When performing Computer based text analysis we sometimes need to shorten our texts by some criterion before we apply machine learning algorithms. One of the reasons could be that a classical vectorization process applied to the original texts would lead to matrices or tensors which are beyond our PC’s memory capabilities.
Another reason for shortening might be that we want to focus our analysis upon words or tokens which are “significant” for the text documents we are dealing with. The individual texts we work with typically are members of a limited collection of texts -a so called text corpus.
What does “significance” mean in this context? Well, words which are significant for a specific text should single out this text among all other texts of the corpus – or vice versa. There should be a strong and specific correlation between the text and its “significant” tokens. Such a kind of distinguishing correlation could be: The significant tokens may appear in the selected specific document, only, or especially often – always in comparison to other texts in the corpus.
It is clear that we need some “measure” for the significance of a token with respect to individual texts. And we somehow need to compare the frequency by which a word/token appears in a text with the frequencies by which the token appears in other texts of our corpus.
For some analysis we might in the end only keep “significant” words which distinguish the texts of corpus from each other. Note that such a shortening procedure would reduce the full vocabulary, i.e. the set of all unique words appearing in the corpus’ texts. And after shortening the statistical basis for “significance” may have changed.
Due to the impact the choice for a “measure” of a token’s significance may have on our eventual analysis results we must be careful and precise when we discuss our results and its presuppositions. We should name the formula used for the measure of significance.
This leads to the question: Do all authors in the filed of Machine Learning [ML] and NLP use the same formula for the “significance” of words or tokens? Are there differences which can be confusing? Oh yes, there are … The purpose of this post is to remind beginners in the field of NLP or text analysis about this fact and to give an overview over the most common approaches. In addition I will discuss some practical aspects and give some snippets of a code which reproduces the TF-IDF values which the fast Keras Tokenizer would give you.
For large corpus the differences of using different formulas for measuring the significance of tokens may be minor and not change fundamental conclusions. But in my opinion the differences should be checked and at least be named.
TF-IDF values as a measure of a token’s significance – dependency on token, text AND corpus
In NLP a measure of a word’s significance with respect to a specific text of a defined corpus is given by a quantity called “TF-IDF” – “term frequency – inverse document frequency” (see below). TF-IDF values are specific for a word (or token) and a selected text (out of the corpus). We will discuss elements of the formulas in a minute.
Note: Significance values as TF-IDF values will in general depend on the corpus, too.
This is due to the fact that “significance” is based on correlations. As said above: We need to compare token frequencies within a specific text with the token’s frequencies in other texts of the given corpus. Significance is thus rooted in a singular text and the collection of other texts in a specific corpus. Keep a specific text, but change the corpus (e.g by eliminating some texts) – and you will change the significance of a token for the selected text.
If you have somehow calculated “TF-IDF”-values for all the words used in a specific text (of the corpus collection), a simple method to shorten the selected text would be to use a “TF-IDF”-threshold: We keep words which have a “TF-IDF”-value above the defined threshold and omit others from our specific text.
Such a shortening procedure would depend on word- and text-specific values. Another way of shortening could be based on an averaged TF-IDF value of each token evaluated over all documents. We then would get a corpus-specific significance value <TF-IDF> for each of our tokens. Such averaged TF-IDF values for our tokens together with a threshold could also be used for text-shortening.
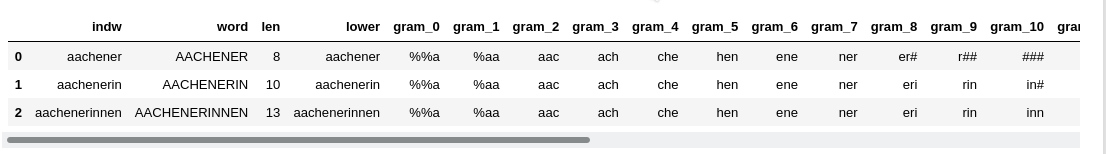
Whatever method for shortening you choose: “TF-IDF”-values require a statistical analysis over the given ensemble of texts, i.e. the text corpus. The basic statistical data are often collected during the application of a tokenizer to the texts of a given corpus. A tokenizer identifies unique tokens appearing in the texts of a corpus and collects them in a long vector. Such a vector represents the tokenizer’s (and the corpus’) “vocabulary”. The tokenizer vocabulary often is sorted by the total frequency of the tokens in the corpus. In a Python environment the tokenizer’s vocabulary often will be represented by one or more Python dictionary objects.
Technical obstacles may require the explicit calculation of TF-IDF values by a coded formula in your programs
NLP frameworks in most cases provide specific objects and methods to automatically calculate TF-IDF values for the tokens and texts of a corpus during certain analysis runs. But things can become problematic because some tokenizers provide “TF-IDF”-data during a vectorization procedure, only. By vectorization we mean a digital encoding of the texts with respect to the tokenizer’s vocabulary in a common way for all texts.
An example is one-hot encoding: Each text can be represented by numbers 0,1..,n at positions in a long vector in which each position represents a specific token. 0,1,..n would then mean: In this text the token appears 0,1 or n times. Such a kind of encoding is especially useful for the training of neural networks.
A tool which gives us TF-IDF values after some vectorization is the Keras Tokenizer. I prefer the Keras Tokenizer in my projects because it really is super fast.
But: You see at once that we may run into severe trouble if we need to feed all of the tokens of a really big corpus into a TF-IDF analysis based on vectorization. For a collection of texts the number of unique tokens may lie in the range of several millions. This in turn leads to very long vectors. The number of vectors to look at is given by the number of texts which in itself may be hundreds of thousands or even millions. You can do the math for RAM requirements with a 16- or 32 Bit-representation yourself. As a consequence you may have to accept that you can do vectorization only batch-wise. And this may be time-consuming and may require intermediate manual interaction with your programs – especially when working with Python code and Jupyter notebooks (see below).
For the analysis of huge text corpora the snake of tokenizing, TF-IDF calculation, reasonable shortening and vectorization for ML may bite in its tail:
We need tf-idf to to shorten texts in a reasonable way and to avoid later memory problems during vectorization for ML tasks. But sometimes our tool-kit provides “tf-idf”-data only after a first vectorization, which may not be feasible due to the size of our corpus and the size of the resulting vocabulary of the tokenizer.
A typical example is given by the Keras tokenizer. If you have a big corpus with millions of texts and tokens vectorization may not be a good recipe to get your TF-IDF values. Especially in the case of post-OCR applications you may not be allowed to throw away any of the identified tokens before you have corrected them for possible OCR errors. And your ML-mechanism for such correction may depend on the results of some kind of TF-IDF analysis.
In such a situation one must invest some (limited) effort into a “manual” calculation of TF-IDF values. You then have to pick some formula from a text book on NLP and/or ML-based text analysis. But having done so you may soon find out that your (text-book) formula for a “TF-IDF”-calculation does not reproduce the values the tokenizer of a selected NLP framework would have given you after a vectorization of your texts. Note that you can always check for such differences by using an artificially and drastically reduced corpus.
A formula for the TF-IDF calculation performed the Keras tokenizer is one of the central topic of this post. I omit the hyphen in TF-IDF below sometimes for convenience reasons.
Is it worth the effort to calculate TF-IDF values by your own code?
Tokenizers of a NLP tool-kit do not only identify individual tokens in a text, but are also capable to vectorize texts. Vectorization leads to the representation of a text by an (ordered) series of integer or float numbers, i.e. a vector, which in a unique way refers to the words of a vocabulary extracted from the text collection:
The indexed position in the vector refers to a specific word in the vocabulary of the text corpus, the value given at this position instead describes the word’s (statistical) appearance in a text in some way.
We saw already that “one-hot”-encoding for vectorization may result in a “bag-of words”-model: A word appearing in a text is marked by a “one” (or integer) in an indexed vector referring to tokens of the ordered vocabulary derived by a tokenizer. But vectorization can be provided in different modes, too: The “ones” (1) in a simple “one-hot-encoded” vector can e.g. be replaced by TF-IDF values (floats) of the words (tfidf-mode). This is the case for the Keras Tokenizer: By using respective Keras tokenizer functions you may get the aspired TFIDF-values for reducing the texts during a vectorization run. However, all one-hot like encodings of texts come with a major disadvantage:
The length of the word vectors depends on the number of words the tokenizer has identified over all texts in a collection for the vocabulary.
If you have extracted 3 million words (tokens) out of hundreds of thousands of texts you may run into major trouble with the RAM of PC (and CPU-time). Most tokenizers allow for a (manual) sequential vectorization approach for a limited number of texts to overcome memory problems under such circumstances. What does this mean practically when working with Jupyter notebooks?
Well, if you work with notebooks on a PC with a small amount of RAM – and small may mean 128 GB (!) in some cases – you may have to perform a sequence of vectorization runs, each with maximally some hundred or thousand texts. Then you may have to export your results and afterward manually reset your notebook kernel to get rid of the RAM consumption – just because the standard garbage collection of your Python environment may not work fast enough.
I recently had this problem in a project with 200,000 texts, the Keras tokenizer and a vocabulary of 2.4 million words (where words with less than 4 characters were already omitted). The Keras tokenizer produces almost all relevant data for a “manual” calculation of TF-IDFF values after it has been applied to a corpus. In my case the CPU-time required to tokenize and build a vocabulary for the 200,000 texts took 20 secs, only. A manual and sequential approach to create all TFIDF values via vectorization, however, required about an hour’s time. This was due both to the time the vectorization and tf-idf calculation needed and the time required for resetting the notebook kernel.
After I had decided to implement the TFIDF calculation on my own in my codes I could work on the full corpus and get the values within a minute. So, if one has to work on a big corpus multiple times with some iterative processes (e.g. in post-OCR procedures) we may talk of a performance difference in terms of hours.
Therefore, I would in general recommend to perform TFIDF-calculations by your own code segments when being confronted with big corpora – and not to rely on (vectorizing) tools of a framework. Besides performance aspects another reason for TF-IDF functions programmed by yourself is that you afterward know exactly what kind of TF-IDF formula you have used.
TF-IDF formulas: The “IDF”-term – and what does the Keras tokenizer use for it?
The TF-IDF data describe the statistical overabundance of a token in a specific text by some formula measuring the token’s frequency in the selected text in comparison to the frequency over all texts in a weighted and normalized way.
During my own “TF-IDF”-calculations based on some Python code and basic tokenizer data, I, of course, wanted to reproduce the values the Keras tokenizer gave me during my previous vectorization approach. To my surprise it was rather difficult to achieve this goal. Just using a reasonable “TF-IDF”-formula taken from some NLP text-book simply failed.
The reason was that “TF-IDF”-data can be and are indeed calculated in different ways both in NLP literature adn in NLP frameworks. The Keras tokenizer does it differently than the tokenizer of SciKit – actually for both the TF and the IDF-part. There is a basic common structure behind a normalized TFIDF-value; there are, however, major differences in the details. Lets look at both points.
Everybody who has once in his/her life programmed a search engine knows that the significance of a word for a specific text (of an ensemble) depends on the number of occurrences of the word inside the specific text, but also on the frequency of the very same word in all the other texts of a given text collection:
If a word also appears very often in all other texts of a text ensemble then the word is not very significant for the specific text we are looking at.
Examples are typical “stop-words” – like “this” or “that” or “and”. Such words appear in very many texts of a corpus of English texts. Therefore, stop-words are not significant for a specific text.
Thus we expect that a measure of the statistical overabundance of a word in a selected text is a combination of the abundance in this specific text and a measure of the frequency in all the other texts of the corpus. The “TF-IDF” quantity follows this recipe. It is a combination of the so so called “term frequency” [tf(t)] with the “inverse document frequency [idf(t)], with “t” representing a special token or term:
tfidf(t) = tf(t) * idf(t)
While the term frequency measures the occurrence or frequency of a word within a selected text, the “idf” factor measures the frequency of a word in multiple texts of the collection. To get some weighing and normalization into this formula, the “idf”-term is typically based on the natural logarithm of the fraction described
- by the number of texts NT comprised by a corpus as the nominator
- and the number of documents ND(t) in which a special word or term appears as the denominator.
Once again: A TF-IDF value is always characteristic of a word or term and the specific text we look at. But, the “idf”-term is calculated in different manners in various text-books on text-analysis. Most variants try to avoid the idf-term becoming negative or want to avoid a division by zero; typical examples are:
-
idf(t) = log( NT / (ND + 1) )
-
idf(t) = log( (1 + NT) / (ND + 1) )
-
idf(t) = log( 1 + NT / (ND + 1) )
-
idf(t) = log( 1 + NT / ND )
-
idf(t) = log( (1 + NT) / (ND + 1) ) + 1
Note: log() represents the natural logarithm above.
Who uses which IDF version?
The second variant appears e.g. in a book of S. Raschka (see below) on “Python Machine Learning” (2016, Packt Publishing).
The fourth version in the list above is used in Scikit according to https://melaniewalsh.github.io/Intro-Cultural-Analytics/05-Text-Analysis/03-TF-IDF-Scikit-Learn.html
This is in so far consistent to Raschka’s version as he himself characterizes the SciKit “TF-IDF” version as:
tfidf(t) = tf(t) * [ idf(t) + 1 ]
Keras: The third variant is the one you find in the source code of the Keras tokenizer. The strange thing is that you also find a reference in the Keras code which points to a section in a Wikipedia article that actually reflects the fourth form (!) given in the list above.
Source code excerpt of the Keras Tokenizer (as of 10/2021):
.....
.....
elif mode == 'tfidf':
# Use weighting scheme 2 in
# https://en.wikipedia.org/wiki/Tf%E2%80%93idf
tf = 1 + np.log(c)
idf = np.log(1 + self.document_count /
(1 + self.index_docs.get(j, 0)))
x[i][j] = tf * idf
.....
.....
What we learn from this is that there are several variants of the “IDF”-term out there. So, if you want to reproduce TFIDF-numbers of a certain NLP framework you should better look into the code of your framework’s classes and functions – if possible.
Variants of the “term frequency” TF? Yes, they do exist!
While I had already become aware of the existence of different IDF-variants, I did not at all know that here were and are even differences regarding the term-frequency “tf(t)“. Normally, one would think that it is just the number describing how often a certain word or term appears in a specific text, i.e. the token’s frequency fro the selected text.
Let us, for example, assume that we have turned a specific text via a tokenizer function into a “sequence” of numbers. An entry in this sequence refers to a unique number assigned to a word of a somehow sorted tokenizer vocabulary. A tokenizer vocabulary is typically represented by a Python dictionary where the key is the word itself (or a hash of it) and the value corresponds to a unique number for the word. (Hint: In my applications for texts I always create a supplementary dictionary, which I call “switched_vocab”, with keys and values switched (number => word). Such a dictionary is useful for a lot of analysis steps)
A sequence can typically be represented by a Python list of numbers “li_seq”: the position in the list corresponds to the word’s position in the text (marked by separators), the number given corresponds to the words unique index number in the vocabulary.
Then, with Python 3, a straight-forward code snippet to get simple tf-values (as we sum of the number’s frequency in the sequence) would be
ind_w = li_seq[i] # with "i" selecting a specific point or word in the sequence d_count = Counter(li_seq) tf = d_count[ind_w]
This code creates a dictionary “d_count” with the word’s unique number appearing in the original sequence and the sum of occurrences of this specific number in the text’s sequence – i.e. in the text we are looking at.
Does the Keras tokenizer calculate and use the tf-term in this manner when it vectorizes texts in tfidf-mode? No, it does not! And this was a major factor for the differences in comparison to the TFIDF-values I naively produced for my texts.
With the terms above the Keras tokenizer instead uses a logarithmic value for tf (= TF):
ind_w = li_seq[i] # i selecting a specific point or word in the sequence d_count = Counter(li_seq) tf = log( 1 + d_count[ind_w] )
This makes a significant difference in the derived “TF-IDF” values – even if one had gotten the “IDF”-term right!
Please note that all of the variants used for the TF- and the IDF-terms have their advantages and disadvantages. You should at least know exactly which formula you use in your analysis. In my project the Keras way of doing TF-IDF was useful, but there may be cases where another choice is appropriate.
Quick and dirty Python code to calculate TF-IDF values manually for a list of texts with the Keras tokenizer
For reasons of completeness, I outline some code fragments below, which may help readers to calculate “TF-IDF”-values, which are consistent with those produced during “sequences to matrix”-vectorization runs with the Keras tokenizer (as of 10/2021).
I assume that you already have a working Keras implementation using either CPU or GPU. I further assume that you have gathered a collection of texts (cleansed by some Regex operations) in a column “txt” of a Pandas dataframe “df_rex”. We first extract all the texts into a list (representing the corpus) and then apply the Keras tokenizer:
from tensorflow.keras import preprocessing
from tensorflow.keras.preprocessing.text import Tokenizer
num_words = 1800000 # or whatever number of words you want to be taken into account from the vocabulary
li_txts = df_rex['txt'].to_list()
tokenizer = Tokenizer(num_words=num_words, lower=True) # converts tokens to lower-case
tokenizer.fit_on_texts(li_txts)
vocab = tokenizer.word_index
w_count = tokenizer.word_counts
w_docs = tokenizer.word_docs
num_tot_vocab_words = len(vocab)
# Switch vocab - key <> value
# ****************************
switched_vocab = dict([(value, key) for key, value in vocab.items()])
Tokenizing should be a matter of seconds or a few ten-second intervals depending on the number of texts and the length of the texts. In my case with 200,000 texts, on average each with 2000 words, it took 25 secs and produced a vocabulary of about 2.4 million words.
In a next step we create “integer sequences” from all texts:
li_seq_full = tokenizer.texts_to_sequences(li_txts) leng_li_seq_full = len(li_seq_full)
Now, we are able to create a super-list of lists – including a list of tf-idf-values per text:
li_all_txts = []
j_end = leng_li_seq_full
for j in range(0, j_end):
li_text = []
li_text.append(j)
leng_seq = len(li_seq_full[j])
li_seq = []
li_tfidf = []
li_words = []
d_count = {}
d_count = Counter(li_seq_full[j])
for i in range(0,leng_seq):
ind_w = li_seq_full[j][i]
word = switched_vocab[ind_w]
# calculation of tf-idf
# ~~~~~~~~~~~~~~~~~~~~~
# https://github.com/keras-team/keras-preprocessing/blob/1.1.2/keras_preprocessing/text.py#L372-L383
# Use weighting scheme 2 in https://en.wikipedia.org/wiki/Tf%E2%80%93idf
dfreq = w_docs[word] # document frequency
idf = np.log( 1.0 + (leng_li_seq_full) / (dfreq + 1.0) )
tf_basic = d_count[ind_w]
tf = 1.0 + np.log(tf_basic)
tfidf = tf * idf
li_seq.append(ind_w)
li_tfidf.append(tfidf)
li_words.append(word)
li_text.append(li_seq)
li_text.append(li_tfidf)
li_text.append(li_words)
li_all_txts.append(li_text)
leng_li_all_txts = len(li_all_txts)
This last run took about 1 minute in my case. When getting the same numbers with a sequential approach calculating Keras vectorization matrices in tf-idf mode for around 6000 texts in each run with in-between memory cleansing it took me around an hour and required continuous manual system interactions. Well, such a run provides the encoded text vectors as one of its major products, but, actually, we do not need these vectors for just evaluating TF-IDF values.
Conclusion
In this article I have demonstrated that “TF-IDF”-values can be calculated almost directly from the output of a tokenizer like the Keras Tokenizer. Such a “manual” calculation by one’s own coded instructions is preferable in comparison to e.g. a Keras based vectorization run in “tf-idf”-mode when both the number of texts and the vocabulary of your text corpus are huge. “tf-idf”-word-vectors may easily get a length of more than a million words with reasonably complex text ensembles. This poses RAM problems on many PC-based systems.
With TF-IDF-values calculated by your code functions you will get a measure for the significance of words or tokens within a reasonable CPU time and without any RAM problems. The evaluated “TF-IDF”- values may afterward help you to shorten your texts in a well founded and reasonable way before you vectorize your texts, i.e. ahead of applying advanced ML-algorithms.
Formulas for TF-IDF values used in the literature and various NLP tool-kits or frameworks do differ with respect to both for the TF-terms and the IDF-terms. You should be aware of this fact and choose one of the formulas given above carefully. You should also specify the formula used during your text analysis work when presenting results to your customers.