In the last posts of this mini-series we have studied if and how we can use three 3-char-grams at defined positions of a string token to identify matching words in a reference vocabulary. We have seen that we should choose some distance between the char-grams and that we should use the words length information to keep the list of possible hits small.
Such a search may be interesting if there is only fragmented information available about some words of a text or if one cannot trust the whole token to be written correctly. There may be other applications. Note: This has so far nothing to do with text analysis based on machine learning procedures. I would put the whole topic more in the field of text preparation or text rebuilding. But, I think that one can combine our simple identification of fitting words by 3-char-grams with ML-methods which evaluate the similarity or distance of a (possibly misspelled) token with vocabulary words: When we get a long hit-list we could invoke ML-methods to to determine the best fitting word.
We saw that we can do a 100,000 search runs with 3-char-grams on a decent vocabulary of around 2 million words in a Pandas dataframe below a 1.3 minutes on one CPU core of an older PC. In this concluding article I want to look a bit at the idea of multiprocessing the search with up to 4 CPU cores.
Points to take into account when using multiprocessing – do not expect too much
Pandas normally just involves one CPU core to do its job. And not all operations on a Pandas dataframe may be well suited for multiprocessing. Readers who have followed the code fragments in this series so far will probably and rightly assume that there is indeed a chance for reasonably separating our search process for words or at least major parts of it.
But even then – there is always some overhead to expect from splitting a Pandas dataframe into segments (or “partitions”) for a separate operations on different CPU cores. Overhead is also expected from the task to correctly to combine the particular results from the different processor cores to a data unity (here: dataframe) again at the end of a multiprocessed run.
A bottleneck for multiprocessing may also arise if multiple processes have to access certain distinct objects in memory at the same time. In our case we this point is to be expected for the access of and search within distinct sub-dataframes of the vocabulary containing words of a specific length.
Due to overhead and bottlenecks we do not expect that a certain problem scales directly and linearly with the number of CPU cores. Another point is that although the Linux OS may recognize a hyperthreading physical core of an Intel processor as two cores – but it may not be able to use such virtual cores in a given context as if they were real separate physical cores.
Code to invoke multiple processor cores
In this article I just use the standard Python “multiprocessing” module. (I did not test Ray yet – as a first trial gave me trouble in some preparing code-segments of my Jupyter notebooks. I did not have time to solve the problems there.)
Following some advice on the Internet I handled parallelization in the following way:
import multiprocessing
from multiprocessing import cpu_count, Pool
#cores = cpu_count() # Number of physical CPU cores on your system
cores = 4
partitions = cores # But actually you can define as many partitions as you want
def parallelize(data, func):
data_split = np.array_split(data, partitions)
pool = Pool(cores)
data = pd.concat(pool.map(func, data_split), copy=False)
pool.close()
pool.join()
return data
The basic function, corresponding to the parameter “func” of function “parallelize”, which shall be executed in our case is structurally well known from the last posts of this article series:
We perform a search via
putting conditions on columns (of the vocabulary-dataframe) containing 3-char-grams at different positions. The search is done on sub-dataframes of the vocabulary containing only words with a given length. The respective addresses are controlled by a Python dictionary “d_df”; see the last post for its creation. We then build a list of indices of fitting words. The dataframe containing the test tokens – in our case a random selection of real vocabulary words – will be called “dfw” inside the function “func() => getlen()” (see below). To understand the code you should be aware of the fact that the original dataframe is split into (4) partitions.
We only return the length of the list of hits and not the list of indices for each token itself.
# Function for parallelized operation
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
def getlen(dfw):
# Note 1: The dfw passed is a segment ("partition") of the original dataframe
# Note 2: We use a dict d_lilen which was defined outside
# and is under the control of the parallelization manager
num_rows = len(dfw)
for i in range(0, num_rows):
len_w = dfw.iat[i,0]
idx = dfw.iat[i,33]
df_name = "df_" + str(len_w)
df_ = d_df[df_name]
j_m = math.floor(len_w/2)+1
j_l = 2
j_r = len_w -1
col_l = 'gram_' + str(j_l)
col_m = 'gram_' + str(j_m)
col_r = 'gram_' + str(j_r)
val_l = dfw.iat[i, j_l+2]
val_m = dfw.iat[i, j_m+2]
val_r = dfw.iat[i, j_r+2]
li_ind = df_.index[ (df_[col_r]==val_r)
& (df_[col_m]==val_m)
& (df_[col_l]==val_l)
]
d_lilen[idx] = len(li_ind)
# The dataframe must be returned - otherwise it will not be concatenated after parallelization
return dfw
While the processes work on different segments of our input dataframe we write results to a Python dictionary “d_lilen” which is under the control of the “parallelization manager” (see below). A dictionary is appropriate as we might otherwise loose control over the dataframe-indices during the following processes.
A reduced dataframe containing randomly selected “tokens”
To make things a bit easier we first create a “token”-dataframe “dfw_shorter3” based on a random selection of 100,000 indices from a dataframe containing long vocabulary words (length ≥ 10). We can derive it from our reference vocabulary. I have called the latter dataframe “dfw_short3” in the last post (because we use three 3-char-grams for longer tokens). “dfw_short3” contains all words of our vocabulary with a length of “10 ≤ length ≤ 30”.
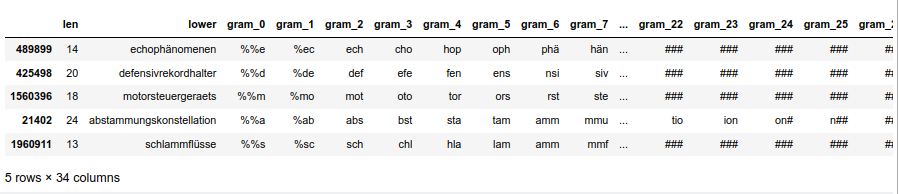
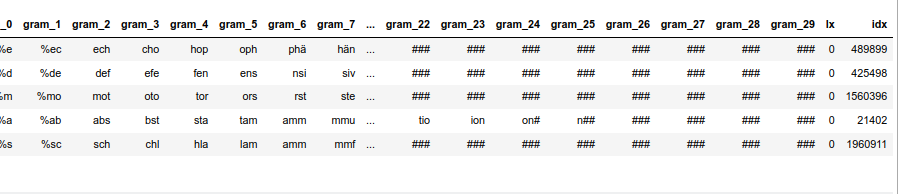
# Prepare a sub-dataframe for of the random 100,000 words # ****************************** num_w = 100000 len_dfw = len(dfw_short3) # select a 100,000 random rows random.seed() # Note: random.sample does not repeat values li_ind_p_w = random.sample(range(0, len_dfw), num_w) len_li_p_w = len(li_ind_p_w) dfw_shorter3 = dfw_short3.iloc[li_ind_p_w, :].copy() dfw_shorter3['lx'] = 0 dfw_shorter3['idx'] = dfw_shorter3.index dfw_shorter3.head(5)
The resulting dataframe “dfw_shorter3” looks like :
nYou see that the index varies randomly and is not in ascending order! This is the reason why we must pick up the index-information during our parallelized operations!
Code for executing parallelized run
The following code enforces a parallelized execution:
manager = multiprocessing.Manager()
d_lilen = manager.dict()
print(len(d_lilen))
v_start_time = time.perf_counter()
dfw_res = parallelize(dfw_shorter3, getlen)
v_end_time = time.perf_counter()
cpu_time = v_end_time - v_start_time
print("cpu : ", cpu_time)
print(len(d_lilen))
mean_length = sum(d_lilen.values()) / len(d_lilen)
print(mean_length)
The parallelized run takes about 29.5 seconds.
cpu : 29.46206265499968 100000 1.25008
How does cpu-time vary with the number of cores of my (hyperthreading) CPU?
The cpu-time does not improve much when the number of cores gets bigger than the number of real physical cores:
1 core : 90.5 secs 2 cores: 47.6 secs 3 cores: 35.1 secs 4 cores: 29.4 secs 5 cores: 28.2 secs 6 cores: 26.9 secs 7 cores: 26.0 secs 8 cores: 25.5 secs
My readers know about this effect already from ML experiments with CUDA and libcublas:
As long a s we use physical processor cores we see substantial improvement, beyond that no real gain in performance is observed on hyperthreading CPUs.
Compared to a run with just one CPU core we seem to gain a factor of almost 3 by parallelization. But, actually, this is no fair comparison: My readers have certainly seen that the CPU-time for the run with one CPU-Core is significantly slower than comparable runs which I described in my last post. At that time we found a cpu-time of around 75 secs, only. So, we have a basic deficit of about 15 secs – without real parallelization!
Overhead and RAM consumption of multiprocessing
Why does run with just one CPU core take so long time? Is it functional overhead for organizing and controlling multiprocessing – which may occur despite using just one core and just one “partition” of the dataframe (i.e. the full dataframe)? Well, we can test this easily by reconstructing the runs of my last post a bit:
# Reformulate Run just for cpu-time comparisons
# **********************************************
b_test = True
# Function
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
def getleng(dfw, d_lileng):
# Note 1: The dfw passed is a segment of the original dataframe
# Note 2: We use a list l_lilen which was outside defined
# and is under the control of the prallelization manager
num_rows = len(dfw)
#print(num_rows)
for i in range(0, num_rows):
len_w = dfw.iat[i,0]
idx = dfw.iat[i,33]
df_name = "df_" + str(len_w)
df_ = d_df[df_name]
j_m = math.floor(len_w/2)+1
j_l = 2
j_r = len_w -1
col_l = 'gram_' + str(j_l)
col_m = 'gram_' + str(j_m)
col_r = 'gram_' + str(j_r)
val_l = dfw.iat[i, j_l+2]
val_m = dfw.iat[i, j_m+2]
val_r = dfw.iat[i, j_r+2]
li_ind = df_.index[ (df_[col_r]==val_r)
& (df_[col_m]==val_m)
& (df_[col_l]==val_l)
]
leng = len(li_ind)
d_lileng[idx] = leng
return d_lileng
if b_test:
num_w = 100000
len_dfw = len(dfw_short3)
# select a 100,000 random rows
random.seed()
# Note: random.sample does not repeat values
li_ind_p_w = random.sample(range(0, len_dfw), num_w)
len_li_p_w = len(li_ind_p_w)
dfw_shortx = dfw_short3.iloc[li_ind_p_
w, :].copy()
dfw_shortx['lx'] = 0
dfw_shortx['idx'] = dfw_shortx.index
d_lileng = {} #
v_start_time = time.perf_counter()
d_lileng = getleng(dfw_shortx, d_lileng)
v_end_time = time.perf_counter()
cpu_time = v_end_time - v_start_time
print("cpu : ", cpu_time)
print(len(d_lileng))
mean_length = sum(d_lileng.values()) / len(d_lileng)
print(mean_length)
dfw_shortx.head(3)
How long does such a run take?
cpu : 77.96989408900026 100000 1.25666
Just 78 secs! This is pretty close to the number of 75 secs we got in our last post’s efforts! So, we see that turning to multiprocessing leads to significant functional overhead! The gain in performance, therefore, is less than the factor 3 observed above:
We (only) get a gain in performance by a factor of roughly 2.5 – when using 4 physical CPU cores.
I admit that I have no broad or detailed experience with Python multiprocessing. So, if somebody sees a problem in my code, please, send me a mail.
RAM is not released completely
Another negative side effect was the use of RAM in my case. Whereas we just get 2.2 GB RAM consumption with all required steps and copying parts of the loaded dataframe with all 3-char-grams in the above test run without multiprocessing, I saw a monstrous rise in memory during the parallelized runs:
Starting from a level of 2.4 GB, memory rose to 12.5 GB during the run and then fell back to 4.5 GB. So, there are copying processes and memory is not completely released again in the end – despite having all and everything encapsulated in functions. Repeating the multiprocessed runs even lead to a systematic increase in memory by about 150 MB per run.
So, when working with the “multiprocessing module” and big Pandas dataframes you should be a bit careful about the actual RAM consumption during the runs.
Conclusion
This series about finding words in a vocabulary by using two or three 3-char-grams may have appeared a bit “academical” – as one of my readers told me. Why the hell should someone use only a few 3-char-grams to identify words?
Well, I have tried to give some answers to this question: Under certain conditions you may only have fragments of words available; think of text transcribed from a recorded, but distorted communication with Skype or think of physically damaged written text documents. A similar situation may occur when you cannot trust a written string token to be a correctly written word – due to misspelling or other reasons (bad OCR SW or bad document conditions for scans combined with OCR).
In addition: character-grams are actually used as a basis for multiple ML methods for text-analysis tasks, e.g. in Facebook’s Fasttext. They give a solid base for an embedded word vector space which can help to find and measure similarities between correctly written words, but also between correctly written words and fantasy words or misspelled words. Looking a bit at the question of how much a few 3-char-grams help to identify a word is helpful to understand their power in other contexts, too.
We have seen that only three 3-char-grams can identify matching words quite well – even if the words are long words (up to 30 characters). The list of matching words can be kept surprisingly small if and when
- we use available or reasonable length information about the words we want to find,
- we define positions for the 3-char-grams inside the words,
- we put some positional distance between the location of the chosen 3-char-grams inside the words.
For a 100,000 random cases with correctly written 3-char-grams the average length of the hit list was below 2 – if the distance between the 3-char-grams was
reasonably large compared to the token-length. Similar results were found for using only two 3-char-grams for short words.
We have also covered some very practical aspects regarding search operation on relatively big Pandas dataframes :
The CPU-time for identifying words in a Pandas dataframe by using 3-char-grams is reasonably small to allow for experiments with around 100,000 tokens even on PCs within minutes or quarters of an hour – but it does not take hours. As using 3-char-grams corresponds to putting conditions on two or three columns of a dataframe this result can be generalized to other similar problems with string comparisons on dataframe columns.
The basic RAM consumption of dataframes containing up to fifty-five 3-char-grams per word can be efficiently controlled by using the dtype “category” for the respective columns.
Regarding cpu-time we saw that working with many searches may get a performance boost by a factor well above 2 by using simple multiprocessing techniques based on Python’s “multiprocessing” module. However, this comes with an unpleasant side effect of enormous RAM consumption – at least temporarily.
I hope you had some fun with this series of posts. In a forthcoming series I will apply these results to the task of error correction. Stay tuned.
Links
https://towardsdatascience.com/staying-sane-while-adopting-pandas-categorical-datatypes-78dbd19dcd8a
https://thispointer.com/python-pandas-select-rows-in-dataframe-by-conditions-