During last weekend I wanted to compare the performance of an old 20-line Keras setup for a simple MLP with the performance of a self-programmed Python- and Numpy-based MLP regarding training epochs on the MNIST dataset. The Keras code was set up in a Jupyter notebook last autumn – at that time for TensorFlow 1 and Cuda 10.0 for my Nvidia graphics card. I thought it might be a good time to move everything to Tensorflow 2 and the latest Cuda libraries on my Opensuse Leap 15.1 system. This was more work than expected – and for some problems I was forced to apply some dirty workarounds. I got it running. Maybe the necessary steps, which are not really obvious, are helpful for others, too.
Install Cuda 10.2 and Cudnn on an Opensuse Leap 15.1
Before you want to use TensorFlow [TF] on a Nvidia graphics card you must install Cuda. The present version is Cuda 10.2. I was a bit naive to assume that this should be the right version – as it has been available for some time already. Wrong! Afterwards I read somewhere that TensorFlow2 [TF2] is working with Cuda 10.1, only, and not yet fully compatible with Cuda 10.2. Well, at least for my purposes [MLP training] it seemed to work nevertheless – with some “dirty” additional library links.
There is a central Cuda repository available at this address: cuda10.2. Actually, the repo offers both cuda10.0, cuda10.1 and cuda10.2 (plus some nvidia drivers). I selected some central cuda10.2 packages for installation – just to find out where the related files were placed in the filesystem. I then ran into a major chain of packet dependencies, which I had to resolve during many tedious steps . Some packages may not have been necessary for a basic installation. In the end I was too lazy to restrict the libs to what is really required for Keras. The bill came afterwards: Cuda 10.2 is huge! If you do not know exactly what you need: Be prepared to invest up to 3 GB on your hard disk.
The Cuda 10.2 RPM packets install most of the the required “*.so”-shared library files and many other additional files in a directory “/usr/local/cuda-10.2/”. To make changes between different versions of Cuda possible we also find a link “/usr/local/cuda” pointing to
“/usr/local/cuda-10.2/” after the installation. Ok, reasonable – we could change the link to point to “/usr/local/cuda-10.0/”. This makes you assume that the Tensorflow 2 libraries and other modules in your virtual Python3 and Jupyter environment would look for required Cuda files in the central directory “/usr/local/cuda” – i.e. without special version attributes of the files. Unfortunately, this was another wrong assumption. 🙁 See below.
In addition to the Cuda packages you must install the present “cudnn” libraries from Nvidia – more precisely: The runtime and the development package. You get the RPMs from here. Be prepared to give Nvidia your private data. 🙁
I should add that I ignored and ignore the Nvidia drivers from the Cuda repository, i.e. I never installed them. Instead, I took those from the standard Nvidia community repository. They worked and work well so far – and make my update life on Opensuse easier.
Installation of Tensorflow2 modules in your (virtual) Python3 environment
I use a virtual Python3 environment and update it regularly via “pip”. Regarding TF2 an update via the command “pip install –upgrade tensorflow” should be sufficient – it will resolve dependencies. (By the way: If you want to bring all Python libs to their present version you can also use “pip-review –auto”. Note that under certain circumstances you may need the “–force” option for special upgrades. I cannot go into
details in this article.)
Multiple complaints about missing libraries …
Unfortunately, the next time I started my virtual Python environment I got the warning that the dynamic library “libnvinfer.so.6” could not be found, but was required in case I planned to use TensorRT. What? Well, you may find some information here
https://blogs.nvidia.com/blog/2016/08/22/difference-deep-learning-training-inference-ai/
https://developer.nvidia.com/tensorrt
I leave it up to you whether you really need TensorRT. You can ignore this message – TF will run for standard purposes without it. But, dear TF-developers: a clear message in the warning would in my opinion have been helpful. Then I checked whether some version of the Nvidia related library “libnvinfer.so” came with Cuda or Cudnn onto my system. Yeah, it did – unfortunately version 7 instead of 6. :-(.
So, we are confronted with a dependency on a specific file version which is older than the present one. I do not like this style of development. Normally, it should be the other way round: If a newer version is required due to new capabilities you warn the user. But under normal circumstances a backward compatibility of libs should be established. You would assume such a backward compatibility and that TF would search for the present version via looking for files “libnvinfer.so” and “libnvinfer_plugin.so” which do exist and point to the latest versions. But, no, in this case they want it explicitly to be version 6 … Makes you wonder whether the old Cudnn version is still available. I did not check it. Ok, ok – backward compatibility is not always possible ….
Just to see how good the internal checking of the alleged dependency is, I did something you normally do not do: I created a link “libnvinfer.so.6” in “/usr/lib64” to “libnvinfer.7.so”. Had to do the same for “libnvinfer_plugin.so.6”. Effect: I got rid of the warning – so much about dependency checking. I left the linking. You see I trust in coming developments sometimes and run some risks ….
Then came the next surprise. I had read a bit about changed statements in TF2 (compared to TF1) – and thought I was prepared for this. But, when I tried to execute some initial commands to configure TF2 from a Jupyter cell as e.g.
import time
import tensorflow as tf
from tensorflow import keras as K
from keras.datasets import mnist
from keras import models
from keras import layers
from tensorflow.python.client import device_lib
import os
#os.environ["CUDA_VISIBLE_DEVICES"] = "-1"
tf.config.optimizer.set_jit(True)
tf.config.threading.set_intra_op_parallelism_threads(4)
tf.config.threading.set_inter_op_parallelism_threads(4)
tf.debugging.set_log_device_placement(True)
device_lib.list_local_devices()
I at once got a complaint in the shell from which I had started the Jupyter notebook – saying that a lib called “libcudart.so.10.1” was missing. Again – an explicit version dependency 🙁 . On purpose or just a glitch? Just one out of many files version dependent? Without a clear information? If this becomes the standard in the interaction between TF2 and Cuda – well, no fun any longer. In my opinion the TF2 developers should not use a search for files via version specific names – but do an analysis of headers and warn explicitly that the present version requires a specific Cuda version. Would be much more convenient for the user and compatible with the link mechanism described above.
Whilst a bunch of other dynamic libs was loaded by their name without a version in this case TF2 asks for a very specific version – although there is a corresponding lib available in the directory “/usr/lib/cuda-10.2″…. Nevertheless with full trust again in a better future
I offered TF2 a softlink “libcudart.so.10.1” in “/usr/lib64/” pointing to the “/usr/local/cuda-10.2/lib64/libcudart.so”. It cleared my way to the next hurdle. And my Keras MLP worked in the end …
Missing “./bin” directory … and other path related problems
When I tried to run specific Keras commands, which TF2 wanted to compile as XLA-supported statements, I again got complaints that files in a local directory “./bin” were missing. This was a first clear indication that Cuda paths were totally ignored in my Python/Jupyter environment. But what directory did the “./” refer to? Some experiments revealed:
I had to link an artificial subdirectory “./bin” in the directory where I kept my Jupyter notebooks to “/usr/local/Cuda-10.2/bin”.
But the next problems with other directories waited directly around the corner. Actually many … To make a long story short – the installation of TF2 in combination with Cuda 1.2 does not evaluate paths or ask for paths when used in a Python3/Jupyter environment. We have to provide and export them as shell environment variables. See below.
Warnings and errors regarding XLA capabilities
Another thing which drove me nuts was that TF2 required information about XLA-flags. It took me a while to find out that this also could be handled via environment variables.
All in all I now start the shell from which I launch my virtual Python environment and Jupyter notebooks with the following command sequence:
myself@mytux:/projekte/GIT/....../ml> export XLA_FLAGS=--xla_gpu_cuda_data_dir=/usr/local/cuda
myself@mytux:/projekte/GIT/....../ml> export TF_XLA_FLAGS=--tf_xla_cpu_global_jit
myself@mytux:/projekte/GIT/....../ml_1> export OPENBLAS_NUM_THREADS=4
myself@mytux:/projekte/GIT/....../ml_1> source bin/activate
(ml) myself@mytux:/projekte/GIT/....../ml_1> jupyter notebook
The first two commands did the magic regarding the path-problems! TF2 worked afterwards both for XLA-capable CPUs and Nvidia GPUs. So, a specific version may or may not have advantages – I do not know – but at least you can get TF2 running with Cuda 10.2.
Changed commands to control threading and memory consumption
Without the use of explicit compatibility commands TF2 does not support commands like
config = tf.ConfigProto(intra_op_parallelism_threads=num_cores,
inter_op_parallelism_threads=num_cores,
allow_soft_placement=True,
device_count = {'CPU' : num_CPU,
'GPU' : num_GPU},
log_device_placement=True
)
config.gpu_options.per_process_gpu_memory_fraction=0.4
config.gpu_options.force_gpu_compatible = True
any longer. But as with TF1 you probably do not want to pick all the memory from your graphics card and you do not want to use all cores of a CPU in TF2. You can circumvent the lack of a “ConfigProto” property in TF2 by the following commands:
# configure use of just in time compiler
tf.config.optimizer.set_jit(True)
# limit use of parallel threads
tf.config.threading.set_intra_op_parallelism_threads(4)
tf.config.threading.set_inter_op_parallelism_threads(4)
# Not required in TF2: tf.enable_eager_execution()
# print out use of certain device (at first run)
tf.debugging.set_log_device_placement(True)
#limit use of graphics card memory
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
try:
tf.config.experimental.set_virtual_device_configuration(gpus[0],
[tf.config.experimental.VirtualDeviceConfiguration(memory_limit=1024)])
except RuntimeError as e:
print(e)
# Not required in TF2: tf.enable_eager_execution()
# print out a list of usable devices
device_lib.list_local_devices()
Addendum, 15.
05.2020:
Well, this actually proved to be correct for the limitation of the GPU memory, only. The limitations on the CPU cores do NOT work. At least not on my system. See also:
tensorflow issues 34415.
I give you a workaround below.
Test run with MNIST
Afterwards the following simple Keras MLP ran without problems and with the expected performance on a GPU and a multicore CPU:
Jupyter cell 1
import time
import tensorflow as tf
#from tensorflow import keras as K
import keras as K
from keras.datasets import mnist
from keras import models
from keras import layers
from keras.utils import to_categorical
from tensorflow.python.client import device_lib
import os
# use to work with CPU (CPU XLA ) only
# os.environ["CUDA_VISIBLE_DEVICES"] = "-1"
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
try:
tf.config.experimental.set_virtual_device_configuration(gpus[0],
[tf.config.experimental.VirtualDeviceConfiguration(memory_limit=1024)])
except RuntimeError as e:
print(e)
# if not yet done elsewhere
tf.config.optimizer.set_jit(True)
tf.debugging.set_log_device_placement(True)
use_cpu_or_gpu = 1 # 0: cpu, 1: gpu
# The following can only be done once - all CPU cores are used otherwise
#if use_cpu_or_gpu == 0:
# tf.config.threading.set_intra_op_parallelism_threads(4)
# tf.config.threading.set_inter_op_parallelism_threads(6)
# function for training
def train(train_images, train_labels, epochs, batch_size):
network.fit(train_images, train_labels, epochs=epochs, batch_size=batch_size)
# setup of the MLP
network = models.Sequential()
network.add(layers.Dense(200, activation='sigmoid', input_shape=(28*28,)))
network.add(layers.Dense(100, activation='sigmoid'))
network.add(layers.Dense(50, activation='sigmoid'))
network.add(layers.Dense(30, activation='sigmoid'))
network.add(layers.Dense(10, activation='sigmoid'))
network.compile(optimizer='rmsprop', loss='categorical_crossentropy', metrics=['accuracy'])
# load MNIST
mnist = K.datasets.mnist
(X_train, y_train), (X_test, y_test) = mnist.load_data()
# simple normalization
train_images = X_train.reshape((60000, 28*28))
train_images = train_images.astype('float32') / 255
test_images = X_test.reshape((10000, 28*28))
test_images = test_images.astype('float32') / 255
train_labels = to_categorical(y_train)
test_labels = to_categorical(y_test)
Jupyter cell 2
# run it
if use_cpu_or_gpu == 1:
start_g = time.perf_counter()
train(train_images, train_labels, epochs=45, batch_size=1500)
end_g = time.perf_counter()
test_loss, test_acc= network.evaluate(test_images, test_labels)
print('Time_GPU: ', end_g - start_g)
else:
start_c = time.perf_counter()
train(train_images, train_labels, epochs=45, batch_size=1500)
end_c = time.perf_counter()
test_loss, test_acc= network.evaluate(test_images, test_labels)
print('Time_CPU: ', end_c - start_c)
# test accuracy
print('Acc: ', test_acc)
Switch to force Tensorflow to use the CPU, only
Another culprit is that – depending on the exact version of TF 2 – you may need to use the following statement to run (parts of) your code on the CPU only:
os.environ["CUDA_VISIBLE_DEVICES"] = "-1"
in the beginning. Otherwise Tensorflow 2.0 and version 2.1 will choose execution on the GPU even if you use a statement like
with tf.device("/CPU:0"):
(which worked in TF1).
It seems that this problem was solved with TF 2.2 (tested it on 15.05.2020)! But you may have to check it yourself.
You can
watch the involvement of the GPU e.g. with “watch -n0.1 nvidia-smi” on a terminal. Another possibility is to set
tf.debugging.set_log_device_placement(True)
and get messages in the shell of your virtual Python environment or in the presently used Jupyter cell.
Addendum 16.05.2020: Limiting the number of CPU cores for Tensorflow 2.0 on Linux
After several trials and tests I think that both TF2 and the Keras version delivered with handle the above given TF2 statements to limit the number of CPU cores to use inefficiently. I addition the behavior of TF2/Keras has changed with the TF2 versions 2.0, 2.1 and now 2.2.
Strange things also happen, when you combine statements of the TF1 compat layer with pure TF2 restriction statements. You should refrain from mixing them.
So, it is either
Option 1: CPU only and limited number of cores
from tensorflow import keras as K
from tensorflow.python.keras import backend as B
import os
os.environ["CUDA_VISIBLE_DEVICES"] = "-1"
...
config = tf.compat.v1.ConfigProto(intra_op_parallelism_threads=4, inter_op_parallelism_threads=1)
B.set_session(tf.compat.v1.Session(config=config))
...
OR
Option 2: Mixture of GPU (with limited memory) and CPU (limited core number) with TF2 statements
import tensorflow as tf
from tensorflow import keras as K
from tensorflow.python.keras import backend as B
from keras import models
from keras import layers
from keras.utils import to_categorical
from keras.datasets import mnist
from tensorflow.python.client import device_lib
import os
tf.config.threading.set_intra_op_parallelism_threads(6)
tf.config.threading.set_inter_op_parallelism_threads(1)
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
try:
tf.config.experimental.set_virtual_device_configuration(gpus[0],
[tf.config.experimental.VirtualDeviceConfiguration(memory_limit=1024)])
except RuntimeError as e:
print(e)
OR
Option 3: Mixture of GPU (limited memory) and CPU (limited core numbers) with TF1 compat statements
import tensorflow as tf
from tensorflow import keras as K
from tensorflow.python.keras import backend as B
from keras import models
from keras import layers
from keras.utils import to_categorical
from keras.datasets import mnist
from tensorflow.python.client import device_lib
import os
#gpu = False
gpu = True
if gpu:
GPU = True; CPU = False; num_GPU = 1; num_CPU = 1
else:
GPU = False; CPU = True; num_CPU = 1; num_GPU = 0
config = tf.compat.v1.ConfigProto(intra_op_parallelism_threads=6,
inter_op_parallelism_threads=1,
allow_soft_placement=True,
device_count = {'CPU' : num_CPU,
'GPU' : num_GPU},
log_device_placement=True
)
config.gpu_options.per_process_gpu_memory_fraction=0.35
config.gpu_options.force_gpu_compatible = True
B.set_session(tf.compat.v1.Session(config=config))
Hint 1:
If you want to test some code parts on the GPU and others on the CPU in the same session, I strongly recommend to use the compat statements in the form given by Option 3 above
The reason is that it – strangely enough – gives you a faster performance on a multicore CPU by more than 25% in comparison to the pure TF2 statements .
Afterwards you can use statements like:
batch_size=64
epochs=5
if use_cpu_or_gpu == 0:
start_g = time.perf_counter()
with tf.device("/GPU:0"):
train(train_imgs, train_labels, epochs, batch_size)
end_g = time.perf_
counter()
print('Time_GPU: ', end_g - start_g)
else:
start_c = time.perf_counter()
with tf.device("/CPU:0"):
train(train_imgs, train_labels, epochs, batch_size)
end_c = time.perf_counter()
print('Time_CPU: ', end_c - start_c)
Hint 2:
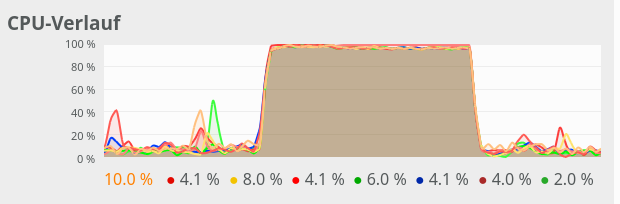
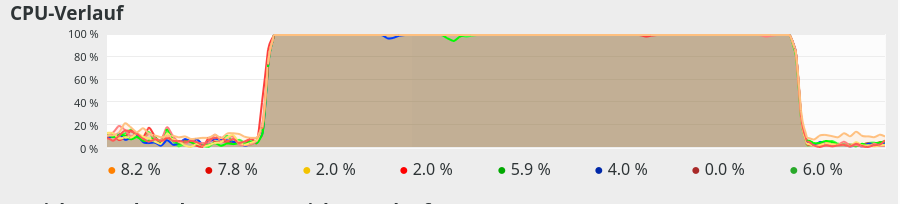
If you check the limitations on CPU cores (threads) via watching the CPU load on tools like “gkrellm” or “ksysguard”, it may appear that all cores are used in parallel. You have to set the update period of these tools to 0.1 sec to see that each core is only used intermittently. In gkrellm you should also see a variation of the average CPU load value with a variation of the parameter “intra_op_parallelism_threads=n”.
Hint 3:
In my case with a Quadcore CPU with hyperthreading the following settings seem to be optimal for a variety of Keras CNN models – whenever I want to train them on the CPU only:
...
config = tf.compat.v1.ConfigProto(intra_op_parallelism_threads=6, inter_op_parallelism_threads=1)
B.set_session(tf.compat.v1.Session(config=config))
...
Hint 4:
If you want to switch settings in a Jupyter session it is best to stop and restart the respective kernel. You can do this via the commands under “kernel” in the Jupyter interface.
Conclusion
Well, my friends the above steps where what I had to do to get Keras working in combination with TF2, Cuda 10.2 and the present version of Cudnn. I regard this not as a straightforward procedure – to say it mildly.
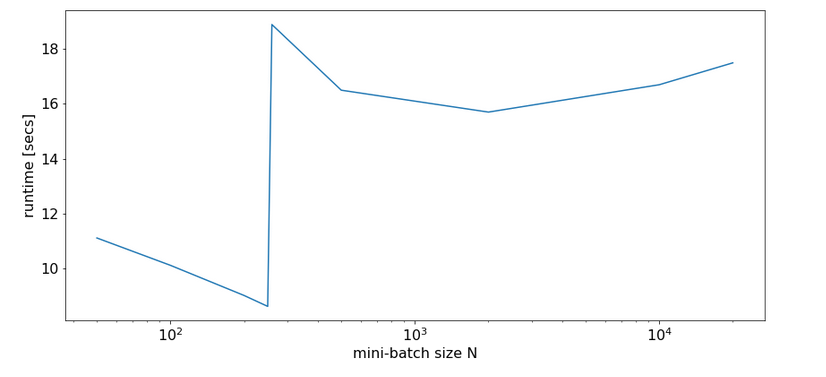
In addition after some tests I might also say that the performance seems to be worse than with Tensorflow 1. Especially when using Keras – whether the Keras included with Tensorflow 2 or Keras in form of separate Python lib. Especially the performance on a GPU is astonishingly bad with Keras for small networks.
This impression of sluggishness stands in a strange contrast to elementary tests were I saw a factor of 5 difference for a series of typical matrix multiplications executed directly with tf.matmul() on a GPU vs. a CPU. But this another story …..
Links
tensorflow-running-version-with-cuda-on-cpu-only