This post requires Javascript to display formulas!
Sometimes when we solve some problems with Deep Neural Networks we have no direct clue about how such a network evolves during training and how exactly it encodes knowledge learned from patterns in the input data. The encoding may not only affect weights of neurons in neural filters or maps, but also the functional mapping of input data onto vectors of some intermediate or target vector space. To get a better understanding it may sometimes seem that experiments with statistical points or vectors in such vector spaces could be helpful. One could e.g. suggest to study the reaction of a trained Artificial Neural Network to input vectors distributed over a multidimensional feature space.
Another important case is the test of an Autoencoder’s Decoder network on data points which we distribute statistically over extended regions of the Autoencoder’s latent space.
An Autoencoder [AEs] is an example of a Encoder-Decoder pair of networks. The Encoder encodes original complex input data into vectors of the so called latent space – a multi-dimensional vector space. The Decoder instead uses vectors of the latent space to reconstruct or generate output data in the feature space of the original input objects. The dimensionality of the latent space may be much smaller than that of the original feature space, but still much bigger than the 2 or 3 dimensions we have intuitive experience with.
Latent spaces play a major role in Machine Learning [ML] whenever a deep neural algorithm encodes information. Embedding input data (like words or images) in a vector space by some special neural layers (e.g. for text or image classification) is nothing else than filling a kind of latent space with encoded data in an optimized way to help the neural network to solve a special task like e.g. text or image classification. But we can also use statistical data in the latent space of an AE together with its trained Decoder to create or better generate new objects. This leads us into the field of using neural networks for generative, creative tasks.
Unfortunately, we know from experience that not all data regions in a latent space support the production of objects which we would consider as being similar to our training objects or categories of such objects. A priori we actually know very little about the shape of the data distribution an encoding network will create during training. Therefore we may be tempted to scan the latent space in some way – and to distribute data points in it in some statistical way. And test a Decoder’s reaction to such points …in the hope to find an interesting region by chance.
In this small series of posts I want to point out that such statistical experiments may be futile. On the one hand side trying to fill a multidimensional space with “statistical” data may lead to unexpected and counter-intuitive results. On the other hand side you may easily miss interesting regions of such a vector space. Especially confined regions. The whole point is just another interesting variant of the curse of high dimensions.
The theoretical and numerical results which we will derive during the next posts underline the necessity to analyze the concrete data distributions a neural network creates in a latent space very carefully. Both regarding position and extension of the filled region – and maybe also with respect to sub-structures. The results will help us to understand better why an Autoencoder most often fails to produce reasonable objects from data points statistically distributed over larger regions of its latent space.
This post series requires an understanding of some basic math. In the first post I will derive some formulas describing data distributions resulting from a constant probability density for each vector component, i.e. a constant probability density per dimension. We will learn that with a growing number of dimensions the data points concentrate more and more in regions which are parts of a multi-dimensional sphere shell. Which is at least in my opinion a bit counter-intuitive and therefore an interesting finding in itself.
Many dimensions …
An AE compresses data very effectively. So the dimension of a latent space will most often be significantly smaller than the number of dimensions M characterizing the feature space of the original training objects. The space spanned by logically independent variables which describe digitized objects may have a number of dimensions in the range of millions and tenths of millions. In the case of latent spaces we instead have to deal with a number of dimensions N in the range of multiples of 10 up to tens of thousands. (For relatively simple objects showing many correlations in their feature space …)
10 ≤ N ≤ 105 << M
Still, such numbers N are far beyond standard experience.
Representation of a N-dimensional space by the ℜN
We can in general represent a multidimensional feature or latent space in Machine Learning by the ℜN. Each point in this space can be defined by a vector whose components correspond to the points’ coordinate values along the N orthogonal axes. A vector is thus represented by a N-tuple of real (coordinate) values:
(x1, x2, …, xk, … xN) .
Formally, the set of all tuples together with some addition and multiplication operations form a vector space over ℜ. Whether vector operations performed with vectors to different data points in a latent space or a feature space really have a solid interpretable meaning in your ML- or AE-context is not our business in this post. Our objective is to cover such a space with statistically defined points (or vectors).
The curse of high dimensionality
It seems to be easy to fill a N-dimensional cube with data points that form a grid. More precisely, a grid with a constant distance between the points in all orthogonal directions parallel to the axes. However, a simple calculation shows that such an approach is doomed in computational practice:
Let us assume that we deal with a space of N = 256 dimensions. Let us also assume that we have reasons to believe that we should focus on the region of a cube around the origin that has a side length of 10 in each dimension k. Then the coordinate values xk for data points within the cube fulfill
\]
Let us further assume that we want to have a grid resolution of 10 points per dimension. Then we would need to create
\]
to fill our 256-dimensional cube with a homogeneous (!) point distribution. This is already a very clear indication that we can not systematically study a multidimensional cube in this way. Multidimensional spaces are huge by their nature.
We have to get and use more information about interesting regions or we must make assumptions on the distribution of points in each coordinate direction.
Constant probability density per coordinate value and dimension
A most simple case for making an assumption is a constant probability distribution of each coordinate value xk within a defined interval along the related coordinate axis. Let us again pick a cube around the origin of a space with N=256:
Let us assume that we only want a resolution of 100 equidistant values per side length of the cube. From a statistical point of view we deal with our dimensions like they were part of a classical experiment in which we pick balls with imprinted values from a (mixing) black box. In our assumed case we have a box with 100 numbers and fetch a ball 256 times to create a 256-tuple for each vector – of course with laying back the picked ball each time. We will come back to this interpretation in a second post. For now let us define a constant probability density and see where we get from there mathematically.
The normalized density distribution in the interval [-b, b] simply is
One may “naively” think the following:
We statistically fetch 256 xk-values, each from our uniform distribution, and fill our vector N-tuples with these values. Repeating this vector generation process for e.g. 200000 times should give us some widespread point distribution covering our cube very well. Let me warn you:
This is a common mistake – although such a procedure seems to work rather well in 2 or 3 dimensions!
It admit: The whole thing is somewhat counter-intuitive. If you create a homogeneous distribution for each of the coordinate values xk then the points should well cover the whole space? The answer unfortunately is: No.
To get a better understanding we need some more math. And we first reduce the problem to statements about a one-dimensional quantity, namely the vector length. But let us first look at results in 3 dimensions.
Point distribution in a 3-dimensional cube for a constant probability density of each coordinate value
Let us create some 200 points in a cube
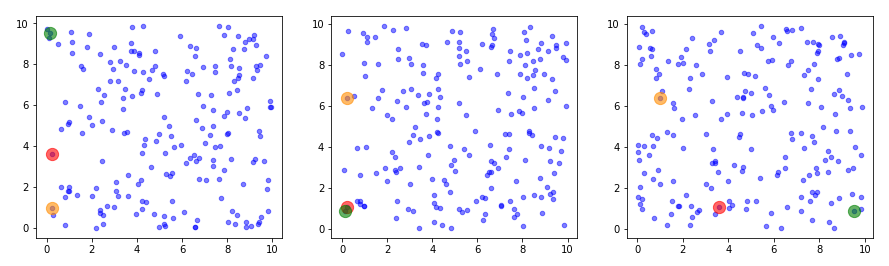
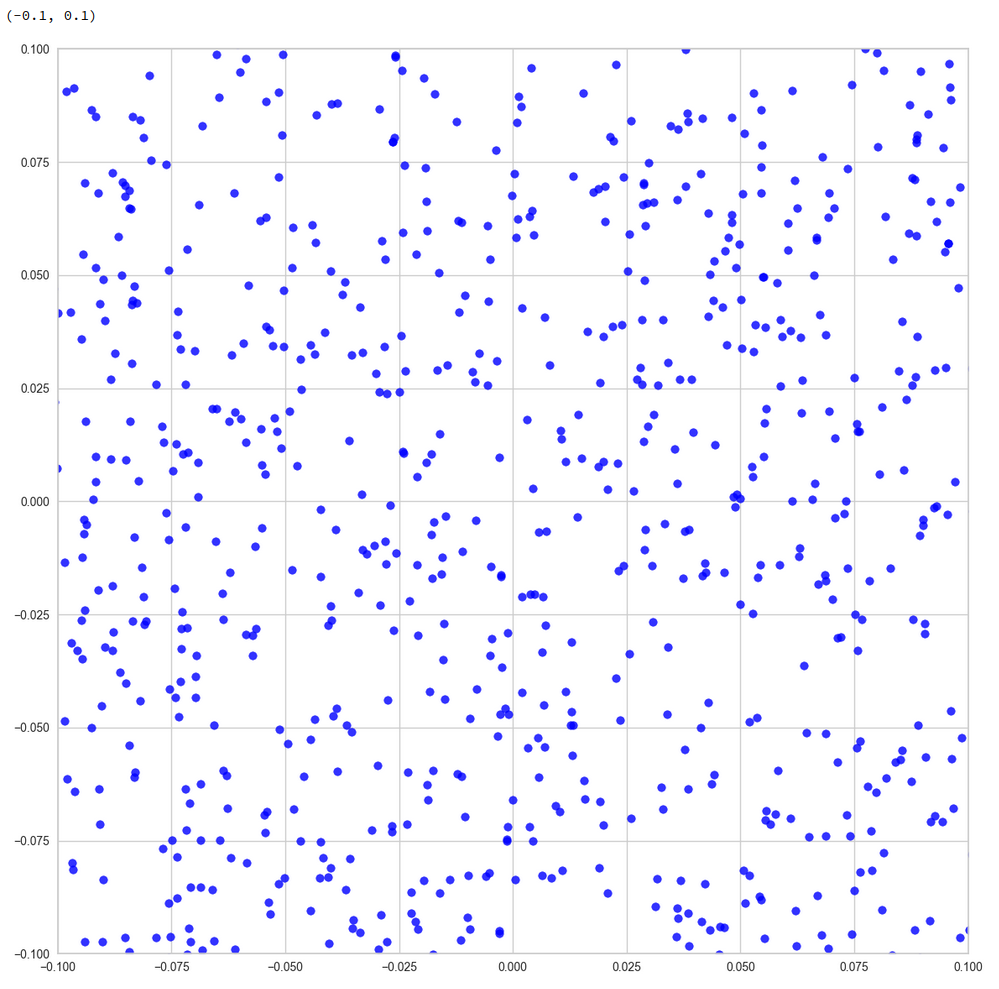
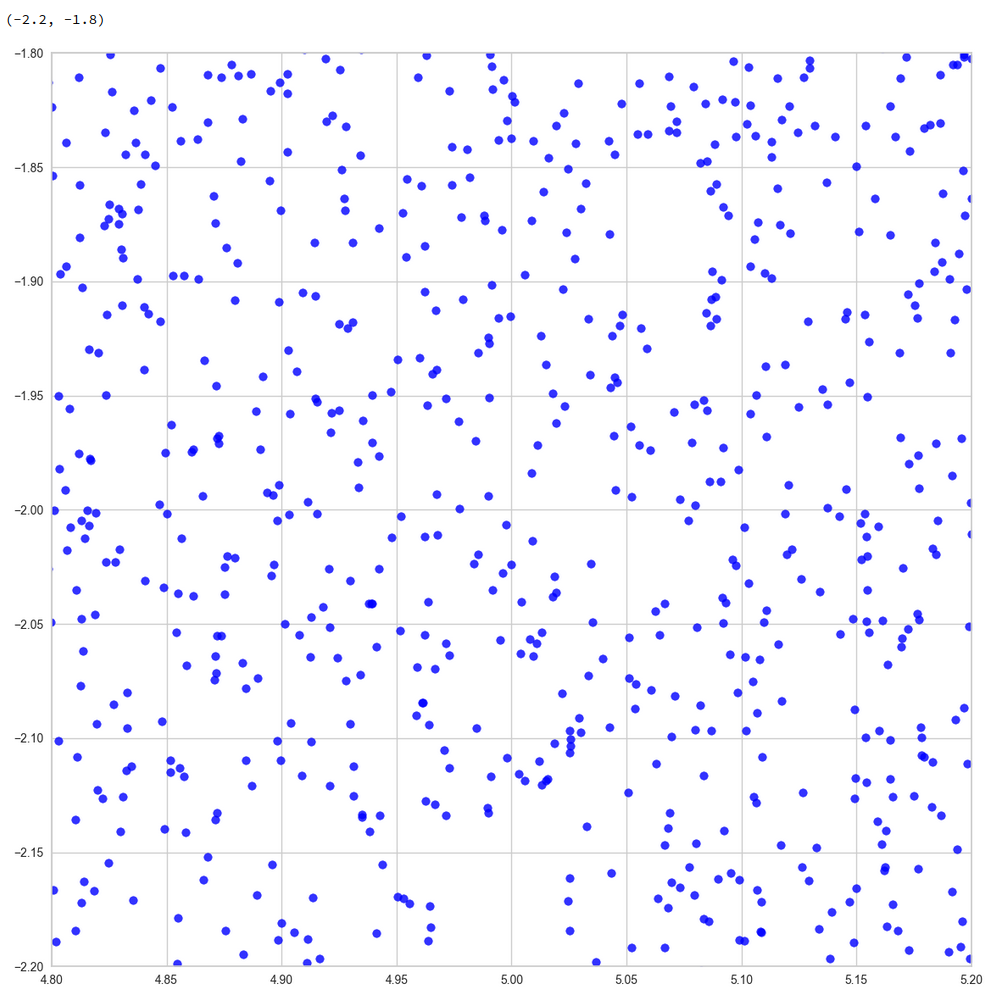
The following plots show the distributions projections on the planes between the coordinate axes:
This looks quite OK. All areas seem to be covered. I have marked some points to illustrate their positions. We see that filling a corner in 2D does not necessarily mean that the points are close to the origin of the 3D coordinate system. Now, let us look at a 3D-plot:
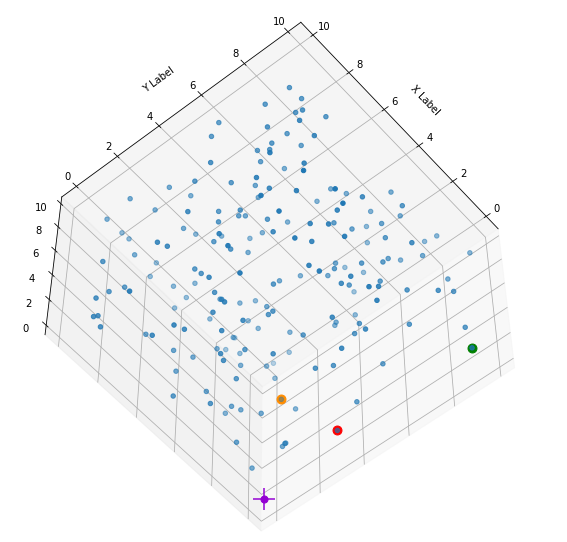
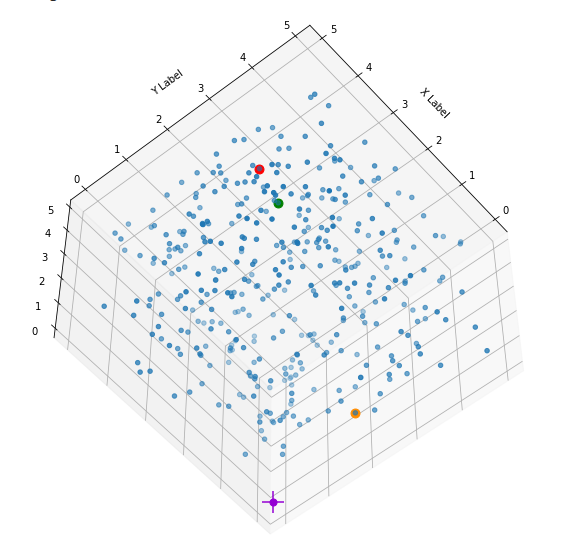
I have marked the origin in violet. Notably, extended elongated stripes at the edges with two coordinate close to extreme values seem strangely depleted. This is no accident. Below you find a similar plot with 400 points and b=5:
What is the reason? Well, it is simply statistics: To fill the special regions named above you have to fix one or two coordinates at the same time. Let us look at a stripe stretching from one corner and parallel to an axis and perpendicular to the plane {x_1, x_2}:
As we deal with a uniform distribution per coordinate value the chances are 1/10 * 1/10 = 1/100 that a randomly constructed point will fulfill these conditions.
Also: For a region with all coordinates around [0,1] the chance is just 0.13 = 0.001.
When see already that the probability to get a point with a limited radius vector becomes rather small. Now, what happens in a real multi-dimensional space, e.g. with N=256?
The mean radius of data points created with a constant probability density for each coordinate value
Let us call the length of a vector directed from the origin to some defined point in our cube the “radius” R of the vector. This radius then is defined as:
What is the length of a vector with all xk = 1 for all k? For N=256 dimensions the answer is:
So, we should not be surprised when we have to deal with large average numbers for vector lengths in the course of our analysis below. We again assume a constant probability density ρ along the each coordinate axis in the interval [-b, b]
Now, what will the mean radius of a “random” point distribution in the cube be when we base the distribution on this probability density? How do we get the expectation value <R> for the length of our vectors?
If we wanted to approach the problem directly we would have to solve an integral over the vector length and a (normalized) probability density. This would lead us to integrals of the type
As our normalized probability density per dimension is a constant, the whole thing looks simple. However, the integrals over the square root are, unfortunately, not really trivial to handle. Try it yourself …
Therefore, I choose a different way: Instead of looking for the expectation value of the vector length <R> I look for the square root of the expectation value of the vector length squared R2 – and assume:
We take care of this assumption later by giving a reference and an approximation formula.
We also use the fact that the expectation value is a linear function for a sum of independent quantities. This leads us to
The resulting integral is much easier to solve. It splits up into simple 1-dimensional integrals. Note: Each of your squared coordinate values (xk)2 is associated with a simple, un-squared (!) probability density of 1/(2b) for an interval [-b, b] or 1/(b-a) for [a, b].
The result for a general common interval [a,b] for all coordinate values a \le; xk ≤ b is :
\]
So, for our special case a = -b we get:
\]
This will give us for b = 5:
Note that
Ok, the mean length value is a bit bigger than half of the length which a diagonal vector from the origin to the outmost edge of our cube (b, b,,…,b) would have. This does not say so much about the point and the radius distributions themselves. We have to look at higher momenta. Let us first look at the variance and standard variation for R2.
Variance and standard deviation of R2
What we actually would like to see is the variance and standard deviation for R. But again it is easier to first solve the problem for the squared quantity. The variance of R2 is defined as:
\]
\]
We can solve this step by step. First we use:
\]
We evaluate (for our cube):
\]
and
\]
For the different contributions to the expectation values we get
\]
Integrating over our constant probability densities for the independent coordinates gives
\]
Thus
\]
Eventually, we get for the ratio of the standard variation of R2 to <R2> :
\]
The ratio of the standard variation of R to <R>
We are basically interested in the ratio
\]
Before you start to fight the complex integrals over square roots think of the following:
\]
From an adaption of this argument we conclude
\]
A very simple relation! It shows that on average the interval for the radius spread systematically becomes smaller in comparison to the average radius with a growing number of dimensions. In the case of N=256 the relative radius spread is only of the order of some percent.
\]
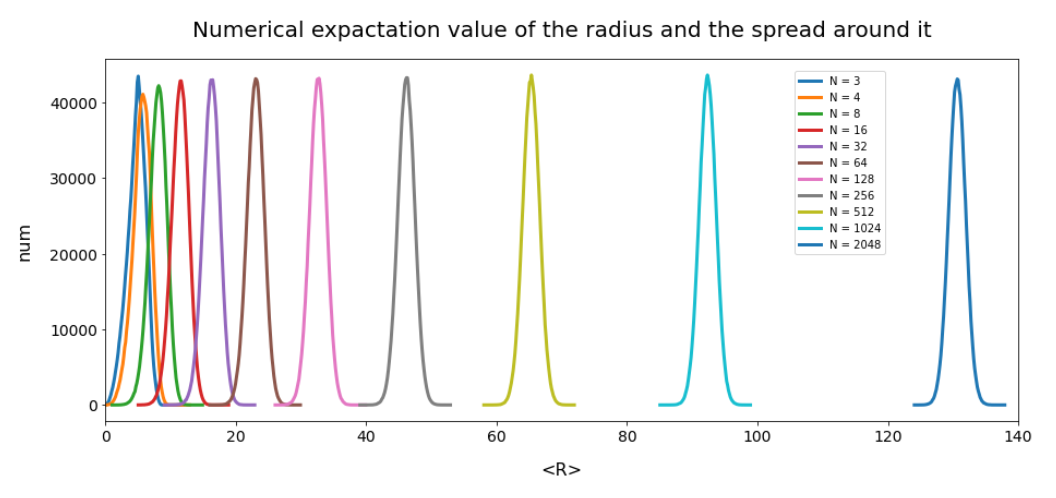
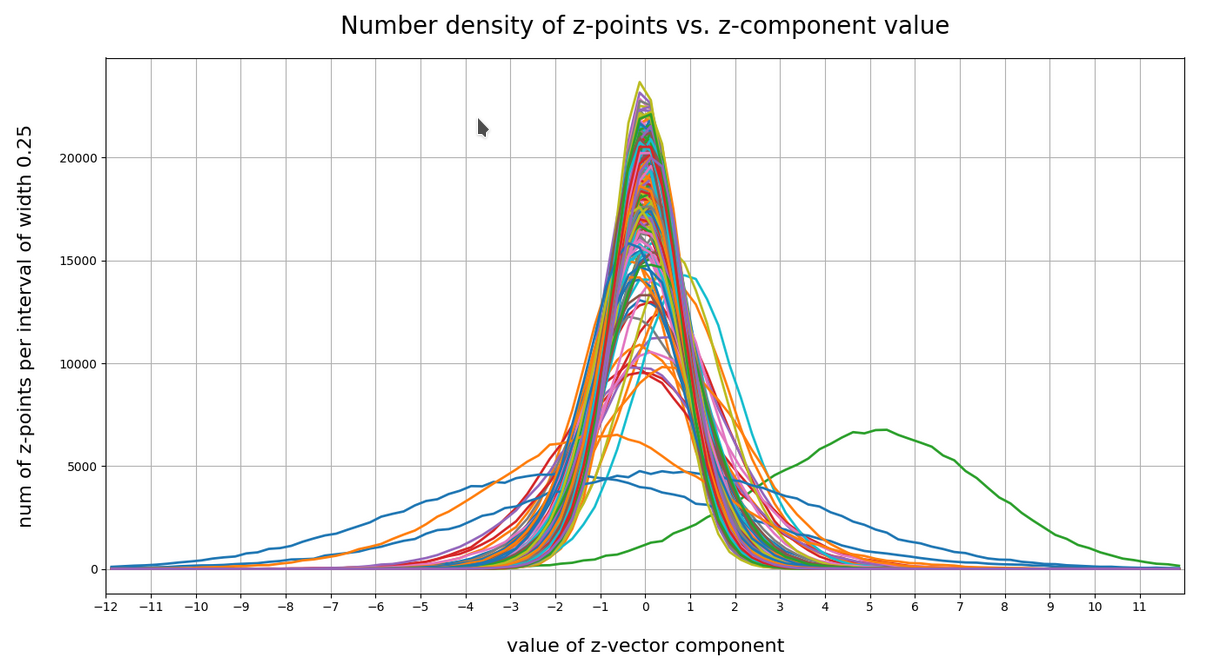
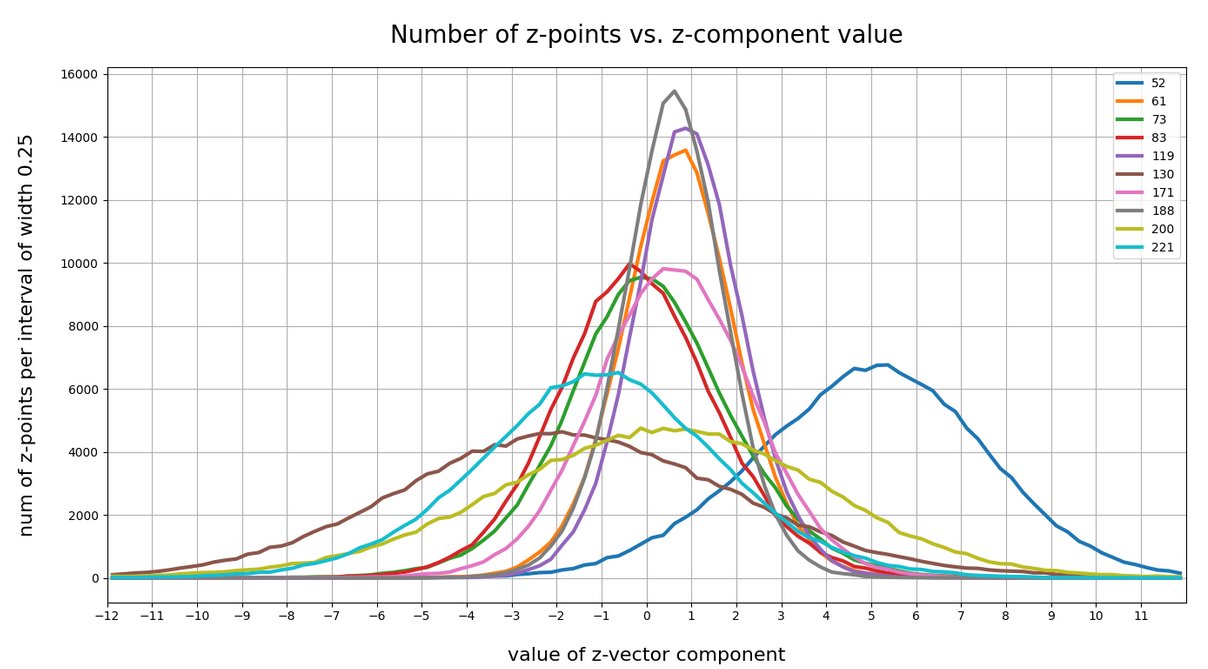
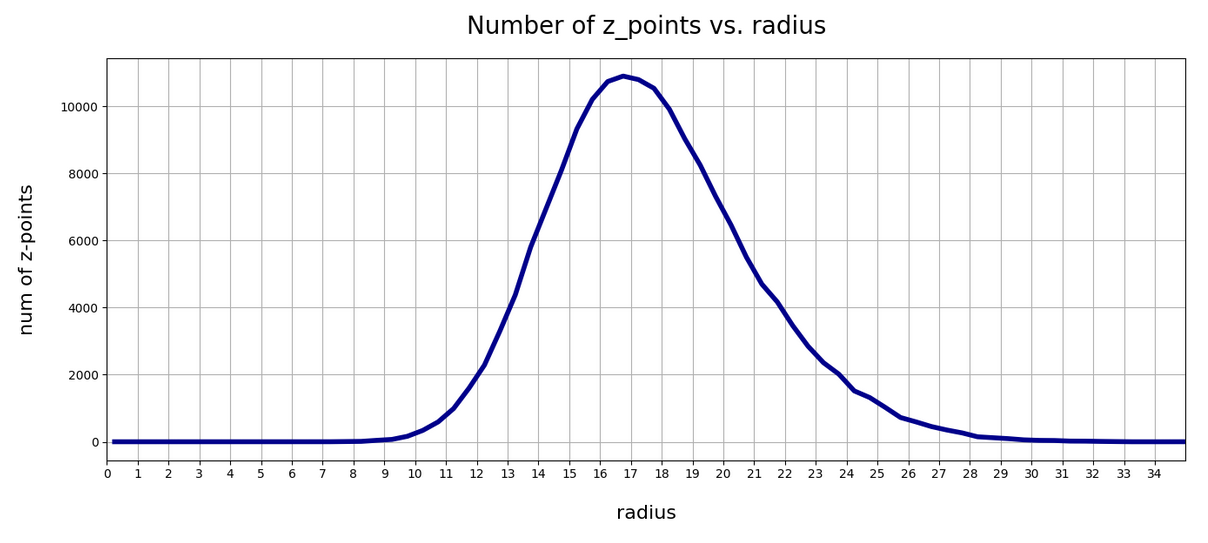
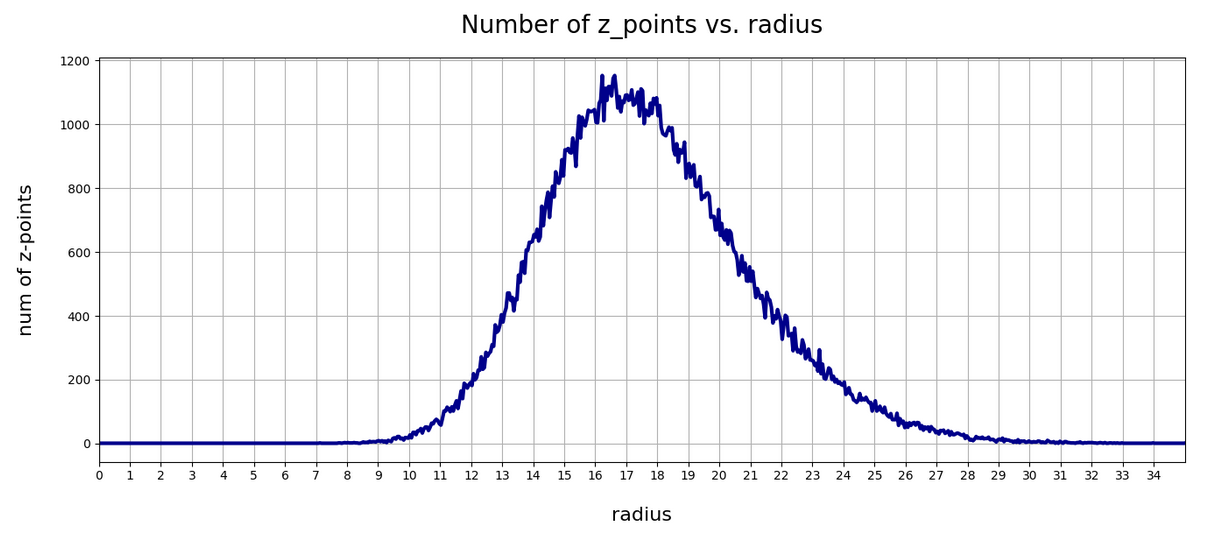
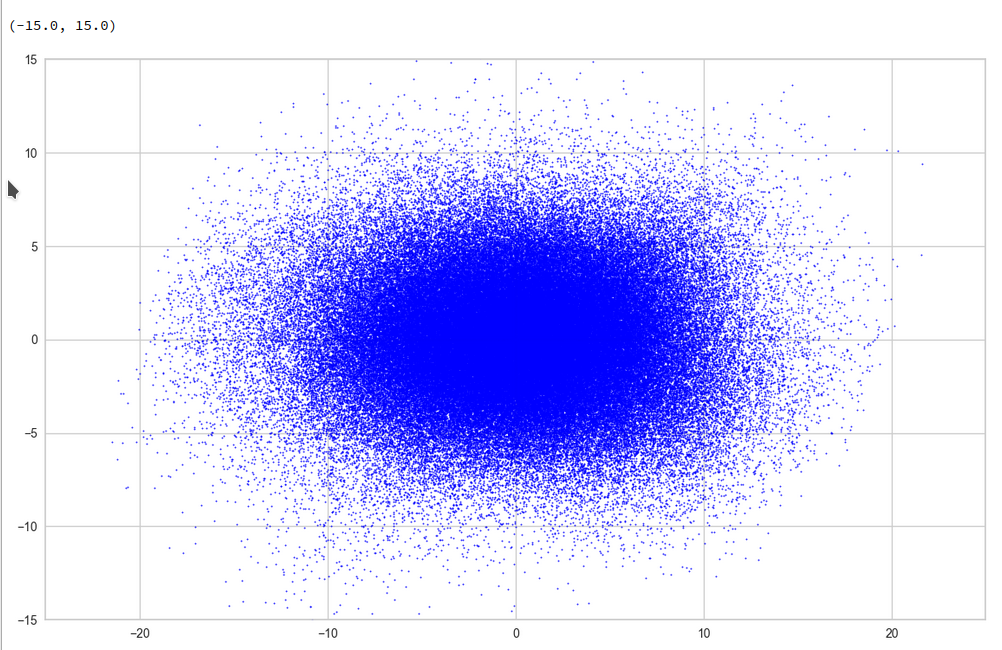
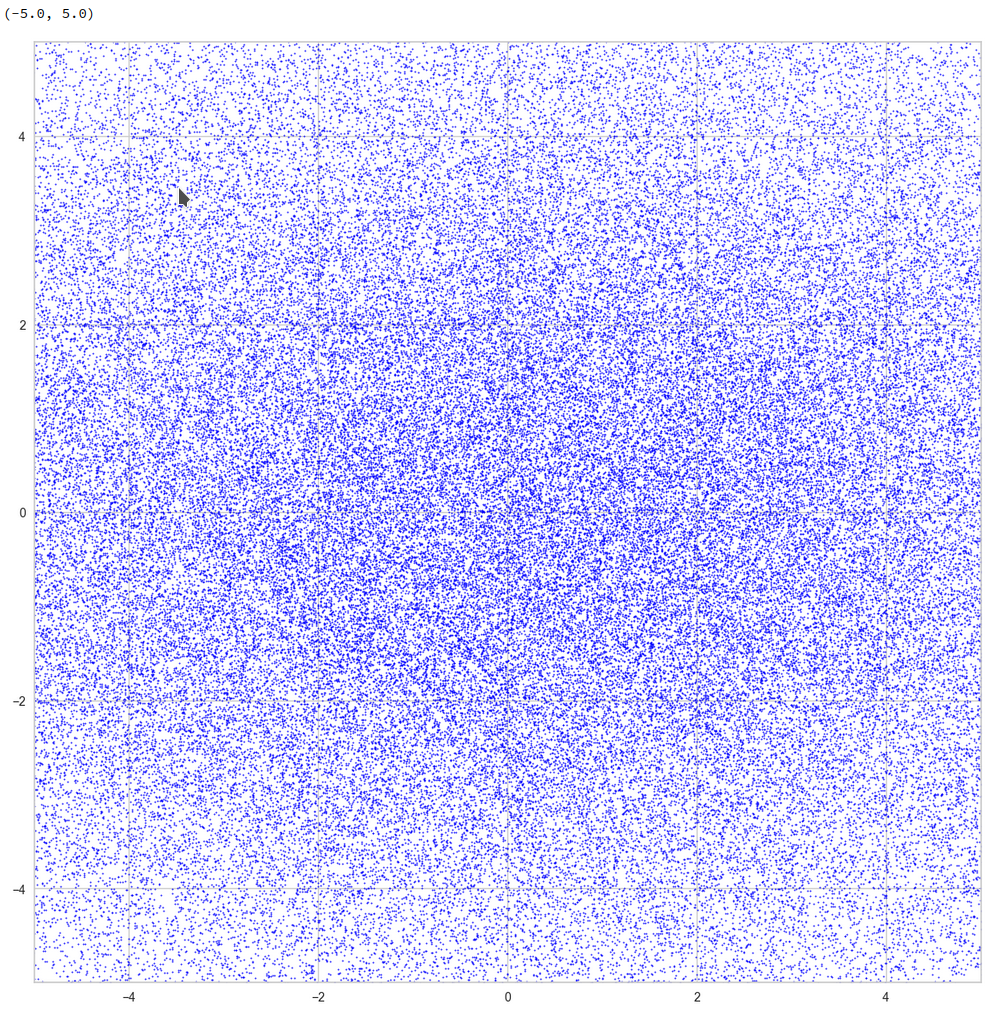
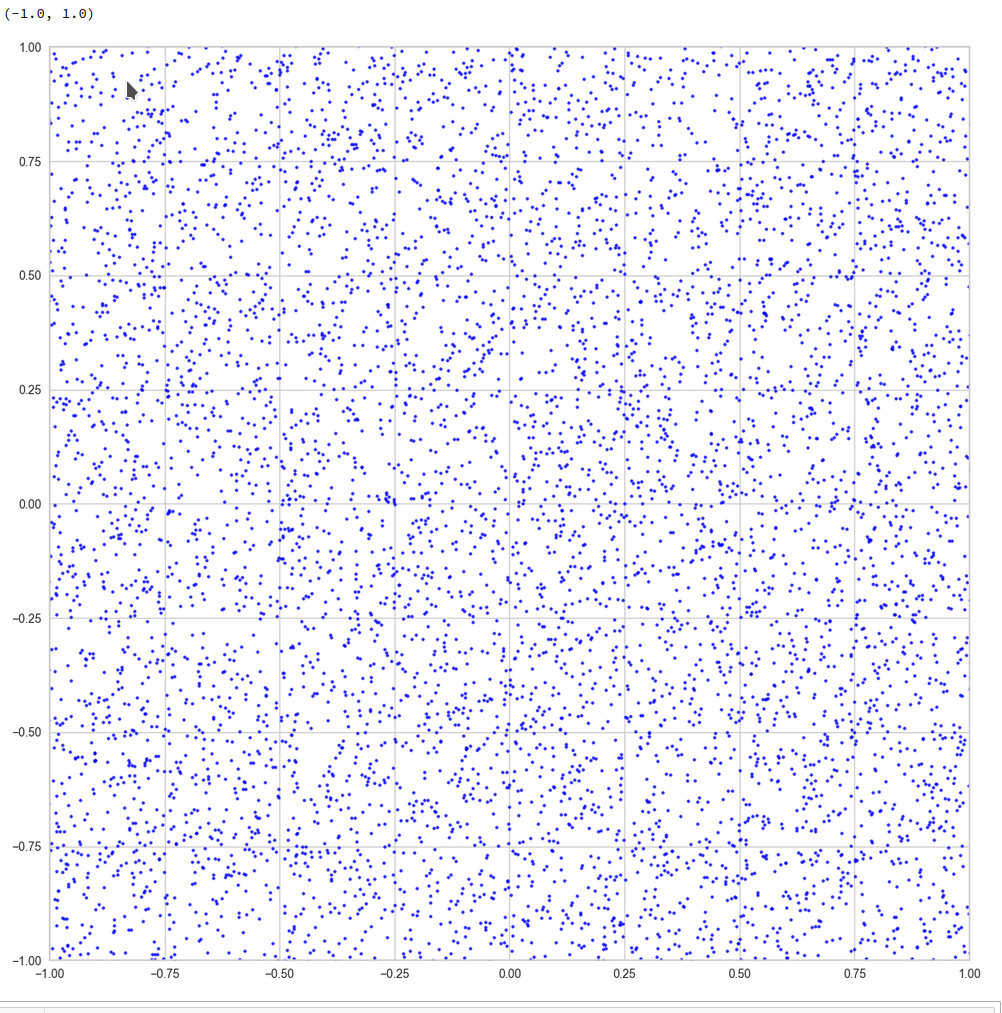
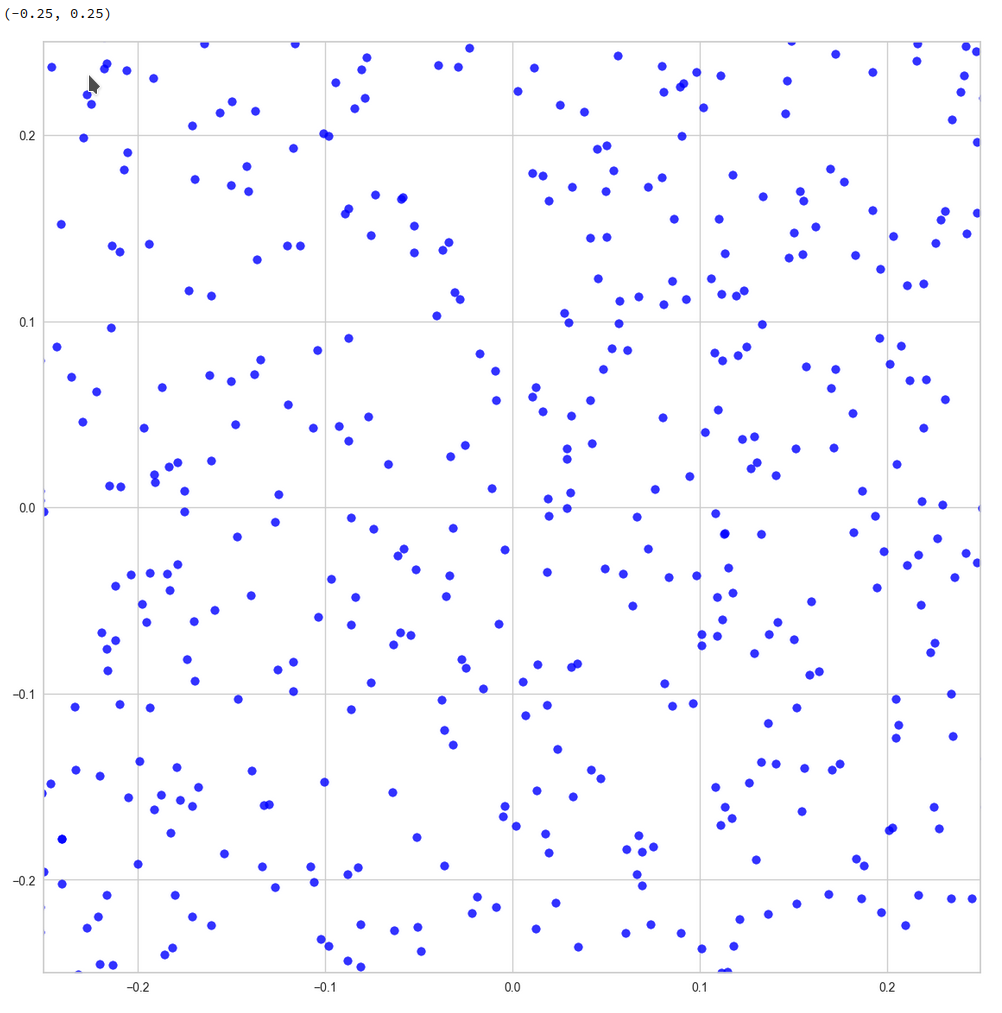
This defines a region or regions within a very narrow multi-dimensional spherical shell. The plot below shows the result of a numerical simulation based on one million data points for dimensions between 3 ≤ N ≤ 2048. You see the radius < R > on the x-axis, the number of points on the y-axis and the spread around the mean radius for the selected numbers of dimensions. Although the spread and the standard deviation remain almost constant the radius value is steadily rising.
Note that one should be careful not to assume more than a point concentration in regions of a multi-dimensional spherical shell. Even within such a shell the created data points may avoid certain regions. I will give some arguments that support this point of view in the next post. But already now it has become very clear that a statistical distribution of vectors with constant probabilities of component values per dimension will not at all fill the latent space homogeneously.
Some better approximations for <R> and the relative standard deviation
The above formulas suffer from the fact that
\]
is only a rough approximation which overestimates the real $lt; R > – value, which comes out of numerical simulations. The deviation rises sharply with a small number of dimensions. For a thorough discussion of how good the approximation is see the answer and links at the following discussion at stackexchange
https://stats.stackexchange.com/ questions/ 317095/ expectation-of-square-root-of-sum-of-independent-squared-uniform-random-variable
Below I give you a better approximation to < R >, which should be sufficient for most purposes and N ≥ 3:
\]
This approximation can in turn be used to improve the ratio of the standard deviation to < N gt;:
\]
Conclusion and outlook on the next post
In this post I have tried to fill a cube around the origin of a multi-dimensional orthogonal coordinate system with statistical points. I have assumed a constant probability density for each component of the respective vectors. A simple mathematical analysis leads to the somewhat surprising conclusion that the resulting data distribution concentrates in regions which are located within a multi-dimensional spherical shell. This shell becomes narrower and narrower with a rising number of dimensions N.
In the next post
Latent spaces – pitfalls of distributing points in multi dimensions – II – missing specific regions
I shall first prove the derived results and given approximations by numerical data. Then I will discuss what kind of regions, that a Neural Network might fill in its latent space, we will miss by such an approach.