPDATEIFCOPY : False dot tCC time = 21.77729867 dot tCF time = 20.68745600 dot tFC time = 21.42704156 dot tFF time = 20.65543837
Actually, most of the tiny differences comes from putting the matrix into a fitting order. This is something Numpy.dot() performs automatically; see the documentation. The matrix operation is fastest for the second matrix being in F-order, but the difference is nothing to worry about at our present discussion level.
Avoiding the bias problem at the input layer
We could now apply one of the two strategies to improve our mechanism of dealing with the bias nodes at the input layer. You would notice a significant acceleration there. But you leave the other layers unchanged. Why?
The reason is quite simple: The matrix multiplications with the weight matrix – done by “np.dot()” – produces the C-contiguous arrays at later layers with the required shapes! E.g., an input array at layer L1 of the suitable shape (70, 10000). So, we can for the moment leave everything at the hidden layers and at the output layer untouched.
However, the discussion above made one thing clear: The whole approach of how we technically treat bias nodes is to be criticized. Can we at least go another way at the input layer?
Yes, we can. Without touching the weight matrix connecting the layers L0 and L1. We need to get rid of unnecessary or inefficient operations in the training loop, but we can afford some bigger operations during the setup of the input data. What, if we added the required bias values already to the input data array?
This would require a column operation on a transposition of the whole dataset “X”. But, we need to perform this operation only once – and before splitting the data set into training and test sets! As a MLP generally works with flattened data such an approach should work for other datasets, too.
Measurements show that adding a bias column will cost us between 0.030 and 0.035 secs. A worthy one time investment! Think about it: We would not need to touch our already fast methods of shuffling and slicing to get the batches – and even the transposed matrix would already have the preferred F-contiguous order for np.dot()! The required code changes are minimal; we just need to adapt our methods “_handle_input_data()” and “_fw_propagation()” by two, three lines:
''' -- Method to handle different types of input data sets
Currently only
different MNIST sets are supported
We can also IMPORT a preprocessed MIST data set --'''
def _handle_input_data(self):
'''
Method to deal with the input data:
- check if we have a known data set ("mnist" so far)
- reshape as required
- analyze dimensions and extract the feature dimension(s)
'''
# check for known dataset
try:
if (self._my_data_set not in self._input_data_sets ):
raise ValueError
except ValueError:
print("The requested input data" + self._my_data_set + " is not known!" )
sys.exit()
# MNIST datasets
# **************
# handle the mnist original dataset - is not supported any more
if ( self._my_data_set == "mnist"):
mnist = fetch_mldata('MNIST original')
self._X, self._y = mnist["data"], mnist["target"]
print("Input data for dataset " + self._my_data_set + " : \n" + "Original shape of X = " + str(self._X.shape) +
# "\n" + "Original shape of y = " + str(self._y.shape))
#
# handle the mnist_784 dataset
if ( self._my_data_set == "mnist_784"):
mnist2 = fetch_openml('mnist_784', version=1, cache=True, data_home='~/scikit_learn_data')
self._X, self._y = mnist2["data"], mnist2["target"]
print ("data fetched")
# the target categories are given as strings not integers
self._y = np.array([int(i) for i in self._y], dtype=np.float32)
print ("data modified")
print("Input data for dataset " + self._my_data_set + " : \n" + "Original shape of X = " + str(self._X.shape) +
"\n" + "Original shape of y = " + str(self._y.shape))
# handle the mnist_keras dataset - PREFERRED
if ( self._my_data_set == "mnist_keras"):
(X_train, y_train), (X_test, y_test) = kmnist.load_data()
len_train = X_train.shape[0]
len_test = X_test.shape[0]
X_train = X_train.reshape(len_train, 28*28)
X_test = X_test.reshape(len_test, 28*28)
# Concatenation required due to possible later normalization of all data
self._X = np.concatenate((X_train, X_test), axis=0)
self._y = np.concatenate((y_train, y_test), axis=0)
print("Input data for dataset " + self._my_data_set + " : \n" + "Original shape of X = " + str(self._X.shape) +
"\n" + "Original shape of y = " + str(self._y.shape))
#
# common MNIST handling
if ( self._my_data_set == "mnist" or self._my_data_set == "mnist_784" or self._my_data_set == "mnist_keras" ):
self._common_handling_of_mnist()
# handle IMPORTED MNIST datasets (could in later versions also be used for other dtaasets
# **************************+++++
# Note: Imported sets are e.g. useful for testing some new preprocessing in a Jupyter environment before implementing related new methods
if ( self._my_data_set == "imported"):
if (self._X_import is not None) and (self._y_import is not None):
self._X = self._X_import
self._y = self._y_import
else:
print("Shall handle imported datasets - but they are not defined")
sys.exit()
#
# number of total records in X, y
self._dim_X = self._X.shape[0]
# ************************
# Common dataset handling
# ************************
# transform to 32 bit
# ~~~~~~~~~~~~~~~~~~~~
self._X = self._X.astype(np.
float32)
self._y = self._y.astype(np.int32)
# Give control to preprocessing - Note: preproc. includes also normalization
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
self._preprocess_input_data() # scaling, PCA, cluster detection ....
# ADDING A COLUMN FOR BIAS NEURONS
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
self._X = self._add_bias_neuron_to_layer(self._X, 'column')
print("type of self._X = ", self._X.dtype, " flags = ", self._X.flags)
print("type of self._y = ", self._y.dtype)
# mixing the training indices - MUST happen BEFORE encoding
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
shuffled_index = np.random.permutation(self._dim_X)
self._X, self._y = self._X[shuffled_index], self._y[shuffled_index]
# Splitting into training and test datasets
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
if self._num_test_records > 0.25 * self._dim_X:
print("\nNumber of test records bigger than 25% of available data. Too big, we stop." )
sys.exit()
else:
num_sep = self._dim_X - self._num_test_records
self._X_train, self._X_test, self._y_train, self._y_test = self._X[:num_sep], self._X[num_sep:], self._y[:num_sep], self._y[num_sep:]
# numbers, dimensions
# *********************
self._dim_sets = self._y_train.shape[0]
self._dim_features = self._X_train.shape[1]
print("\nFinal dimensions of training and test datasets of type " + self._my_data_set +
" : \n" + "Shape of X_train = " + str(self._X_train.shape) +
"\n" + "Shape of y_train = " + str(self._y_train.shape) +
"\n" + "Shape of X_test = " + str(self._X_test.shape) +
"\n" + "Shape of y_test = " + str(self._y_test.shape)
)
print("\nWe have " + str(self._dim_sets) + " data records for training")
print("Feature dimension is " + str(self._dim_features))
# Encode the y-target labels = categories // MUST happen AFTER encoding
# **************************
self._get_num_labels()
self._encode_all_y_labels(self._b_print_test_data)
#
return None
.....
.....
''' -- Method to handle FW propagation for a mini-batch --'''
def _fw_propagation(self, li_Z_in, li_A_out):
'''
Parameter:
li_Z_in : list of input values at all layers - li_Z_in[0] is already filled -
other elements of this list are to be filled during FW-propagation
li_A_out: list of output values at all layers - to be filled during FW-propagation
'''
# index range for all layers
# Note that we count from 0 (0=>L0) to E L(=>E) /
# Careful: during BW-propagation we need a clear indexing of the lists filled during FW-propagation
ilayer = range(0, self._n_total_layers-1)
# do not change if you use vstack - shape may vary for predictions - cannot take self._no_ones yet
# np_bias = np.ones((1,li_Z_in[0].shape[1]))
# propagation loop
# ***************
for il in ilayer:
# Step 1: Take input of last layer and apply activation function
# ******
#ts=time.perf_counter()
if il == 0:
A_out_il = li_Z_in[il] # L0: activation function is identity !!!
else:
A_out_il = self._act_func( li_Z_in[il] ) # use real activation function
# Step 2: Add bias node
# ******
# As we have taken care of this for the input layer already at data setup we
perform this only for hidden layers
if il > 0:
A_out_il = self._add_bias_neuron_to_layer(A_out_il, 'row')
li_A_out[il] = A_out_il # save data for the BW propagation
# Step 3: Propagate by matrix operation
# ******
Z_in_ilp1 = np.dot(self._li_w[il], A_out_il)
li_Z_in[il+1] = Z_in_ilp1
# treatment of the last layer
# ***************************
il = il + 1
A_out_il = self._out_func( li_Z_in[il] ) # use the output function
li_A_out[il] = A_out_il # save data for the BW propagation
return None
The required change of the first method consists of adding just one effective line
self._X = self._add_bias_neuron_to_layer(self._X, 'column')
Note that I added the column for the bias values after pre-processing. The bias neurons – more precisely – their constant values should not be regarded or included in clustering, PCA, normalization or whatever other things we do ahead of training.
In the second method we just had to eliminate a statement and add a condition, which excludes the input layer from an (additional) bias neuron treatment. That is all we need to do.
Improvements ???
How much of an improvement can we expect? Assuming that the forward propagation consumes around 40% of the total computational time of an epoch, and taking the introductory numbers we would say that we should gain something like 0.40*0.43*100 %, i.e. 17.2%. However, this too much as the basic effect of our change varies non-linearly with the batch-size.
So, something around a 15% reduction of the CPU time for a training run with 35 epochs and a batch size of only 500 would be great.
However, we should expect a much bigger effect on the FW-propagation of the complete training set (though the test data set may be more interesting otherwise). OK, let us do 2 test runs – the first without a special verification of the accuracy on the training set, the second with a verification of the accuracy via propagating the training set at the end of each and every epoch.
Results of the first run:
------------------ Starting epoch 35 Time_CPU for epoch 35 0.2717692229998647 Total CPU-time: 9.625694645001204 learning rate = 0.0009994051838157095 total costs of training set = -1.0 rel. reg. contrib. to total costs = -1.0 total costs of last mini_batch = 65.10513 rel. reg. contrib. to batch costs = 0.121494114 mean abs weight at L0 : -10.0 mean abs weight at L1 : -10.0 mean abs weight at L2 : -10.0 avg total error of last mini_batch = 0.00805 presently batch averaged accuracy = 0.99247 ------------------- Total training Time_CPU: 9.625974849001068
And the second run gives us :
------------------ Starting epoch 35 Time_CPU for epoch 35 0.37750117799805594 Total CPU-time: 13.164013020999846 learning rate = 0.0009994051838157095 total costs of training set = 5929.9297 rel. reg. contrib. to total costs = 0.0013557569 total costs of last mini_batch = 50.148125 rel. reg. contrib. to batch costs = 0.16029811 mean abs weight at L0 : 0.064023666 mean abs weight at L1 : 0.38064405 mean abs weight at L2 : 1.320015 avg total error of last mini_batch = 0.00626 presently reached train accuracy = 0.99045 presently batch averaged accuracy = 0.99267 ------------------- Total training Time_CPU: 13.16432525900018
The small deviation of the accuracy values determined by error averaging over batches vs. the test on the complete training set stems from slightly different measurement methods as discussed in the first sections of this article.
What do our results mean with respect to performance?
Well, we went down from 11.33 secs to 9.63 secs for the CPU time of the training run. This is a fair 15% improvement. But remember that we came from something like 50 secs at the beginning of our optimization, so all in all we have gained an improvement by a factor of 5 already!
In our last article we found a factor of 1.68 between the runs with a full propagation of the complete training set at each and every epoch for accuracy evaluation. Such a run lasted roughly for 19 secs. We now went down to 13.16 secs. Meaning: Instead of 7.7 secs we only consumed 3.5 secs for propagating all 60000 samples 35 times in one sweep.
We reduced the CPU time for the FW propagation of the training set (plus error evaluation) by 54%, i.e. by more than a factor of 2! Meaning: We have really achieved something for the FW-propagation of big batches!
By the way: Checking accuracy on the test dataset instead on the training dataset after each and every epoch requires 10.15 secs.
------------------ Starting epoch 35 Time_CPU for epoch 35 0.29742689200065797 Total CPU-time: 10.150781942997128 learning rate = 0.0009994051838157095 total costs of training set = -1.0 rel. reg. contrib. to total costs = -1.0 total costs of last mini_batch = 73.17834 rel. reg. contrib. to batch costs = 0.10932728 mean abs weight at L0 : -10.0 mean abs weight at L1 : -10.0 mean abs weight at L2 : -10.0 avg total error of last mini_batch = 0.00804 presently reached test accuracy = 0.96290 presently batch averaged accuracy = 0.99269 ------------------- Total training Time_CPU: 10.1510079389991
You see the variation in the accuracy values.
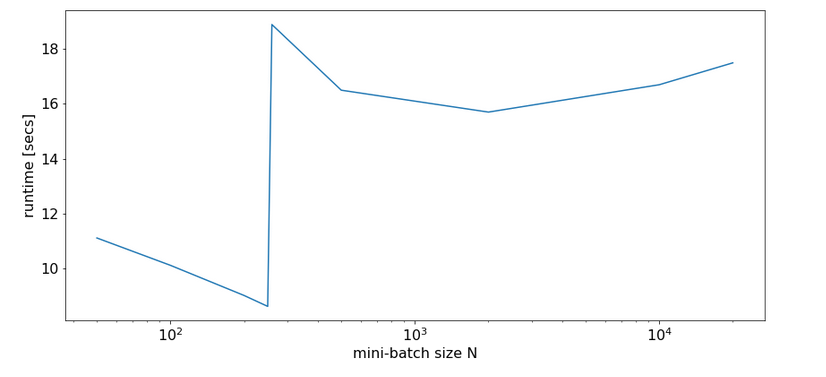
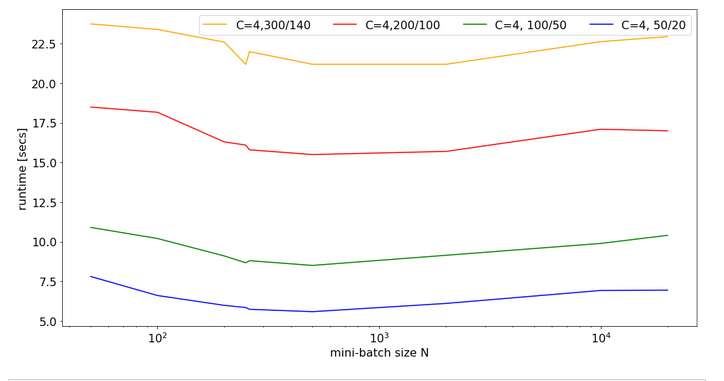
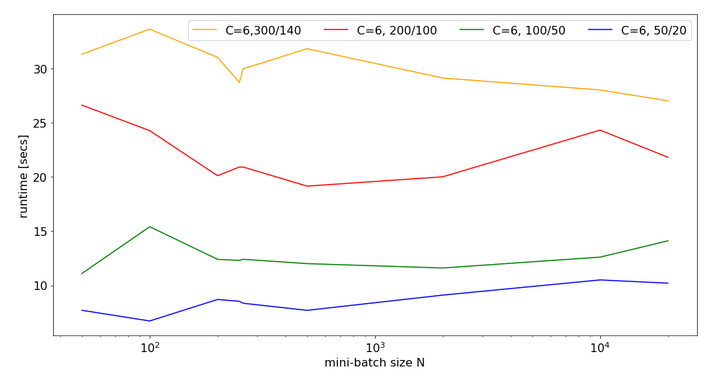
Eventually, I give you run times for 35 epochs of the MLP for larger batch sizes:
bs = 500 => t(35) = 9.63 secs bs = 5000 => t(35) = 8.75 secs bs = 10000 => t(35) = 8.55 secs bs = 20000 => t(35) = 8.68 secs bs = 30000 => t(35) = 8.65 secs
So, we get not below a certain value – despite the fact that FW-propagation gets faster with batch-size. So, we have some more batch-size dependent impediments in the BW-propagation, too, which compensate our gains.
Plots
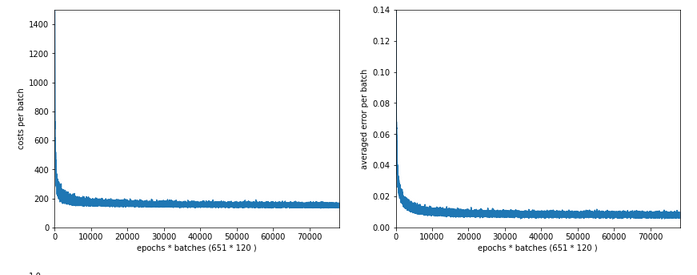
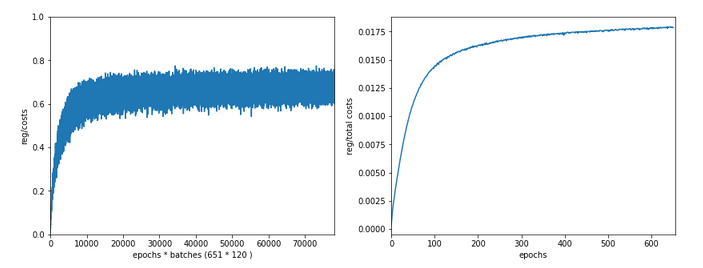
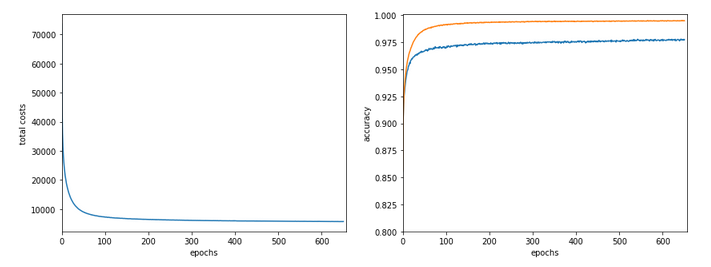
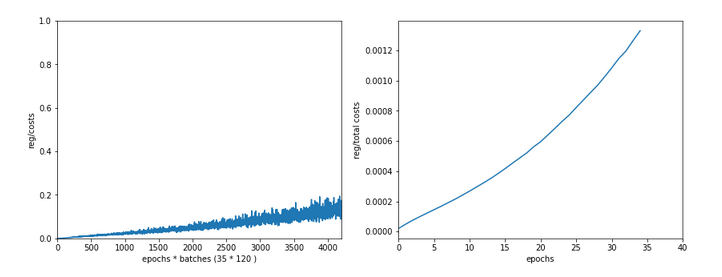
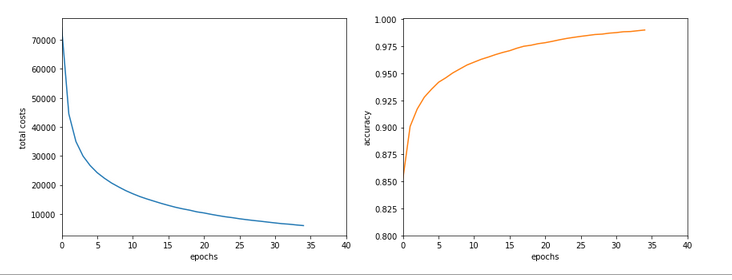
Just to show that our modified program still produces reasonable results after 650 training steps – here the plot and result data :
------------------ Starting epoch 651 .... .... avg total error of last mini_batch = 0.00878 presently reached train accuracy = 0.99498 presently reached test accuracy = 0.97740 presently batch averaged accuracy = 0.99214 ------------------- Total training Time_CPU: 257.541123711002
The total time was to be expected as we checked accuracy values at each and every epoch both for the complete training and the test datasets (635/35*14 = 260 secs = 2.3 min!).
Conclusion
This was a funny ride today. We found a major
impediment for a fast FW-propagation. We determined its cause in the inefficient combination of two differently ordered matrices which we used to account for bias nodes in the input layer. We investigated some optimization options for our present approach regarding bias neurons at layer L0. But it was much more reasonable to circumvent the whole problem by adding bias values already to the input array itself. This gave us a significant improvement for the FW-propagation of big batches – roughly by a factor of 2.5 for the complete training data set as an extreme example. But also testing accuracy on the full test data set at each and every epoch is no major performance factor any longer.
However, our whole analysis showed that we must put a big question mark behind our present approach to bias neurons. But before we attack this problem, we shall take a closer look at BW-propagation in the next article:
MLP, Numpy, TF2 – performance issues – Step III – a correction to BW propagation
And there we shall replace another stupid time wasting part of the code, too. It will give us another improvement of around 15% to 20%. Stay tuned …
Links
Performance of class methods vs. pure Python functions
stackoverflow : how-much-slower-python-classes-are-compared-to-their-equivalent-functions
Shuffle columns?
stackoverflow: shuffle-columns-of-an-array-with-numpy
Numpy arrays or matrices?
stackoverflow : numpy-np-array-versus-np-matrix-performance