I continue with my series on the treatment of the KL loss of Variational Autoencoders in a Keras / TF2.8 environment:
Variational Autoencoder with Tensorflow – I – some basics
Variational Autoencoder with Tensorflow – II – an Autoencoder with binary-crossentropy loss
Variational Autoencoder with Tensorflow – III – problems with the KL loss and eager execution
Variational Autoencoder with Tensorflow – IV – simple rules to avoid problems with eager execution
In the last post it became clear that it might be a good idea to delegate the KL loss calculation to a specific layer within the Encoder model. In this post I discuss the code for such a solution. I am going to encapsulate the construction of a suitable Keras model for the VAE in a class. The class will in further posts be supplemented by more methods for different approaches compatible with TF2.x and eager execution.
The code’s structure has been influenced by the work or books of several people which I want to name explicitly: D. Foster, F. Chollet and Louis Tiao. See the references in the last section of this post.
For the data sets I later want to work with both the Encoder and the Decoder parts of the VAE shall be based upon “convolutional networks” [CNNs] and respective Keras layers. Based on a suggestions of D. Foster and F. Chollet I use a classes interface to provide the parameters of all invoked Conv2D and Conv2DTranspose layers. But in contrast to D. Foster I also indicate how to include different activation functions (e.g. SeLU). In general I also will use the Keras functional API to define and add layers to the VAE model.
Imports to make Keras model and layer classes work
Below I discuss step by step parts of the code I put into a Python module to be used later in Jupyter notebooks. First we need to import some Python modules; note that you may have to add further statements which import personal modules from paths at your local machine:
import sys
import numpy as np
import os
import tensorflow as tf
from tensorflow.keras.layers import Layer, Input, Conv2D, Flatten, Dense, Conv2DTranspose, Reshape, Lambda, \
Activation, BatchNormalization, ReLU, LeakyReLU, ELU, Dropout, AlphaDropout
from tensorflow.keras.models import Model
# to be consistent with my standard loading of the Keras backend in Jupyter notebooks:
from tensorflow.keras import backend as B
from tensorflow.keras.optimizers import Adam
A class for a special Encoder layer
Following the ideas discussed in my last post I now add a class which later allows for the setup of a special customized Keras layer in the Encoder model. This layer will calculate the KL loss for us. To be able to do so, the implementation interface “call()” receives a variable “inputs” which contains references to the mu and var_log layers of the Encoder (see the two last posts in this series).
class My_KL_Layer(Layer):
'''
@note: Returns the input layers ! Required to allow for z-point calculation
in a final Lambda layer of the Encoder model
'''
# Standard initialization of layers
def __init__(self, *args, **kwargs):
self.is_placeholder = True
super(My_KL_Layer, self).__init__(*args, **kwargs)
# The implementation interface of the Layer
def call(self, inputs, fact = 4.5e-4):
mu = inputs[0]
log_var = inputs[1]
# Note: from other analysis we know that the backend applies tf.math.functions
# "fact" must be adjusted - for MNIST reasonable values are in the range of 0.65e-4 to 6.5e-4
kl_mean_batch = - fact * B.mean(1 + log_var - B.square(mu) - B.exp(log_var))
# We add the loss via the layer's add_loss() - it will be added up to other losses of the model
self.add_loss(kl_mean_batch, inputs=inputs)
# We add the loss information to the metrics displayed during training
self.add_metric(kl_mean_batch, name='kl', aggregation='mean')
return inputs
An important point is that a layer based on this class must return its input, namely the mu and var_log layers, for the z-point calculations in the final Encoder layer.
Note that we do not only add the loss to other losses of an eventual VAE model via the layer’s “add_loss()” method, but that we also ensure to get some information about the the size of the KL loss during training by adding the loss to the metrics.
A general class to setup a VAE build on CNNs for Encoder and Decoder
We now build a class to create the essential parts of a VAE. The class will provide the required flexibility and allow for future extensions comprising other TF2.x compatible solutions for KL loss calculations. (In this post we only use a customized layer to get the KL loss).
We start with the classes “__init__” function, which basically transfers saves parameters into class variables.
# The Main class
# ~~~~~~~~~~~~~~
class MyVariationalAutoencoder():
'''
Coding suggestions of D. Foster and F. Chollet were modified and extended by RMO
@version: V0.1, 25.04
@change: added b_build_all
@version: V0.2, 08.05
@change: Handling of the KL-loss via functions (partially not working)
@version: V0.3, 29.05
@change: Handling of the KL-loss function via a customized Encoder layer
'''
def __init__(self
, input_dim # the shape of the input tensors (for MNIST (28,28,1))
, encoder_conv_filters # number of maps of the different Conv2D layers
, encoder_conv_kernel_size # kernel sizes of the Conv2D layers
, encoder_conv_strides # strides - here also used to reduce spatial resolution avoid pooling layers
# used instead of Pooling layers
, decoder_conv_t_filters # number of maps in Con2DTranspose layers
, decoder_conv_t_kernel_size # kernel sizes of Conv2D Transpose layers
, decoder_conv_t_strides # strides for Conv2dTranspose layers - inverts spatial resolution
, z_dim # A good start is 16 or 24
, solution_type = 0 # Which type of solution for the KL loss calculation ?
, act = 0 # Which type of activation function?
, fact = 0.65e-4 # Factor for the KL loss (0.5e-4 < fact < 1.e-3is reasonable)
, use_batch_norm = False # Shall BatchNormalization be used after Conv2D layers?
, use_dropout = False # Shall statistical dropout layers be used for tregularization purposes ?
, b_build_all = False # Added by RMO - full Model is build in 2 steps
):
'''
Input:
The encoder_... and decoder_.... variables are Python lists,
whose length defines the number of Conv2D and Conv2DTranspose layers
input_dim : Shape/dimensions of the input tensor - for MNIST (28,28,1)
encoder_conv_filters: List with the number of maps/filters per Conv2D layer
encoder_conv_kernel_size: List with the kernel sizes for the Conv-Layers
encoder_conv_strides: List with the strides used for the Conv-Layers
act : determines activation function to use (0: LeakyRELU, 1:RELU , 2: SELU)
!!!! NOTE: !!!!
If SELU is used then the weight kernel initialization and the dropout layer need to be special
https://github.com/christianversloot/machine-learning-articles/blob/main/using-selu-with-tensorflow-and-keras.md
AlphaDropout instead of Dropout + LeCunNormal for kernel initializer
z_dim : dimension of the "latent_space"
solution_type : Type of solution for KL loss calculation (0: Customized Encoder layer,
1: model.add_loss()
2: definition of training step with Gradient.Tape()
use_batch_norm = False # True : We use BatchNormalization
use_dropout = False # True : We use dropout layers (rate = 0.25, see Encoder)
b_build_all = False # True : Full VAE Model is build in 1 step;
False: Encoder, Decoder, VAE are build in separate steps
'''
self.name = 'variational_autoencoder'
# Parameters for Layers which define the Encoder and Decoder
self.input_dim = input_dim
self.encoder_conv_filters = encoder_conv_filters
self.encoder_conv_kernel_size = encoder_conv_kernel_size
self.encoder_conv_strides = encoder_conv_strides
self.decoder_conv_t_filters = decoder_conv_t_filters
self.decoder_conv_t_kernel_size = decoder_conv_t_kernel_size
self.decoder_conv_t_strides = decoder_conv_t_strides
self.z_dim = z_dim
# Check param for activation function
if act < 0 or act > 2:
print("Range error: Parameter " + str(act) + " has unknown value ")
sys.exit()
else:
self.act = act
# Factor to scale the KL loss relative to the Binary Cross Entropy loss
self.fact = fact
# Check param for solution approach
if solution_type < 0 or solution_type > 2:
print("Range error: Parameter " + str(solution_type) + " has unknown value ")
sys.exit()
else:
self.solution_type = solution_type
self.use_batch_norm = use_batch_norm
self.use_dropout = use_dropout
# Preparation of some variables to be filled later
self._encoder_input = None # receives the Keras object for the Input Layer of the Encoder
self._encoder_output = None # receives the Keras object for the Output Layer of the Encoder
self.shape_before_flattening = None # info of the Encoder => is used by Decoder
self._decoder_input = None # receives the Keras object for the Input Layer of the Decoder
self._decoder_output = None # receives the Keras object for the Output Layer of the Decoder
# Layers / tensors for KL loss
self.mu = None # receives special Dense Layer's tensor for KL-loss
self.log_var = None # receives special Dense Layer's tensor for KL-loss
# Parameters for SELU - just in case we may need to use it somewhere
# https://keras.io/api/layers/activations/ see selu
self.selu_scale = 1.05070098
self.selu_alpha = 1.67326324
# The number of Conv2D and Conv2DTranspose layers for the Encoder / Decoder
self.n_layers_encoder = len(encoder_conv_filters)
self.n_layers_decoder = len(decoder_conv_t_filters)
self.num_epoch = 0 # Intialization of the number of epochs
# A matrix for the values of the losses
self.std_loss = tf.TensorArray(tf.float32, size=0, dynamic_size=True, clear_after_read=False)
# We only build the whole AE-model if requested
self.b_build_all = b_build_all
if b_build_all:
self._build_all()
Note that for the present post we (can) only use “solution_type = 0” !
A method to build the Encoder
The class shall provide a method to build the Encoder. For our present purposes including a customized layer based on the class “My_KL_Layer”. This layer just returns its input – namely the layers “mu” and “var_log” for the variational calculation of z-points, but it also calculates the KL loss which is added to other model losses.
# Method to build the Encoder
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~
def _build_enc(self, solution_type = 0, fact=-1.0):
'''
Encoder
@summary: Method to build the Encoder part of the AE
This will be a CNN defined by the parameters to __init__
@note: For self.solution = 0, we add an extra layer to calculate the KL loss
@note: The last layer uses a sigmoid activation to create the output
This may not be compatible with some scalers applied to the input data (images)
'''
# Check whether "fact" for the KL loss shall be overwritten
if fact < 0:
fact = self.fact
# Preparation: We later need a function to calculate the z-points in the latent space
# this function will be used by an eventual Lambda layer of the Encoder
def z_point_sampling(args):
'''
A point in the latent space is calculated statistically
around an optimized mu for each sample
'''
mu, log_var = args # Note: These are 1D tensors !
epsilon = B.random_normal(shape=B.shape(mu), mean=0., stddev=1.)
return mu + B.exp(log_var / 2) * epsilon
# Input "layer"
self._encoder_input = Input(shape=self.input_dim, name='encoder_input')
# Initialization of a running variable x for individual layers
x = self._encoder_input
# Build the CNN-part with Conv2D layers
# Note that stride>=2 reduces spatial resolution without the help of pooling layers
for i in range(self.n_layers_encoder):
conv_layer = Conv2D(
filters = self.encoder_conv_filters[i]
, kernel_size = self.encoder_conv_kernel_size[i]
, strides = self.encoder_conv_strides[i]
, padding = 'same' # Important ! Controls the shape of the layer tensors.
, name = 'encoder_conv_' + str(i)
)
x = conv_layer(x)
# The "normalization" should be done ahead of the "activation"
if self.use_batch_norm:
x = BatchNormalization()(x)
# Selection of activation function (out of 3)
if self.act == 0:
x = LeakyReLU()(x)
elif self.act == 1:
x = ReLU()(x)
elif self.act == 2:
# RMO: Just use the Activation layer to use SELU with predefined (!) parameters
x = Activation('selu')(x)
# Fulfill some SELU requirements
if self.use_dropout:
if self.act == 2:
x = AlphaDropout(rate = 0.25)(x)
else:
x = Dropout(rate = 0.25)(x)
# Last multi-dim tensor shape - is later needed by the decoder
self._shape_before_flattening = B.int_shape(x)[1:]
# Flattened layer before calculating VAE-output (z-points) via 2 special layers
x = Flatten()(x)
# "Variational" part - create 2 Dense layers for a statistical distribution of z-points
self.mu = Dense(self.z_dim, name='mu')(x)
self.log_var = Dense(self.z_dim, name='log_var')(x)
if solution_type == 0:
# Customized layer for the calculation of the KL loss based on mu, var_log data
# We use a customized layer accoding to a class definition
self.mu, self.log_var = My_KL_Layer()([self.mu, self.log_var], fact=fact)
# Layer to provide a z_point in the Latent Space for each sample of the batch
self._encoder_output = Lambda(z_point_sampling, name='encoder_output')([self.mu, self.log_var])
# The Encoder Model
self.encoder = Model(self._encoder_input, self._encoder_output)
A method to build the Decoder
The following function should be self-evident; it reverses the Encoder’s operations and uses z-points of the latent space as input.
# Method to build the Decoder
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~
def _build_dec(self):
'''
Decoder
@summary: Method to build the Decoder part of the AE
Normally this will be a reverse CNN defined by the parameters to __init__
'''
# Input layer - aligned to the shape of the output layer
self._decoder_input = Input(shape=(self.z_dim,), name='decoder_input')
# Here we use the tensor shape info from the Encoder
x = Dense(np.prod(self._shape_before_flattening))(self._decoder_input)
x = Reshape(self._shape_before_flattening)(x)
# The inverse CNN
for i in range(self.n_layers_decoder):
conv_t_layer = Conv2DTranspose(
filters = self.decoder_conv_t_filters[i]
, kernel_size = self.decoder_conv_t_kernel_size[i]
, strides = self.decoder_conv_t_strides[i]
, padding = 'same' # Important ! Controls the shape of tensors during reconstruction
# we want an image with the same resolution as the original input
, name = 'decoder_conv_t_' + str(i)
)
x = conv_t_layer(x)
# Normalization and Activation
if i < self.n_layers_decoder - 1:
# Also in the decoder: normalization before activation
if self.use_batch_norm:
x = BatchNormalization()(x)
# Choice of activation function
if self.act == 0:
x = LeakyReLU()(x)
elif self.act == 1:
x = ReLU()(x)
elif self.act == 2:
#x = self.selu_scale * ELU(alpha=self.selu_alpha)(x)
x = Activation('selu')(x)
# Adaptions to SELU requirements
if self.use_dropout:
if self.act == 2:
x = AlphaDropout(rate = 0.25)(x)
else:
x = Dropout(rate = 0.25)(x)
# Last layer => Sigmoid output
# => This requires scaled input => Division of pixel values by 255
else:
x = Activation('sigmoid')(x)
# Output tensor => a scaled image
self._decoder_output = x
# The Decoder model
self.decoder = Model(self._decoder_input, self._decoder_output)
Note that we do not include any loss calculations in the Decoder model. The main loss – namely according to the “binary cross entropy” will later be added to the “fit()” method of the full Keras based VAE model.
The full VAE model
We have already created two Keras models for the Encoder and Decoder. We now combine them to the full VAE model and save this model in a variable of the object derived from our class.
# Function to build the full AE
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
def _build_VAE(self):
model_input = self._encoder_input
model_output = self.decoder(self._encoder_output)
self.model = Model(model_input, model_output, name="vae")
# Function to build full AE in one step if requested
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
def _build_all(self):
self._build_enc()
self._build_dec()
self._build_VAE()
Compilation
For our present solution with the customized layer for the KL loss we now provide a matching “compile()” function:
# Function to compile VA-model with a KL-layer in the Encoder
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
def compile_for_KL_Layer(self, learning_rate):
if self.solution_type != 0:
print("The compile_L() function is only compatible with solution_type = 0")
sys.exit()
self.learning_rate = learning_rate
# Optimizer
optimizer = Adam(learning_rate=learning_rate)
self.model.compile(optimizer=optimizer, loss="binary_crossentropy",
metrics=[tf.keras.metrics.BinaryCrossentropy(name='bce')])
This is the place where we include the main contribution to the loss – namely by a “binary cross-entropy” calculation with respect to the differences between the original input tensor top our model and its output tensor. We had to use the function BinaryCrossentropy(name=’bce’) to be able to give the respective output during training a short name. All in all we expect an output during training comprising:
- the total loss
- the contribution from the binary_crossentropy
- the KL contribution
A method for training
We are almost finished. We just need a matching method for starting the training via calling the “fit()“-function of our Keras based VAE model:
def train_model_with_KL_Layer(self, x_train, batch_size, epochs, initial_epoch = 0):
self.model.fit(
x_train
, x_train
, batch_size = batch_size
, shuffle = True
, epochs = epochs
, initial_epoch = initial_epoch
)
Note that we called the same “x_train” batch of samples twice: The standard “y” output “labels” actually are the input samples (which is, of course, the core characteristic of AEs). We shuffle data during training.
Why use a special function of the class at all and not directly call fit() from Jupyter notebook cells?
Well, at this point we could include multiple other things as custom callbacks (e.g. for special output or model saving) and a scheduler. See e.g. the code of D. Foster at his Github site for variants. For the sake of briefness I skip these techniques in my post.
Jupyter cells to use our class
Let us see how we can use our carefully crafted class with a Jupyter notebook. As I personally gather Python modules (via Eclipse PyDev) in some special folders, I first have to add a path:
Cell 1:
import sys
# !!! ADAPT to YOUR needs !!!!!
sys.path.append("/projects/GIT/ml_4/")
print(sys.path)
Of course, you must adapt this path to your personal situation.
The next cell contains module imports
Cell 2
import numpy as np import time import os import sklearn # could be used for scalers import matplotlib as mpl from matplotlib import pyplot as plt from matplotlib.colors import ListedColormap import matplotlib.patches as mpat # tensorflow and keras import tensorflow as tf from tensorflow import keras as K from tensorflow.python.keras import backend as B from tensorflow.keras import models from tensorflow.keras import layers from tensorflow.keras import regularizers from tensorflow.keras import optimizers from tensorflow.keras import metrics from tensorflow.keras.datasets import mnist from tensorflow.keras.optimizers import schedules from tensorflow.keras.utils import to_categorical from tensorflow.python.client import device_lib from tensorflow.keras.datasets import mnist # My VAE-class from my_AE_code.models.My_VAE import MyVariationalAutoencoder
I then suppress some warnings regarding my Nvidia card and list the available Cuda devices.
Cell 3
# Suppress some TF2 warnings on negative NUMA node number
# see https://www.programmerall.com/article/89182120793/
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3' # or any {'0', '1', '2'}
tf.config.experimental.list_physical_devices()
We then control resource usage:
Cell 4
# Restrict to GPU and activate jit to accelerate
# IMPORTANT NOTE: To change any of the following values you MUT restart the notebook kernel !
b_tf_CPU_only = False # we want to work on a GPU
tf_limit_CPU_cores = 4
tf_limit_GPU_RAM = 2048
if b_tf_CPU_only:
tf.config.set_visible_devices([], 'GPU') # No GPU, only CPU
# Restrict number of CPU cores
tf.config.threading.set_intra_op_parallelism_threads(tf_limit_CPU_cores)
tf.config.threading.set_inter_op_parallelism_threads(tf_limit_CPU_cores)
else:
gpus = tf.config.experimental.list_physical_devices('GPU')
tf.config.experimental.set_virtual_device_configuration(gpus[0],
[tf.config.experimental.VirtualDeviceConfiguration(memory_limit = tf_limit_GPU_RAM)])
# JiT optimizer
tf.config.optimizer.set_jit(True)
Let us load MNIST for test purposes:
Cell 5
def load_mnist():
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train = x_train.astype('float32') / 255.
x_train = x_train.reshape(x_train.shape + (1,))
x_test = x_test.astype('float32') / 255.
x_test = x_test.reshape(x_test.shape + (1,))
return (x_train, y_train), (x_test, y_test)
(x_train, y_train), (x_test, y_test) = load_mnist()
Provide the VAE setup variables to our class:
Cell 6
z_dim = 2
vae = MyVariationalAutoencoder(
input_dim = (28,28,1)
, encoder_conv_filters = [32,64,128]
, encoder_conv_kernel_size = [3,3,3]
, encoder_conv_strides = [1,2,2]
, decoder_conv_t_filters = [64,32,1]
, decoder_conv_t_kernel_size = [3,3,3]
, decoder_conv_t_strides = [2,2,1]
, z_dim = z_dim
, act = 0
, fact = 5.e-4
)
Set up the Encoder:
Cell 7
# overwrite the KL fact from the class fact = 2.e-4 vae._build_enc(fact=fact) vae.encoder.summary()
Build the Decoder:
Cell 8
vae._build_dec() vae.decoder.summary()
Build the VAE model:
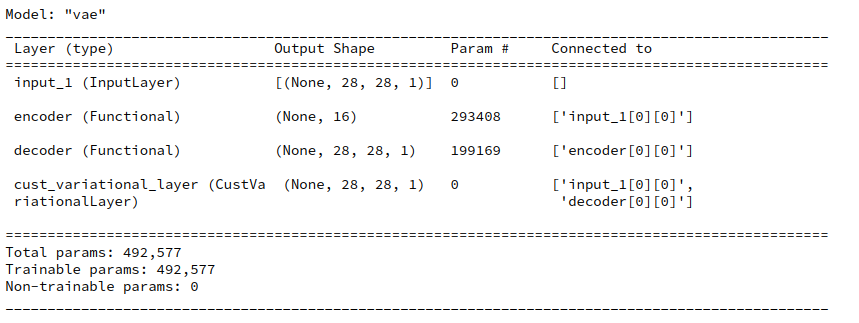
Cell 9
vae._build_VAE() vae.model.summary()
Compile
Cell 10
LEARNING_RATE = 0.0005 vae.compile_for_KL_Layer(LEARNING_RATE)
Train / fit the model to the training data
Cell 11
BATCH_SIZE = 128
EPOCHS = 6 # for real runs ca. 40
INITIAL_EPOCH = 0
vae.train_model_with_KL_Layer(
x_train[0:60000]
, batch_size = BATCH_SIZE
, epochs = EPOCHS
, initial_epoch = INITIAL_EPOCH
)
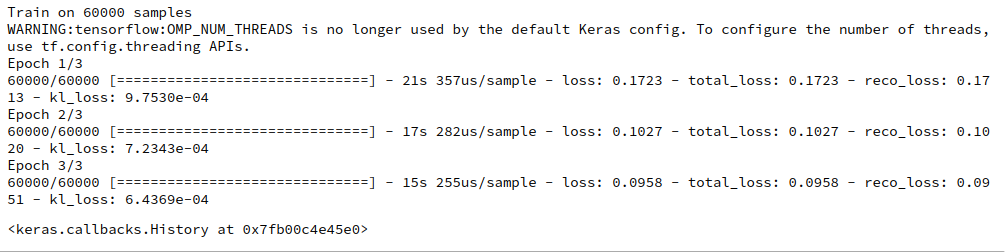
For the given parameters I got the following output on my old GTX960
Epoch 1/6 469/469 [==============================] - 12s 24ms/step - loss: 0.2613 - bce: 0.2589 - kl: 0.0024 Epoch 2/6 469/469 [==============================] - 12s 25ms/step - loss: 0.2174 - bce: 0.2159 - kl: 0.0015 Epoch 3/6 469/469 [==============================] - 11s 23ms/step - loss: 0.2100 - bce: 0.2085 - kl: 0.0015 Epoch 4/6 469/469 [==============================] - 11s 23ms/step - loss: 0.2057 - bce: 0.2042 - kl: 0.0015 Epoch 5/6 469/469 [==============================] - 11s 23ms/step - loss: 0.2034 - bce: 0.2019 - kl: 0.0015 Epoch 6/6 469/469 [==============================] - 11s 23ms/step - loss: 0.2019 - bce: 0.2004 - kl: 0.0015
So 11 secs for an epoch of 60,000 samples with batch-size = 128 is a reference point. Note that this is obviously faster than what we got for the solution discussed in the last post.
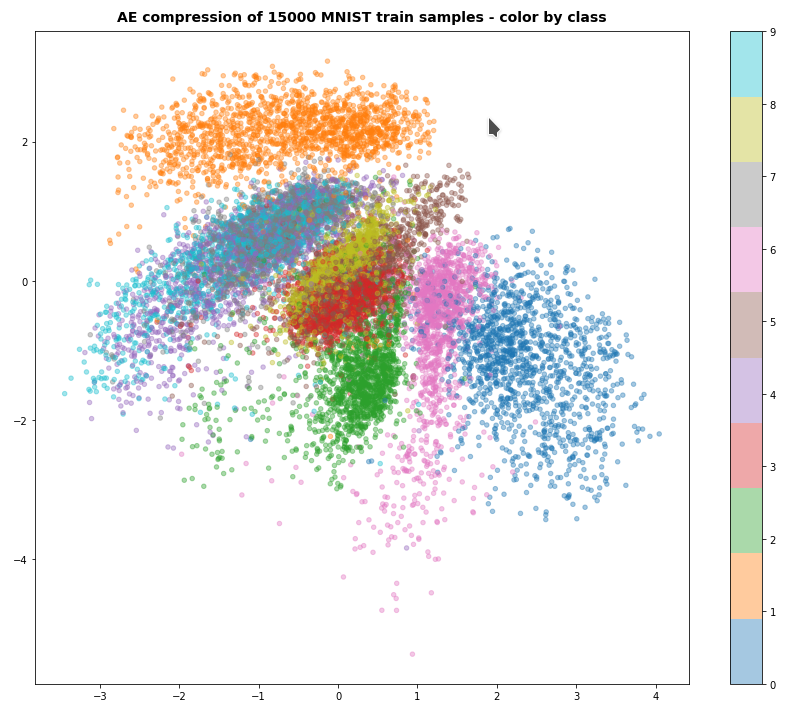
Just to give you an impression of other results:
For z_dim = 2, fact = 2.e-4 and 60 epochs I got something like the following data point distribution in the latent space:
I shall discuss more results – also for other test data sets – in future posts in this blog.
Conclusion
In this post we have build a class to set up a VAE based on an Encoder and a Decoder model with Conv2D and Conv2dTranspose layers. We delegated the calculation of the KL loss to a customized layer of the Encoder, whilst the main loss contribution was defined in form of a binary-crossentropy evaluation with the help of the fit()-function of the VAE model. All loss contributions were displayed as “metrics” elements during training. The presented solution is fully compatible with Tensorflow 2.8 and eager execution. It is in my opinion also elegant and very Keras oriented as all important operations are encapsulated in a continuous sequence of layers. We also found this to be a relatively fast solution.
In the next post of this series
Variational Autoencoder with Tensorflow – VI – KL loss via tensor transfer and multiple output
we are going to use our class to adapt an older suggestion of D.Foster to the requirements of TF2.8.
References
F. Chollet, Deep Learning mit Python und Keras, 2018, 1-te dt. Auflage, mitp Verlags GmbH & Co.KG, Frechen
D. Foster, “Generatives Deep Learning”, 2020, 1-te dt. Auflage, dpunkt Verlag, Heidelberg in Kooperation mit Media Inc.O’Reilly, ISBN 978-3-960009-128-8. See Kap. 3 and the VAE code published at
https://github.com/davidADSP/GDL_code/
Louis Tiao, “Implementing Variational Autoencoders in Keras: Beyond the Quickstart Tutorial”, 2017, http://louistiao.me/posts/implementing-variational-autoencoders-in-keras-beyond-the-quickstart-tutorial/
Recommendation: The article of L. Tiao is not only interesting regarding Keras modularity. I like it very much also for his mathematical depth. I highly recommend his article as a source of inspiration, especially with respect to alternative divergences. Please, also follow Tiao’s list of well selected literature references.
And before I forget it:
Ceterum censeo: The worst living fascist and war criminal today, who must be isolated, denazified and imprisoned, is the Putler.