This post requires Javascript to display formulas!
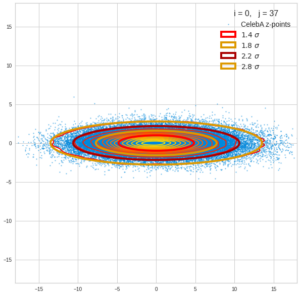
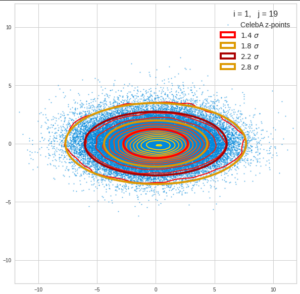
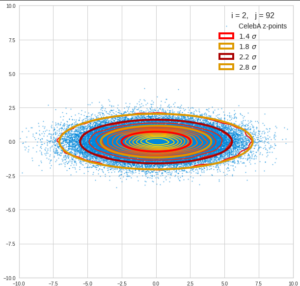
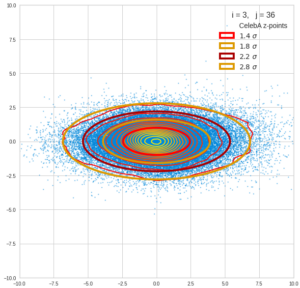
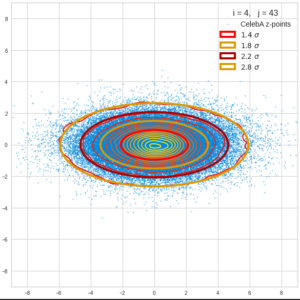
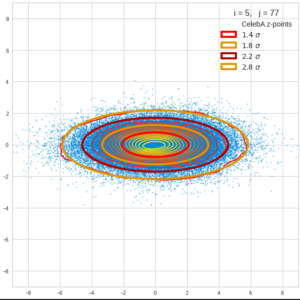
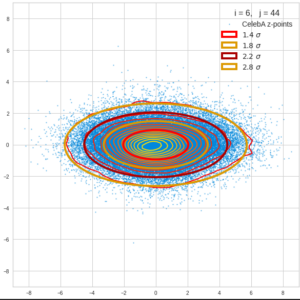
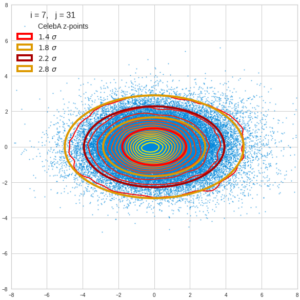
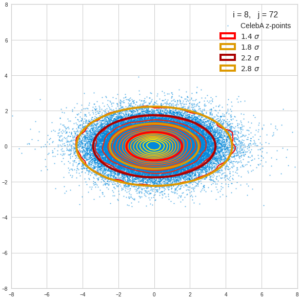
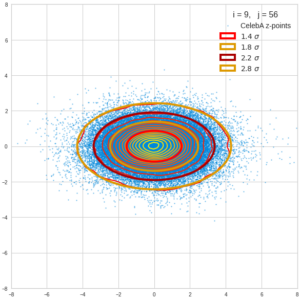
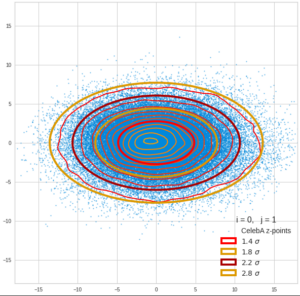
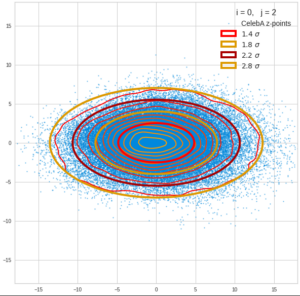
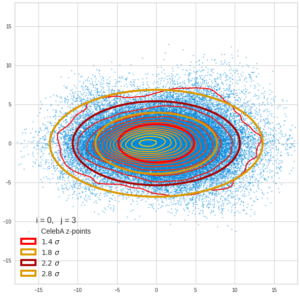
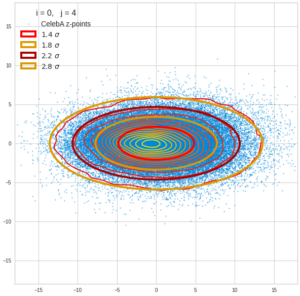
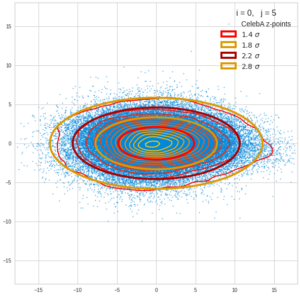
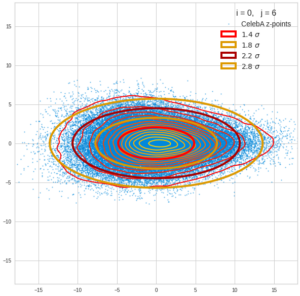
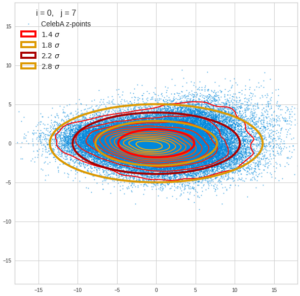
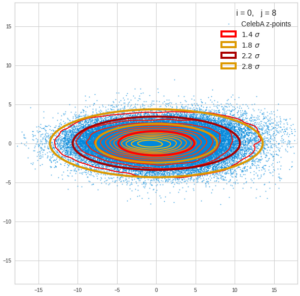
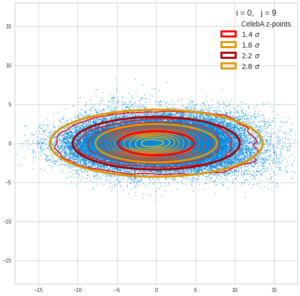
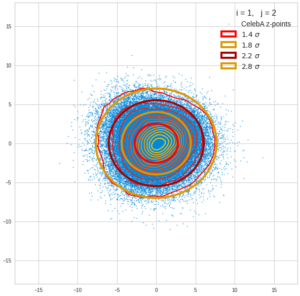
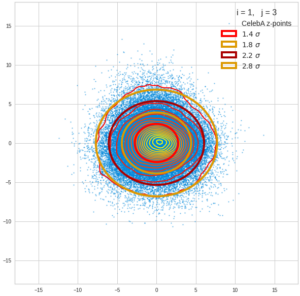
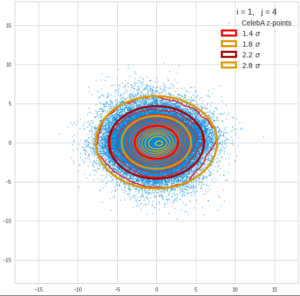
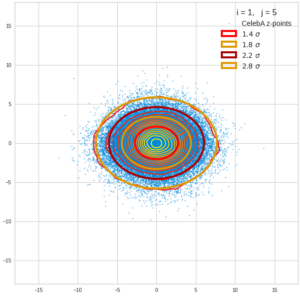
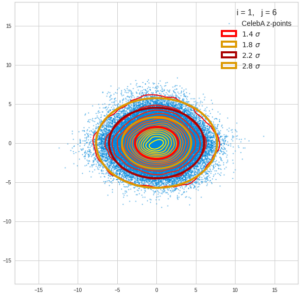
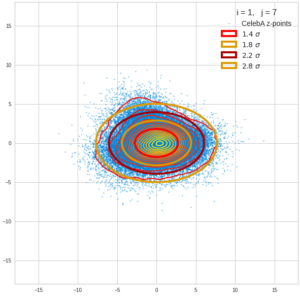
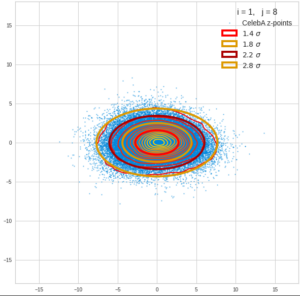
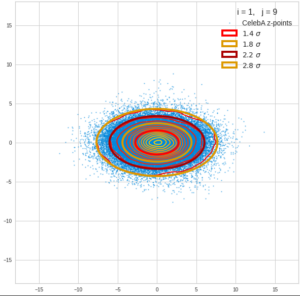
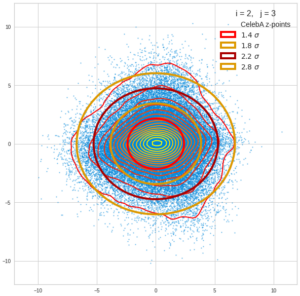
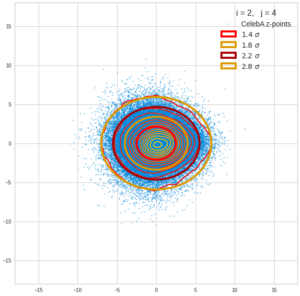
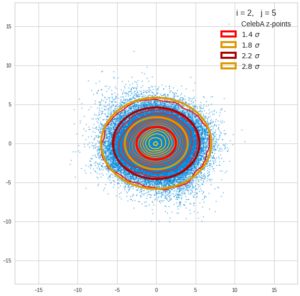
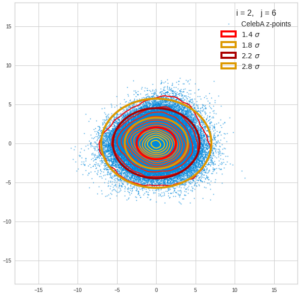
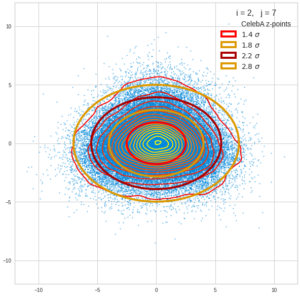
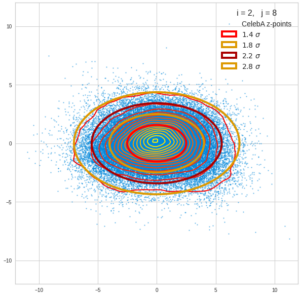
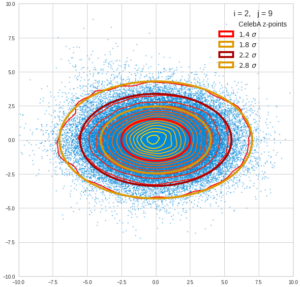
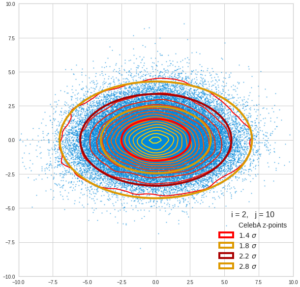
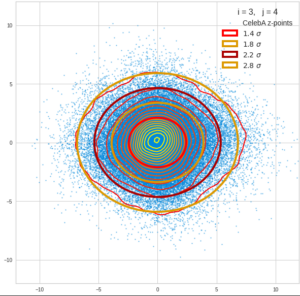
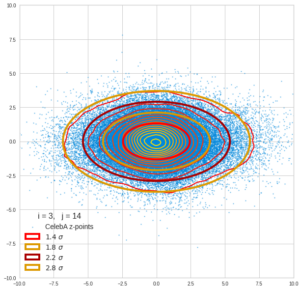
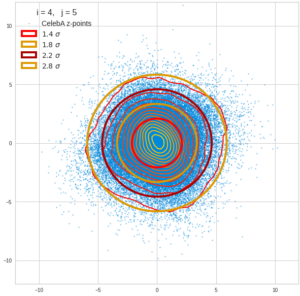
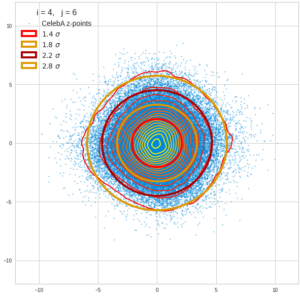
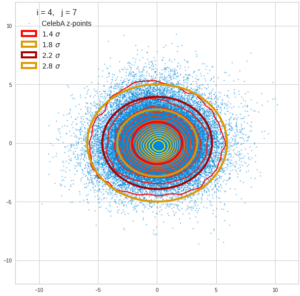
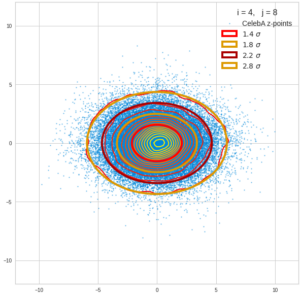
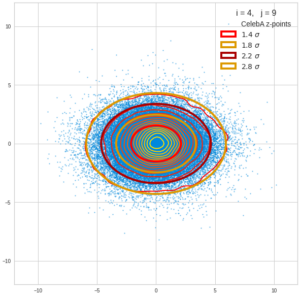
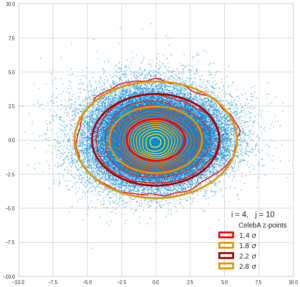
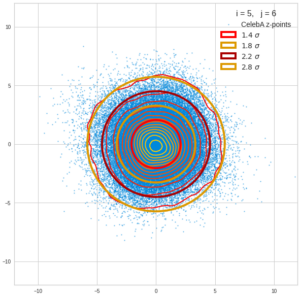
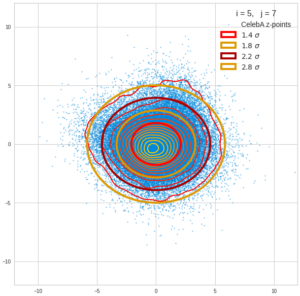
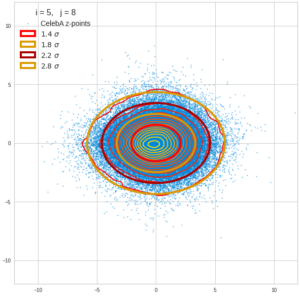
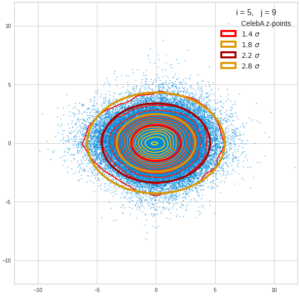
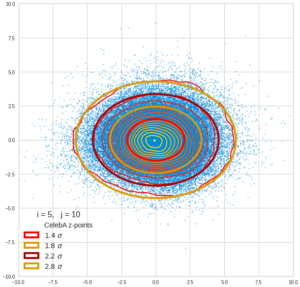
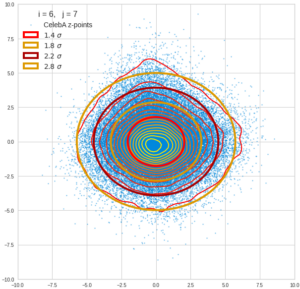
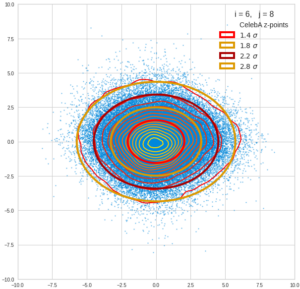
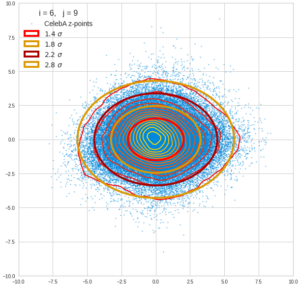
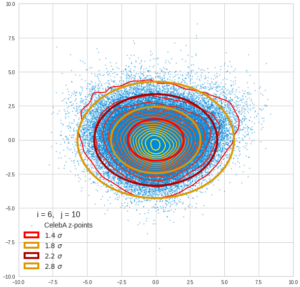
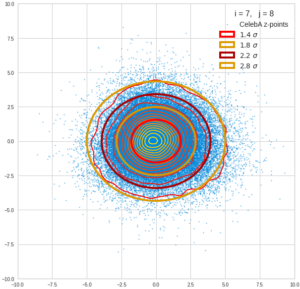
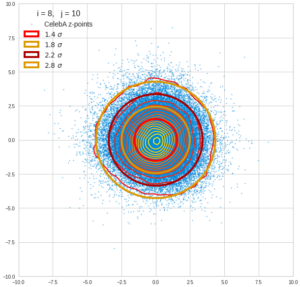
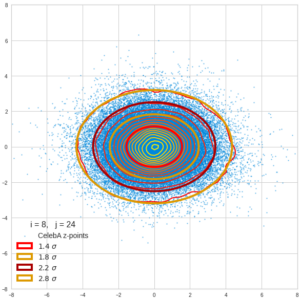
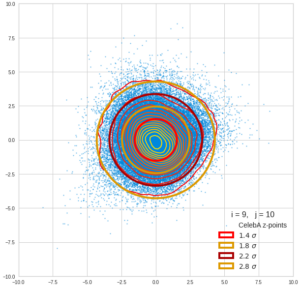
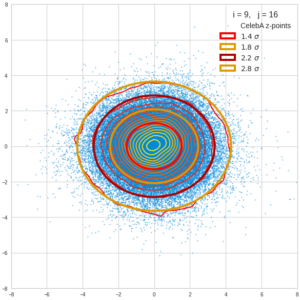
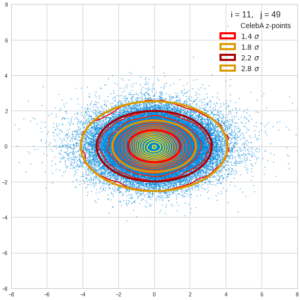
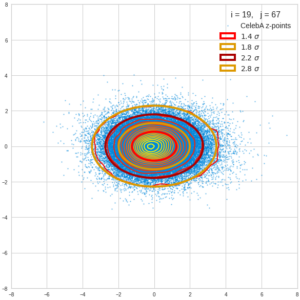
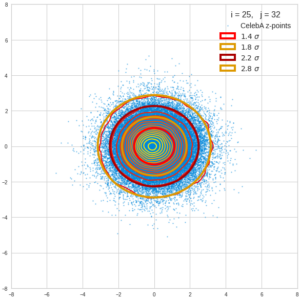
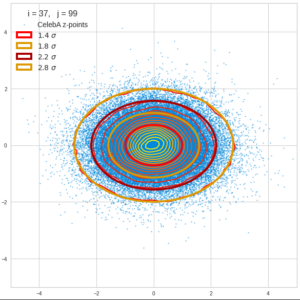
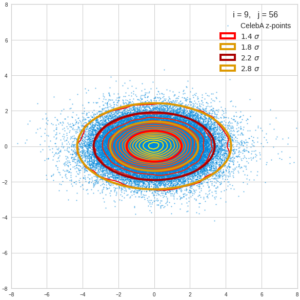
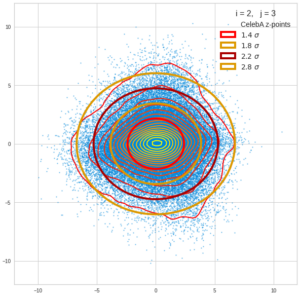
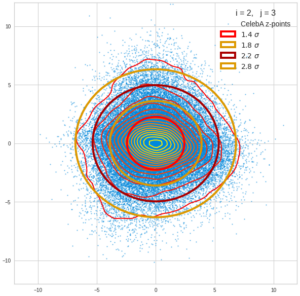
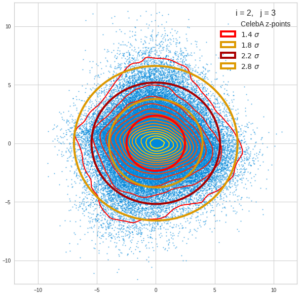
This post series is based on notes I took when I recently studied the mathematical properties of Multivariate Normal Distributions [MNDs]. A motivation to publish the notes came from the analysis of a Machine Learning experiment: A Convolutional Autoencoder [CAE] trained on human face images had produced a MND in its latent space. But MNDs are, of course, relevant in other contexts, too.
In this post we take some preparatory steps: I want to introduce the term “random vector” to describe statistical multivariate distributions in vector form. We then have a look at the definition of a probability density function [pdf] in multiple dimensions. Furthermore we discuss linear transformations of a random vector, its expectation values and its covariance matrix. The latter both marks and quantifies (linear) relations between its components. I will also point out a simple but useful equation for the behavior of the covariance matrix when a matrix operates on its random vector. Eventually we also have a look at marginal distributions and their probability density function.
I will proceed not too formally and my discussion of the topics will not be really complete in a pure mathematical sense. On our way I will also remind you of some basic ideas and equations from statistics. People who are familiar with all the terms named above can jump over this post.
Random Vectors and multivariate distributions
Let us assume that we can describe an object of interest, e.g. in the context of an ML problem, by a set of n indexed, real-value variables sj. Instead of a manifest object we could also use n variables to characterize the result of a well defined experiment. When I speak of “objects below I include this more abstract option.
Each object can mathematically be represented by a data point in the ℝn. We simply relate each of the indexed coordinate axes of a corresponding Euclidean coordinate system with a certain object property. The coordinates of the data points are then given by the variable values.
Let us now order a complete set of parallel measured property values sjc for a concrete object as indexed components of a vector sc. The vector then represents our object:
\]
The T indicates the transposition operation. Note: Throughout this post series we assume that untransposed vectors are written in vertical form. I.e. the components are arranged as rows.
Samples
Let us assume that we have m concrete observations of individual objects (or experimental results). The values of the n real variables for each object are always measured or qualified in parallel:
- We either randomly pick a bunch of m concrete objects out of a wider population of available objects and determine the values of their n characteristic sj-variables.
- Or we repeat a concrete experiment, whose outcome is described by values for a set of n measurable sj-variables, many times. The setup for our experiment defines precisely what the variables are and how we measure their values. The experimental result can be interpreted as an abstract “object”.
Thus we get samples of object data. Our experimental setup and the properties of our objects define a sample space Ω we have a clear description of how we produce one or multiple vectors s. A particular observation maps Ω to an individual vector.
Let us assume that at least in principle our vector components can take any real value. Producing a vector s can then formally be regarded as a function to the ℝn:
FS: Ω => ℝn, s = FS(Ω, experiment number, … ).
A sequence of many observations, i.e. creating a sample, formally corresponds to a sequence of many independent applications of this function. An implicit (and sometime problematic) assumption is that picking multiple objects or our experiments do not influence Ω and the statistics of the basic population.
The outcome, namely a set of resulting s-vectors, depends on the statistical properties of the underlying object population. We index the k-th vector by sk.
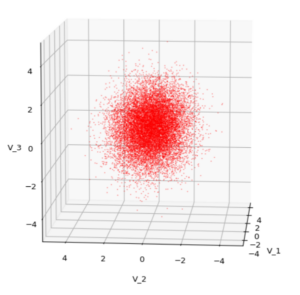
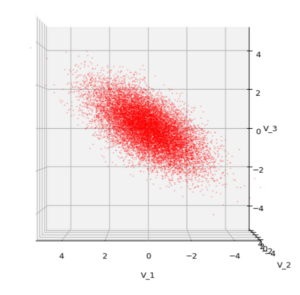
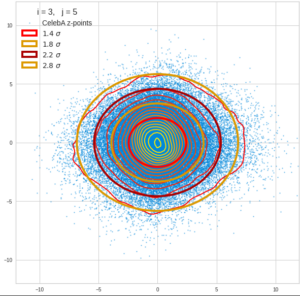
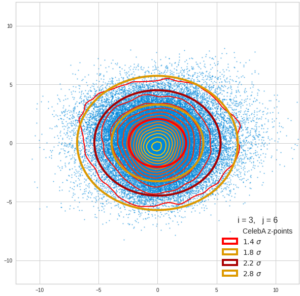
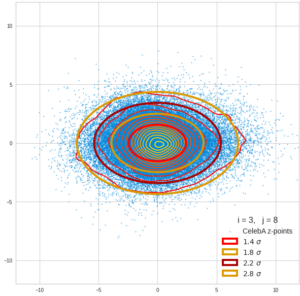
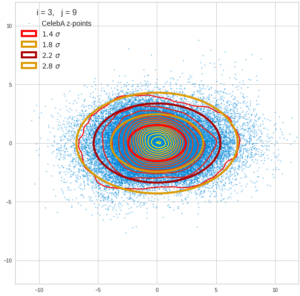
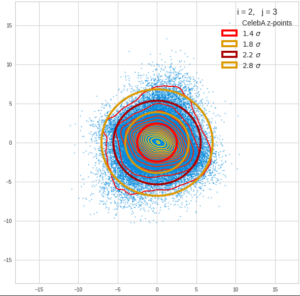
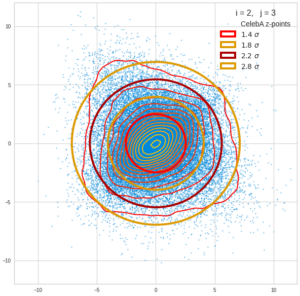
Each sample can be visualized by a bunch of data points in an n-dimensional space. A concrete sample in 3 dimensions may e.g. look like this from two perspectives:
But our sample can also be represented by a data matrix. We can e.g. arrange the columns to be given by the sk vectors. If we have a sample of m objects then we get a (n x m)- data matrix MS ∈ ℝnxm. A row with index j then provides us with a sample statistics for the j-th object property.
\pmb{\operatorname{D}}_S \: = \: \begin{pmatrix}
s^1_1 & s^2_1 &\cdots & s^m_1 \\
s^1_2 & s^2_2 &\cdots & s^m_2 \\
\vdots &\vdots &\ddots &\vdots \\
s^1_n & s^2_n &\cdots & s^m_n \end{pmatrix}
\]
\]
Often you will find the transposed matrix for reasons of an efficient notation. I prefer this version as it later allows for matrix operations on the vector components from the left side, e.g. in a Python program.
Random vectors
We indicate the statistical element in creating a concrete vector from an underpinning population by a vector symbol S. By using this symbol we understand implicitly that any concrete vector S = sc is created by applying our well defined function FS.
It is natural to split S up into n individual univariate random variables Sj: Ω => ℝ, which represent the statistics per vector component. We write S in vector form:
\]
So:
\]
The vector variable S gets concrete component values by a well defined statistical process. But, if you like to see it more globally, S represents any object from the population and thus in a certain meaning the whole population – and its statistics.
We call S a multivariate random variable or a random vector.
Thus, our statistical multivariate distribution can now be represented by a random vector – and we know how to create samples by turning S repeatedly into concrete vectors. Why all the fuss?
Linear operations on random vectors
The vector notation allows us to combine a random vector with mathematical vector operations, in particular linear transformations. We implicitly understand that such an operation is applied to any vector of the population (and in certain contexts also to elements of our samples). Such a transformation consists of a mapping of vectors of the original population or a sample to new vectors in the same or a different vector space by one and the same linear transformation. The mapping gives us new component values.
In linear algebra we learn that a linear vector transformation can be represented by a matrix plus a translation vector. If the transformation maps a random vector S of the ℝm to a random vector X into the ℝn then the linear (affine) transformation is defined by a (n x m)-matrix A ∈ ℝnxm and a vector b ∈ ℝn:
\pmb{x} \,&=\, (x_1, x_2, …, x_n)^T \,=\, \pmb{\operatorname{A}} \pmb{s} \, + \, \pmb{b} \\
&\, \\
\pmb{X} \, &= \, \pmb{\operatorname{A}S} + \pmb{b} \end{align}
\]
In close similarity to standard operations in vector spaces a linear transformation of a random vector can have two meanings:
- The operation either actively transforms our random vector (and the underlying vector distribution) – e.g. by a rotation in the ℝn.
- Or we perform a passive switch to another Euclidean coordinate system (e.g. by a rotation of the coordinate axes) and the transformation adapts the component values accordingly.
Invertible transformations?
If and when we have a transformation within the ℝn the matrix A would typically be a real valued quadratic (n x n)-matrix. In many cases we will assume that the transformation and its matrix A are invertible. This imposes certain conditions on the matrix: A must then be positive definite and have a determinant det(A) ≠ 0.
\, \ne \, 0
\]
But note that in the more general case also rectangular matrices can have a “left”- or a “right”-inverse matrices; see https://en.wikipedia.org/ wiki/ Generalized_ inverse.
Rectangular matrices, e.g. with m (< n) rows and n columns have a rank r ≤ m < n, and not all of the column vectors of the matrix are linearly independent.
Probability density functions in the ℝn
Let us assume that the population behind a random vector covers the ℝn completely and in a continuous way. By “continuous” we mean that a transition to infinitesimal small volumes at any position in the ℝn is possible: We still will find vectors of the population there and component values change smoothly between adjacent elements.
We can apply a frequency analysis to the vector component values of the objects of a huge sample or of a number of samples. E.g. we can define intervals into which the components may fall or not. The analysis gives us a probability P(S=sΔV) for the occurrence of position vectors sΔV which point into a certain finite region ΔV of the ℝn.
If you better like to think in terms of data points:
We get a probability that a data point representing a statistically picked object lies within a certain defined and finite region of the ℝn.
How we split up the ℝn into separate volume regions depends on available information about the object population or on an analysis of sufficiently large samples.
If each component of the vectors s can take any real value we may under certain circumstances make a transition to probability densities. Probability densities can be distilled from huge samples or alternatively very many samples picked from the underlying population. To get an idea of the density function we, in practice, raise the number of observations and reduce the volume regions of the ℝn to a proper averaging size that avoids wild fluctuations. The latter is called “sampling”.
If we by many experiments can assume a converging limiting process to infinitesimal real value intervals dsj for each of the vector components then we end up with a bunch of n continuous 1-dim probability density functions pj(sj): ℝ => ℝ. A concrete probability value for a component must be derived via integration of pj() over a finite interval a ≤ s_j ≤ b:
\]
An interpretation as a probability requires, of course, a normalization of such an integral over the whole ℝ.
It is clear that the full probability density function pS(s) of the vector distribution varies continuously with the 1-dim probability density functions of its components.
In this post series we are interested in examples of vector distributions which can be described by a continuous multidimensional probability density function [p.d.f] for infinitesimal volume elements dVS = ds1, ds2…dsn:
\]
&= \, \int\int…\int_V p_{\pmb{S}} \left( \, s_1, \, s_2, \, ….\, s_n\, \right) \, ds_1ds_2 \cdots ds_n
\end{align}
\]
where V marks some finite volume element in ℝn. Note again: A normalization of such an integral over the whole ℝn is required to get well defined probability values. The normalization factor above is assumed to be integrated in the mathematical expression for pS(s).
For the purposes of this post series we assume that both pS and the pj() are differentiable.
Correlations of random vector components
There may be dependencies which control how the components of a random vector vary together. So the variation of pS(s) may depend in a rather complicated way on the density functions of their components pj(sj)
\]
The variation of components may e.g. happen in pairwise correlated or uncorrelated ways. I.e. we can in general not assume that the total probability density function pS(s) for a random vector is e.g. a product of the 1-dimensional density probability functions pj(sj for its components.
Such dependencies make the statistics of multivariate vector distributions more complex than the statistics of their constituting (marginal) univariate components. The relation between pS and the individual pj can be difficult to derive and it may even reflect non-linear component relations. Luckily we will see that MNDs contain only linear couplings of the components and thus simple correlations described by constant coefficients.
In ML we have only samples with a limited number of individual vectors available. The investigated samples have to be big enough to reveal the properties of the total probability density function pS(s), which assumedly describes the variation of the underlying continuous vector population in the ℝn.
Probability density after linear transformations and contour hyper-surfaces
A transformation of the components of a random vector has, of course, also an impact on the resulting probability density:
\]
and for a finite volume VX in the target space
\]
In case of a well defined distribution S we can hope to find an analytical expression for the multidimensional probability pX(x) of the transformed random vector X. But as we will see we have to invest some work into the calculation. And we will answer the question by what parameters the probability density of a MND is defined in a unique way.
We expect that the concrete form of pX is influenced in a characteristic way by the properties of the original pj(sj)-pdfs and, of course, by our transformation matrix A. This is the case for MNDs and we will study the effects of linear transformations on MNDs and their projections to sub-spaces in more detail in forthcoming posts.
As the matrix A mixes components we may also expect the creation of (linear) relations between the X-components. Therefore, the shape of the density distribution may change due to scaling, mirroring, shearing and rotation effects. This results from the fact that linear transformations are affine transformations in geometrical terms.
This leads to the question by what method we can describe the “shape” of a multivariate distribution. A simple method is to determine hyper-surfaces of constant probability density values. I.e. contour-surfaces:
\]
This actually is the definition of a (n-1)-dimensional hypersurface. The dependency on the component values can be complicated. But for the special case of a MND we will find a rather comprehensible form which we can interpret. So we will get a chance to understand and qualify the effects of a linear transformation by its impact on the contour hyper-surfaces of the probability density of a MND.
Expectation value and covariance of a statistical vector distribution
The covariance of two univariate distributions X and Y is defined as
\begin{align}
\operatorname{cov}(X, Y)
&= \operatorname{\mathbb{E}}\left[\,\left(\,X \,-\, \operatorname{\mathbb{E}}\left(X\right)\,\right) \, \left(\,Y \,-\, \operatorname{\mathbb{E}}\left(Y\right)\, \right)\,\right] \\
&= \operatorname{\mathbb{E}}\left(X Y\right) \,-\, \operatorname{\mathbb{E}}\left(X\right) \operatorname{\mathbb{E}}\left(Y\right)
\end{align}
\]
The expectation value and the covariance of a general vector distribution S are defined by:
\operatorname{Cov}\left(\pmb{S}\right) \: &:= \: \operatorname{\mathbb{E}}\left[\left(\pmb{S} – \operatorname{\mathbb{E}}(\pmb{S}) \right)\, \left(\pmb{S} – \operatorname{\mathbb{E}}(\pmb{S}) \right)^T \right] \end{align}
\]
Note the order of transposition in the definition of Cov! A (vertical) vector is combined with a transposed (horizontal) vector. The rules of a matrix-multiplication then give you a matrix as the result! And the expectation value has to be taken for every element of the matrix.
Thus, the interpretation of the notation given above is:
Pick all pairwise combinations (Sj, Sk) of the component distributions. Calculate the covariance of the pair cov(S_j, S_k) and put it at the (j,k)-place of the matrix.
Meaning:
\operatorname{Cov}\left(\pmb{S}\right)\:&=\:\operatorname{\mathbb{E}} {\begin{pmatrix} (S_{1}-\mu_{1})^{2}&(S_{1}-\mu_{1})(S_{2}-\mu_{2})&\cdots &(S_{1}-\mu_{1})(S_{n}-\mu_{n})\\(S_{2}-\mu_{2})(S_{1}-\mu_{1})&(S_{2}-\mu_{2})^{2}&\cdots &(S_{2}-\mu _{2})(S_{n}-\mu_{n})\\\vdots &\vdots &\ddots &\vdots \\(S_{n}-\mu_{n})(S_{1}-\mu_{1})&(S_{n}-\mu_{n})(S_{2}-\mu _{2})&\cdots &(X_{n}-\mu_{n})^{2} \end{pmatrix}}
\\\\
&=\quad {\begin{pmatrix} \operatorname{Var} (S_{1})&\operatorname{Cov} (S_{1},S_{2})&\cdots &\operatorname{Cov} (S_{1},S_{n})\\ \operatorname{Cov} (S_{2},S_{1})&\operatorname{Var} (S_{2})&\cdots &\operatorname{Cov} (S_{2},S_{n})\\\vdots &\vdots &\ddots &\vdots \\ \operatorname{Cov} (S_{n},S_{1})&\operatorname{Cov} (S_{n},S_{2})&\cdots &\operatorname {Var} (S_{n}) \end{pmatrix}}
\end{align}
\]
Keep the details of this definition in mind!
The above matrix is the (variance-) covariance matrix. As Wikipedia tells you, some people call it a bit different. I refer to it later on just as the covariance matrix (of a random vector).
Useful properties of the covariance matrix of a random vector
Note that the diagonal elements of the covariance matrix are just the variances of the individual component distributions, whilst the off diagonal elements contain the pairwise covariances of the vector components. In case of a vector distribution with uncorrelated (or even independent) Sj the matrix becomes diagonal:
\begin{pmatrix}
(\sigma_1^s)^2 & 0 &\cdots & 0 \\
0 & (\sigma_2^s)^2 &\cdots & 0 \\
\vdots &\vdots &\ddots &\vdots \\
0 & 0 &\cdots & (\sigma_n^s)^2 \end{pmatrix}
\]
Other properties can be derived from the linearity of expectation values and the fact that
\]
It is relatively easy to show that the defined expectation value of a vector distribution behaves linearly with respect to linear transformations of S with a matrix A:
\operatorname{\mathbb{E}}\left( \pmb{\operatorname{A}S} + \pmb{b} \right) \, = \pmb{\operatorname{A}} \operatorname{\mathbb{E}}\left(\pmb{S} \right) \,+\, \pmb{b} \vphantom{(}
\]
And the covariance matrix Cov transforms as follows:
\operatorname{Cov}\left(\pmb{\operatorname{A}S}\right) \: &= \: \operatorname{\mathbb{E}}\left[\, \left( \pmb{\operatorname{A}} (\pmb{S} \,-\, \mu_S ) \right) \, \left( \pmb{\operatorname{A}} (\pmb{S} \,-\, \mu_S ) \right)^T \, \right] \\ &= \: \operatorname{\mathbb{E}}\left[\, \pmb{\operatorname{A}} ( \pmb{S} \,-\, \mu_S ) \, (\pmb{S} \,-\, \mu_S)^T \pmb{\operatorname{A}}^T \,\right] \\
&= \: \pmb{\operatorname{A}} \operatorname{\mathbb{E}}\left[\, ( \pmb{S} \,-\, \mu_S ) \, (\pmb{S} \,-\, \mu_S)^T \,\right] \pmb{\operatorname{A}}^T \\
\operatorname{Cov}\left(\pmb{\operatorname{A}S}\right) \: &= \: \pmb{\operatorname{A}} \operatorname{Cov}\left(\pmb{S}\right) \pmb{\operatorname{A}}^T \end{align}
\]
The intermediate steps for the Cov – and in particular the 3rd one – are in my opinion best understood when you just write the stuff down for e.g. (3×3)-matrices and carefully regard the Cov-definition for random vectors. You will see that the definition of Cov just allows for the right index juggling! The extension to more dimensions is straightforward and could formally be done by induction.
See e.g. also https://cds.nyu.edu/ wp-content/ uploads/ 2021/05/ covariance_matrix.pdf and https://stats.stackexchange.com/ questions/ 113700/ covariance-of-a-random-vector-after-a-linear-transformation.
Other useful properties are
\begin{align}
\operatorname{Cov}\left(V_j,\, V_k\right) \,&=\, \operatorname{\mathbb{E}}\left(V_j, \,V_k\right) \,-\, \operatorname{\mathbb{E}}\left(V_j\right)\operatorname{\mathbb{E}}\left(V_k\right) \\
\operatorname{Cov}\left(\pmb{S}\right) \,=\,
\operatorname{Cov}\left(\pmb{S},\, \pmb{S}\right) \,&=\, \operatorname{\mathbb{E}}\left(\pmb{S}, \,\pmb{S}^T\right) \,-\, \operatorname{\mathbb{E}}\left(\pmb{S}\right)\operatorname{\mathbb{E}}\left(\pmb{S}\right)^T
\end{align}
\]
By the way, for two different random vectors X and Y the matrix
\operatorname{Cov}\left(\pmb{X},\, \pmb{Y}\right) \,=\, \operatorname{\mathbb{E}}\left(\pmb{X}, \,\pmb{Y}^T\right) \,-\, \operatorname{\mathbb{E}}\left(\pmb{X}\right)\operatorname{\mathbb{E}}\left(\pmb{Y}\right)^T
\]
is called the cross-covariance matrix.
Remark on independence and correlation
Two univariate random variables X and Y are statistically (!) independent when their joint probability, i.e. their probability for the occurrence of a value pair (x,y) is the product of the individual probabilities for the occurrence of x and y:
p_{X,Y}(x,y) \,=\, p_X(x)*p_Y(y)
\]
However, uncorrelatedness does mean something else, namely that the covariance of the distributions is 0:
\operatorname{Cov}\left(X,\, Y\right) \,&=\, 0 \\
\Rightarrow \: \operatorname{\mathbb{E}}\left(X Y\right) \,&=\, \operatorname{\mathbb{E}}\left(X\right) \operatorname{\mathbb{E}}\left(Y\right)
\end{align}
\]
Independence and uncorrelatedness are two different (!) things and uncorrelatedness does in general not imply statistical independence!
Thus the uncorrelatedness of a pair of components (Sj, Sk) of a random vector S does unfortunately not automatically imply their independence!
We shall later in this series that this is actually a bit different for a MND.
Conclusion
In this post we have collected and clarified some basic terms which we will use in forthcoming texts about multivariate normal distributions. An important step was to describe a statistical vector distribution in form of a random vector. We also have seen that a random vector can be subject to linear transformations.
We have discussed a limiting transition to probability density functions for statistical vector distributions. We understood that dependencies and correlations may make it difficult to derive the form of the probability density for the random vector from the probability density functions for the vector components.
In addition we have defined the mean value and the covariance matrix for a random vector. We saw how a linear transformation affects the covariance matrix.
In the next post we will apply our knowledge to a simple random vector based on Gaussian probability density functions without correlations. We will study what effects orthogonal and shearing transformations have on such a distribution.