In my last article of my introductory series on “Convolutional Neural Networks” [CNNs] I described how we can visualize the output of different maps at convolutional (or pooling) layers of a CNN.
A simple CNN for the MNIST dataset – IV – Visualizing the activation output of convolutional layers and maps
A simple CNN for the MNIST dataset – III – inclusion of a learning-rate scheduler, momentum and a L2-regularizer
A simple CNN for the MNIST datasets – II – building the CNN with Keras and a first test
A simple CNN for the MNIST datasets – I – CNN basics
We are now well equipped to look a bit closer at the maps of a trained CNN. The output of the last convolutional layer is of course of special interest: It is fed (in the form of a flattened input vector) into the MLP-part of the CNN for a classification analysis. As an MLP detects “patterns” the question arises whether we actually can see common “patterns” in the visualized maps of different images belonging to the same class. In our case we shall have a look at the maps of different MNIST images of a handwritten “4”.
Note for my readers, 20.08.2020:
This article has recently been revised and completely rewritten. It required a much more careful description of what we mean by “patterns” and “features” – and what we can say about them when looking at images of activation outputs on higher convolutional layers. I also postponed a thorough “philosophical” argumentation against a humanized usage of the term “features” to a later article in this series.
Objectives
We saw already in the last article that the images of maps get more and more abstract when we move to higher convolutional layers – i.e. layers deeper inside a CNN. At the same time we loose resolution due to intermediate pooling operations. It is quite obvious that we cannot see much of any original “features” of a handwritten “4” any longer in a (3×3)-map, whose values are produced by a sequence of complex transformation operations.
Nevertheless people talk about “feature detection” performed by CNNs – and they refer to “features” in a very concrete and descriptive way (e.g. “eyes”, “spectacles”, “bows”). How can this be? What is the connection of abstract activation patterns in low resolution maps to original “features” of an image? What is meant when CNN experts claim that neurons of higher CNN layers are allegedly able to “detect features”?
We cannot give a full answer, yet. We still need some more Python programs tools. But, wat we are going to do in this article are three things:
- Objective 1: I will try to describe the assumed relation between maps and “features”. To start with I shall make a clear distinction between “feature” patterns in input images and patterns in and across the maps of convolutional layers. The rest of the discussion will remain a bit theoretical; but it will use the fact that convolutions at higher layers combine filtered results in specific ways to create new maps. For the time being we cannot do more. We shall actually look at visualizations of “features” in forthcoming articles of this series. Promised.
- Objective 2: We follow three different input images, each representing a “4”, as they get processed from one convolutional layer to the next convolutional layer of our CNN. We shall compare the resulting outputs of all feature maps at each convolutional layer.
- Objective 3: We try to identify common “patterns” for our different “4” images across the maps of the highest convolutional layer.
We shall visualize each “map” by an image – reflecting the values calculated by the CNN-filters for all points in each map. Note that an individual value at a map point results from adding up many weighted values provided by the maps of lower layers and feeding the result into an activation function. We speak of “activation” values or “map activations”. So our 2-nd objective is to follow the map activations of an input image up to the highest convolutional layer. An interesting question will be if the chain of complex transformation operations leads to visually detectable similarities across the map outputs for the different images of a “4”.
The eventual classification of a CNN is done by its embedded MLP which analyzes information collected at the last convolutional layer. Regarding this input to the MLP we can make the following statements:
The convolutions and pooling operations project information of relatively large parts of the original image into a representation space of very low dimensionality. Each map on the third layer provides a 3×3 value tensor, only. However, we combine the points of all (128) maps together in a flattened input vector to the MLP. This input vector consists of more nodes than the original image itself.
Thus the sequence of convolutional and pooling layers in the end transforms the original images into another representation space of somewhat higher dimensionality (9×128 vs. 28×28). This transformation is associated with the hope that in the new representation space a MLP may find patterns which allow for a better classification of the original images than a direct analysis of the image data. This explains objective 3: We try to play the MLPs role by literally looking at the eventual map activations. We try to find out which patterns are representative for a “4” by comparing the activations of different “4” images of the MNIST dataset.
Enumbering the layers
To distinguish a higher Convolutional [Conv] or Pooling [Pool] layer from a lower one we give them a number “Conv_N” or “Pool_N”.
Our CNN has a sequence of
- Conv_1 (32 26×26 maps filtering the input image),
- Pool_1 (32 13×13 maps with half the resolution due to max-pooling),
- Conv_2 (64 11×11 maps filtering combined maps of Pool_1),
- Pool_2 (64 5×5 maps with half the resolution due to max-pooling),
- Conv_3 (128 3×3 maps filtering combined maps of Pool_2).
Patterns in maps?
We have seen already in the last article that the “patterns” which are displayed in a map of a higher layer Conv_N, with N ≥ 2, are rather abstract ones. The images of the maps at Conv_3 do not reflect figurative elements or geometrical patterns of the input images any more – at least not in a directly visible way. It does not help that the activations are probably triggered by some characteristic pixel patterns in the original images.
The convolutions and the pooling operation transform the original image information into more and more abstract representation spaces of shrinking dimensionality and resolution. This is due to the fact that the activation of a point in a map on a layer Conv_(N+1) results
- from a specific combination of multiple maps of a layer Conv_N or Pool_N
- and from a loss of resolution due to intermediate pooling.
It is not possible to directly guess in what way active points or activated areas within
a certain map at the third convolutional layer relate to or how they depend on “original and specific patterns in the input image”. If you do not believe me: Well, just look at the maps of the 3rd convolutional layer presented in the last article and tell me: What patterns in the initial image did these maps react to? Without some sophisticated numerical experiments you won’t be able to figure that out.
Patterns in the input image vs. patterns within and across maps
The above remarks indicate already that “patterns” may occur at different levels of consideration and abstraction. We talk about patterns in the input image and patterns within as well as across the maps of convolutional (or pooling) layers. To avoid confusion I already now want to make the following distinction:
- (Original) input patterns [OIP]: When I speak of (original) “input patterns” I mean patterns or figurative elements in the input image. In more mathematical terms I mean patterns within the input image which correspond to a kind of fixed and strong correlation between the values of pixels distributed over a sufficiently well defined geometrical area with a certain shape. Examples could be line-like elements, bow segments, two connected circles or combined rectangles. But OIPs may be of a much more complex and abstract kind and consist of strange sub-features – and they may not reflect a real world entity or a combination of such entities. An OIP may reside at one or multiple locations in different input images.
- Filter correlation patterns [FCP]: A CNN produces maps by filtering input data (Conv level 1) or by filtering maps of a lower layer and combining the results. By doing so a higher layer may detect patterns in the filter results of a lower layer. I call a pattern across the maps of a convolutional or pooling layer Conv_N or Pool_N as seen by Conv_(N+1) a FCP.
Note: Because a 3×3 filter for a map of Conv_(N+1) has fixed parameters per map of the previous layer Conv_N or Pool_N, it combines multiple maps (filters) of Conv_N in a specific, unique way.
Anybody who ever worked with image processing and filters knows that combining basic filters may lead to the display of weirdly looking, combined information residing in complex regions on the original image. E.g., a certain combination of filters may emphasize diagonal lines or bows with some distance in between and suppress all other features. Therefore, it is at least plausible that a map of a higher convolutional layer can be translated back to an OIP. Meaning:
A high activation of certain or multiple points inside a map on Conv_3 may reflect some typical OIP pattern in the input image.
But: At the moment we have no direct proof for such an idea. And it is not at all obvious what kind of OIP pattern this may be for a distinct map – and whether it can directly be described in terms of basic geometrical elements of a figurative number representation in the MNIST case. By just looking at the maps of a layer and their activated points we do not get any clue about this.
If, however, activated maps somehow really correspond to OIPs then a FCP over multiple maps may be associated with a combination of distinct OIPs in an input image.
What are “features” then?
In many textbooks maps are also called “feature maps“. As far I understand it the authors call a “feature” what I called an OIP above. By talking about a “feature” the authors most often refer to a pattern which a CNN somehow detects or identifies in the input images.
Typical examples of “features” text-book authors often discuss and even use in illustrations are very concrete: ears, eyes, feathers, wings, a mustache, leaves, wheels, sun-glasses … I.e., a lot of authors typically name features which human beings identify as physical entities or as entities, for which we have clear conceptual ideas in our mind. I think such examples trigger ideas about CNNs which are too far-fetched and which “humanize” stupid algorithmic processes.
The arguments in favor of the detection of features in the sense of conceptual entities are typically a bit nebulous – to say the least. E.g. in a relatively new book on “Generative Deep Learning” you see a series of CNN neuron layers associated with rather dubious and unclear images of triangles etc. and at the last convolutional layer we suddenly see pretty clear sketches of a mustache, a certain hairdress, eyes, lips, a shirt, an ear .. “. The related text goes like follows (I retranslated the text from the German version of the book): “Layer 1 consists of neurons which activate themselves stronger, when they recognize certain elementary and basic features in the input image, e.g. borders. The output of these neurons is then forwarded to the neurons of layer 2 which can use this information to detect more complex features – and so on across the following layers.” Yeah, “neurons activate themselves” as they “recognize” features – and suddenly the neurons at a high enough layer see a “spectacle”. 🙁
I think it would probably be more correct to say the following:
The activation of a map of a high convolutional layer may indicate the appearance of some kind of (complex) pattern or a sequence of patterns within an input image, for which a specific filter combination produces relatively high values in a low dimensional output space.
Note: At our level of analyzing CNNs even this carefully formulated idea is speculation. Which we will have to prove somehow … Where we stand right now, we are unfortunately not yet ready to identify OIPs or repeated OIP sequences associated with maps. This will be the topic of forthcoming articles.
It is indeed an interesting question whether a trained CNN “detects” patterns in the sense of entities with an underlying “concept”. I would say: Certainly not. At least not pure CNNs. I think, we should be very careful with the use of the term “feature”. Based on the filtering convolutions perform we might say:
A “feature” (hopefully) is a pattern in the sense of defined geometrical pixel correlation in an image.
Not more, not less. Such a “feature” may or may not correspond to entities, which a human being could identify and for which he or she has a concept for. A feature is just a pixel correlation whose appearance triggers output neurons in high level maps.
By the way there are 2 more points regarding the idea of feature detection:
- A feature or OIP may be located at different places in different images of something like a “5”. Due to different sizes of the depicted “5” and translational effects. So keep in mind that if maps do indeed relate to features it has to be explained how convolutional filtering can account for any translational invariance of the “detection” of a pattern in an image.
- The concrete examples given for “features” by many authors imply that the features are more or less the same for two differently trained CNNs. Well, regarding the point that training corresponds to finding a minimum on a rather complex multidimensional hyperplane this raises the question how well defined such a (global) minimum really is and whether it or other valid side minima are approached.
Keep these points in mind until we come back to related experiments in further articles.
From “features” to FCPs on the last Conv-layer?
However and independent of how a CNN really reacts to OIPs or “features”, we should not forget the following:
In the end a CNN – more precisely its embedded MLP – reacts to FCPs on the last convolutional level. In our CNN an FCP on the third convolutional layer with specific active points across 128 (3×3)-maps obviously can obviously tell the MLP something about the class an input image belongs to: We have proven already that the MLP part of our simple CNN guesses the class the original image belongs to with a surprisingly high accuracy. And by construction it obviously does so by just analyzing the 128 (3×3)-activation values of the third layer – arranged into a flattened vector.
From a classification point of view it, therefore, seems to be legitimate to look out for any FCP across the maps on Conv_3. As we can visualize the maps it is reasonable to literally look for common activation patterns which different images of handwritten “4”s may trigger on the maps of the last convolutional level. The basic idea behind this experimental step is:
OIPs which are typical for images of a “4” trigger and activate certain maps or points within certain maps. Across all maps we then may see a characteristic FCP for a “4”, which not only a MLP but also we intelligent humans could identify.
Or: Multiple characteristic features in images of a “4” may trigger characteristic FCPs which in turn can be used indicators of a class an image belongs to by an MLP. Well, let us see how far we get with this kind of theory.
Levels of “abstractions”
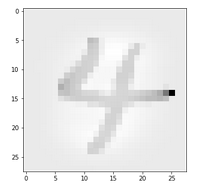
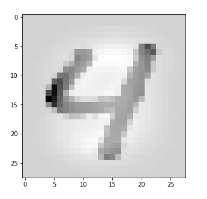
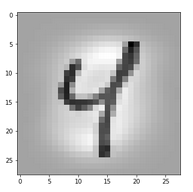
Let us take a MNIST image which represents something which a European would consider to be a clear representation of a “4”.
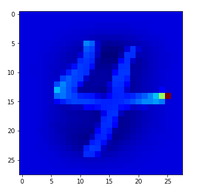
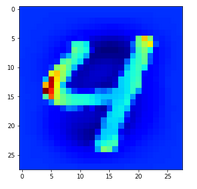
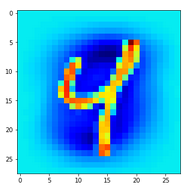
In the second image I used the “jet”-color map; i.e. dark blue indicates a low intensity value while colors from light blue to green to yellow and red indicate growing intensity values.
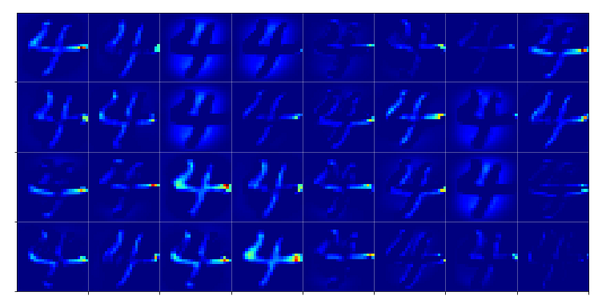
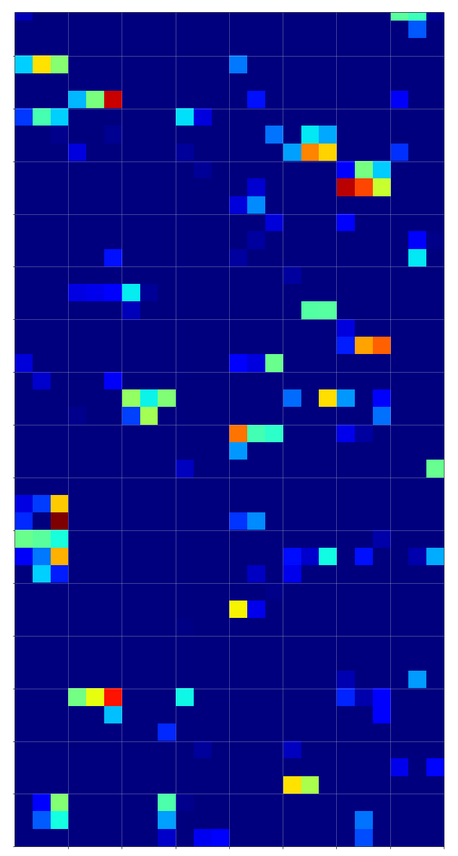
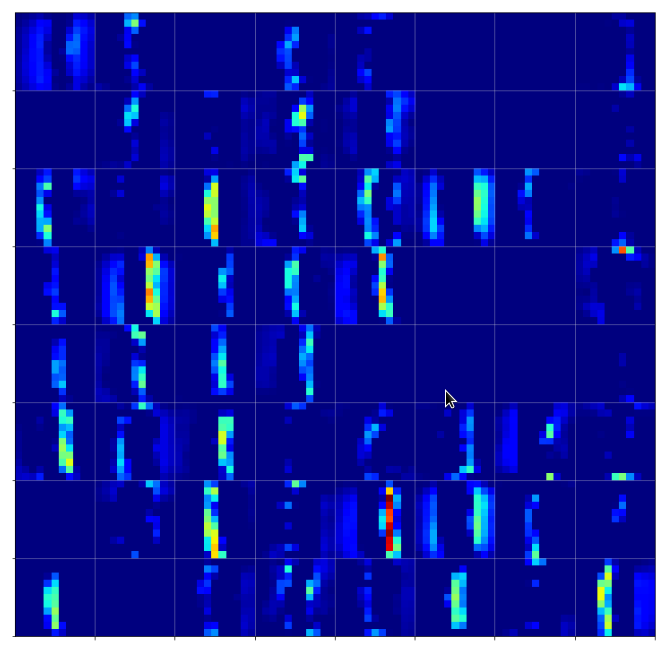
The first conv2D-layer (“Conv2d_1”) produces the following 32 maps of my chosen “4”-image after training:
We see that the filters, which were established during training emphasize general contours but also focus on certain image regions. However, the original “4” is still clearly visible on very many maps as the convolution does not yet reduce resolution too much.
By the way: When looking at the maps the first time I found it surprising that the application of a simple linear 3×3 filter with stride 1 could emphasize an overall oval region and suppress the pixels which formed the “4” inside of this region. A closer look revealed however that the oval region existed already in the original image data. It was emphasized by an inversion of the pixel values …
Pooling
The second Conv2D-layer already combines information of larger areas of the image – as a max (!) pooling layer was applied before. We loose resolution here. But there is a gain, too: the next convolution can filter (already filtered) information over larger areas of the original image.
But note: In other types of more advanced and modern CNNs pooling only is involved after two or more successive convolutions have happened. The direct succession of convolutions corresponds to a direct and unique combination of filters at the same level of resolution.
The 2nd convolution
As we use 64 convolutional maps on the 2nd layer level we allow for a multitude of different new convolutions. It is to be understood that each new map at the 2nd cConv layer is the result of a special unique combination of filtered information of all 32 previous maps (of Pool_1). Each of the previous 32 maps contributes through a specific unique filter and respective convolution operation to a single specific map at layer 2. Remember that we get 3×3 x 32 x 64 parameters for connecting the maps of Pool_1 to maps of Conv_2. It is this unique combination of already filtered results which enriches the analysis of the original image for more complex patterns than just the ones emphasized by the first convolutional filters.
As the max-condition of the pooling layer was applied first and because larger areas are now analyzed we are not too astonished to see that the filters dissolve the original “4”-shape and indicate more general geometrical patterns – which actually reflect specific correlations of map patterns on layer Conv_1.
I find it interesting that our “4” triggers more horizontally activations within some maps on this already abstract level than vertical ones. One should not confuse these patterns with horizontal patterns in the original image. The relation of original patterns with these activations is already much more complex.
The third convolutional layer applies filters which now cover almost the full original image and combine and mix at the same time information from the already rather abstract results of layer 2 – and of all the 64 maps there in parallel.
We again see a dominance of horizontal patterns. We see clearly that on this level any reference to something like an arrangement of parallel vertical lines crossed by a horizontal line is completely lost. Instead the CNN has transformed the original distribution of black (dark grey) pixels into multiple abstract configuration spaces with 2 axes, which only coarsely reflecting the original image area – namely by 3×3 maps; i.e. spaces with a very poor resolution.
What we see here are “correlations” of filtered and transformed original pixel clusters over relatively large areas. But no constructive concept of certain line arrangements.
Now, if this were the level of “FCP-patterns” which the MLP-part of the CNN uses to determine that we have a “4” then we would bet that such abstract patterns (active points on 9×9 grids) appear in a similar way on the maps of the 3rd Conv layer for other MNIST images of a “4”, too.
Well, how similar do different map representations of “4”s look like on the 3rd Conv2D-layer?
What makes a four a four in the eyes of the CNN?
The last question corresponds to the question of what activation outputs of “4”s really have in common. Let us take 3 different images of a “4”:
The same with the “jet”-color-map:
Already with our eyes we see that there are similarities but also quite a lot of differences.
Different “4”-representations on the 2nd Conv-layer
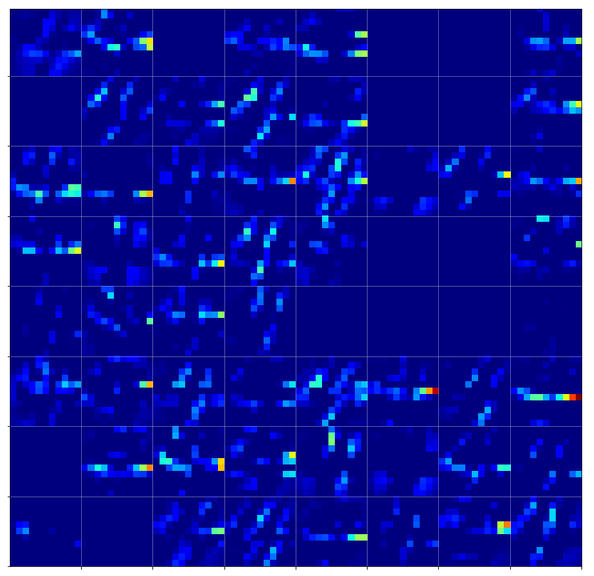
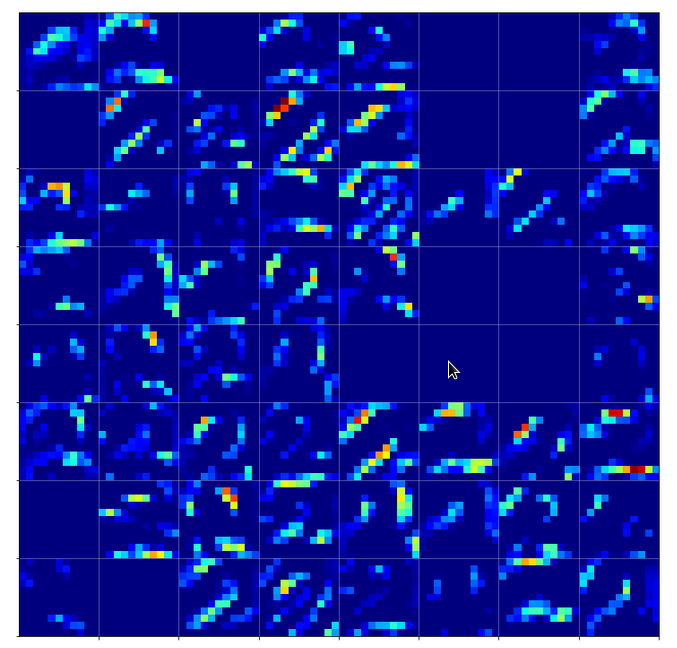
Below we see comparison of the 64 maps on the 2nd Conv-layer for our three “4”-images.
If you move your head backwards and ignore details you see that certain maps are not filled in all three map-pictures. Unfortunately, this is no common feature of “4”-representations. Below you see images of the activation of a “1” and a “2”. There the same maps are not activated at all.
We also see that on this level it is still important which points within a map are activated – and not which map on average. The original shape of the underlying number is reflected in the maps’ activations.
Now, regarding the “4”-representations you may say: Well, I still recognize some common line patterns – e.g. parallel lines in a certain 75 degree angle on the 11×11 grids. Yes, but these lines are almost dissolved by the next pooling step:
Consider in addition that the next (3rd) convolution combines 3×3-data of all of the displayed 5×5-maps. Then, probably, we can hardly speak of a concept of abstract line configurations any more …
“4”-representations on the third Conv-layer
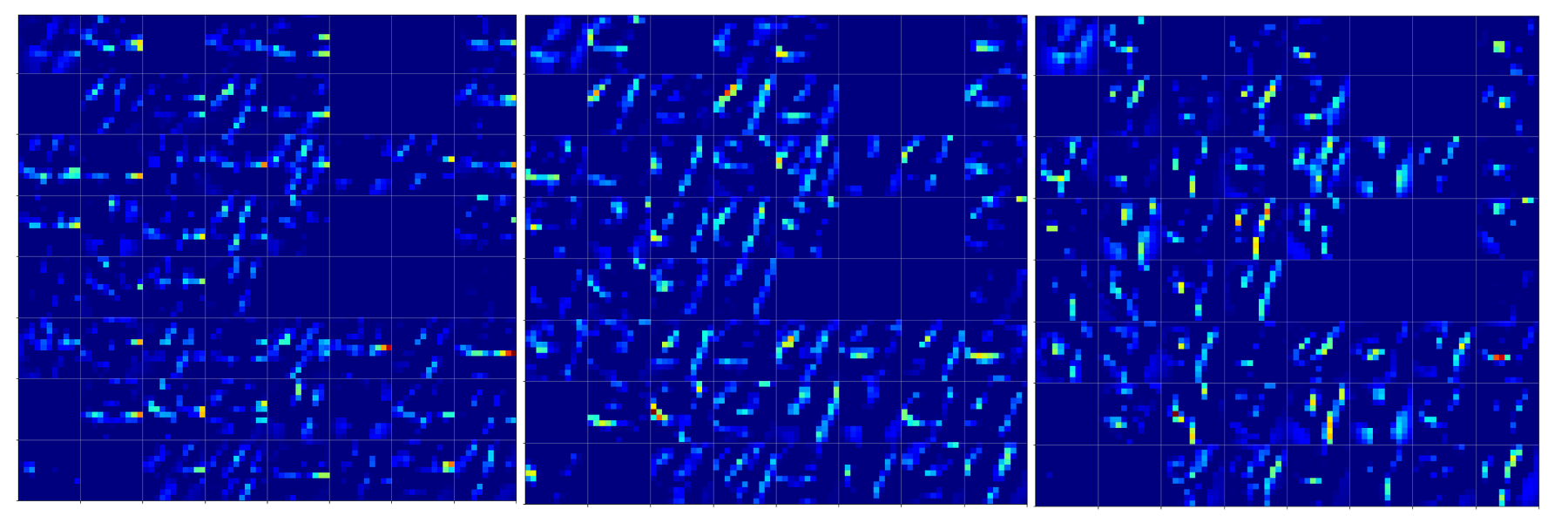
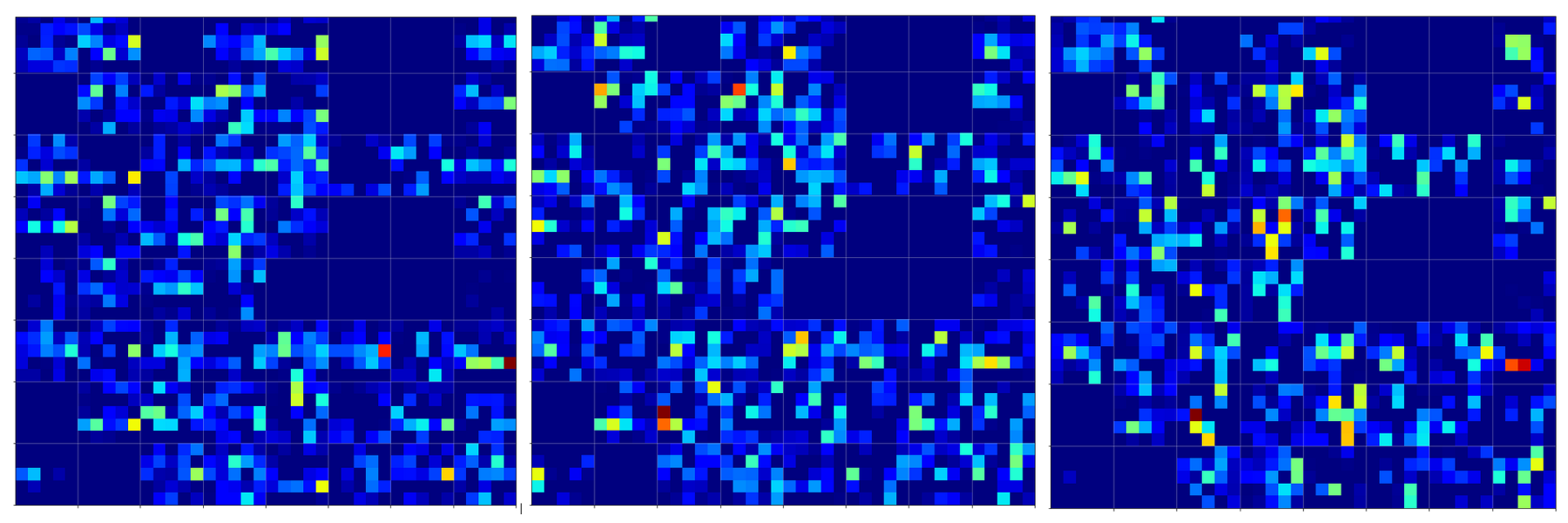
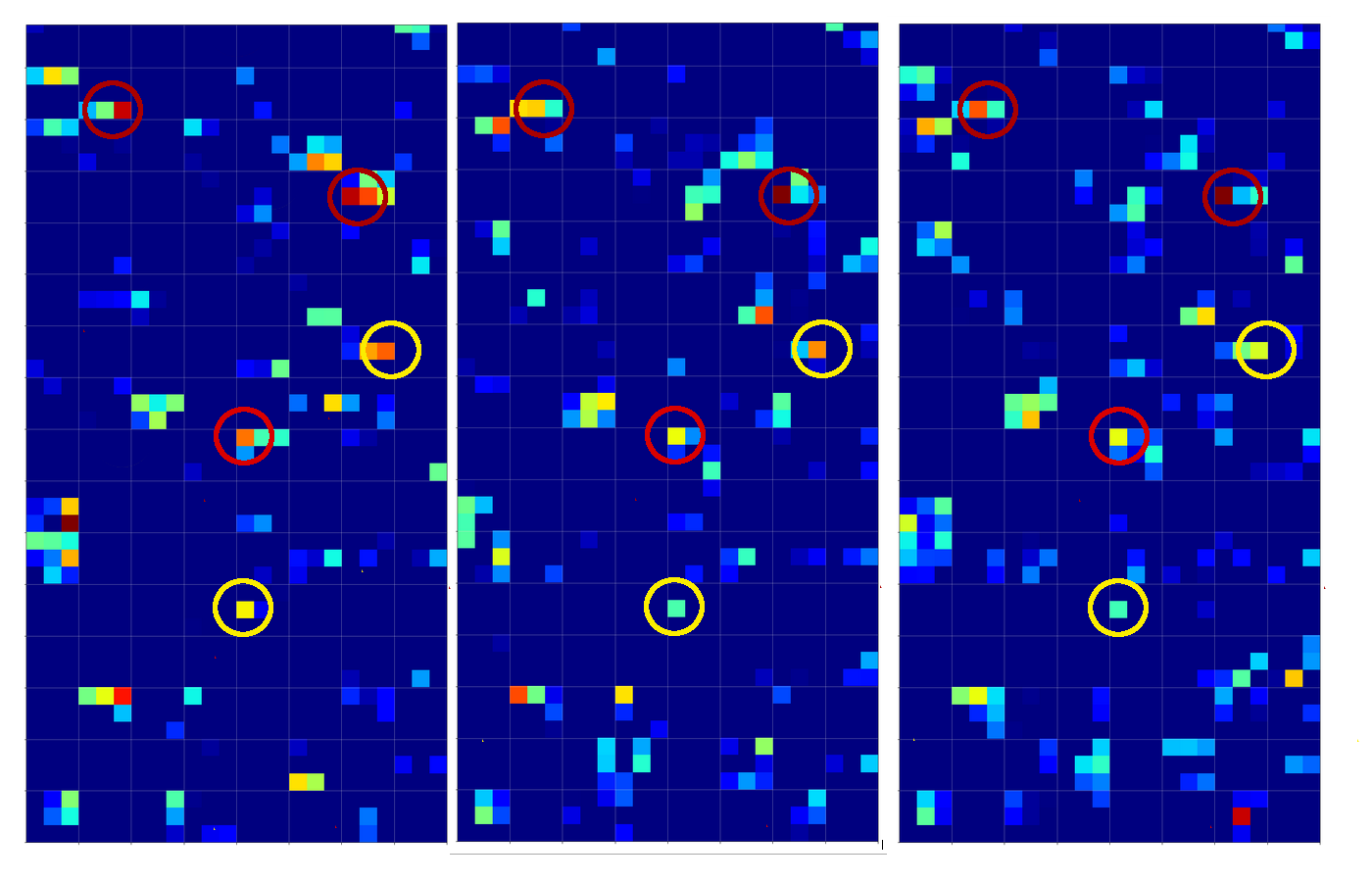
Below you find the activation outputs on the 3rd Conv2D-layer for our three different “4”-images:
When we look at details we see that prominent “features” in one map of a specific 4-image do NOT appear in a fully comparable way in the eventual convolutional maps for another image of a “4”. Some of the maps (i.e. filters after 4 transformations) produce really different results for our three images.
But there are common elements, too: I have marked only some of the points which show a significant intensity in all of the maps. But does this mean these individual common points are decisive for a classification of a “4”? We cannot be sure about it – probably it is their combination which is relevant.
So, what we ended up with is that we find some common points or some common point-relations in a few of the 128 “3×3”-maps of our three images of handwritten “4”s.
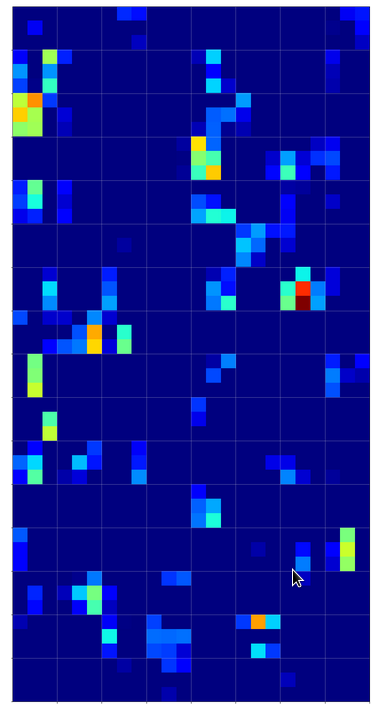
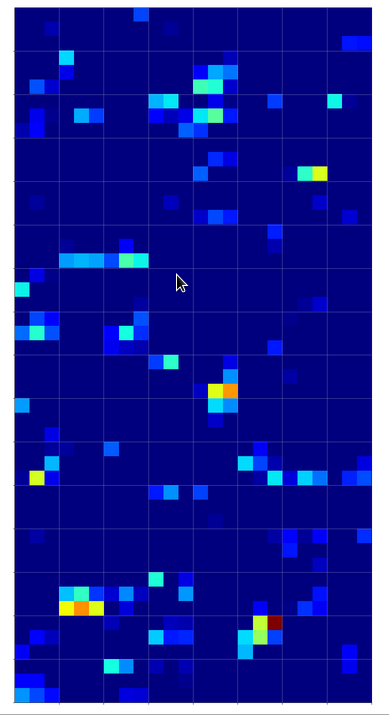
But how does this compare with maps of images of other digits? Well, look at he maps on the 3rd layer for images of a “1” and a “2” respectively:
On the 3rd layer it becomes more important which maps are not activated at all. But still the activation patterns within certain maps seem to be of importance for an eventual classification.
Conclusion
The maps of a CNN are created by an effective and guided optimization process. The results indicate the eventual detection of rather abstract patterns within and across filter maps on higher convolutional layers.
But these patterns (FCP-patterns) should not be confused with figurative elements or “features” in the original input images. Activation patterns at best vaguely remind of the original image features. At our level of analysis of a CNN we can only speculate about some correspondence of map activations with original features or patterns in an input image.
But it seems pretty clear that patterns in or across maps do not indicate any kind of constructive concept which describes how to build a “4” from underlying more elementary features in the sense of combine-able independent entities. There is no sign of conceptual constructive idea of how to denote a “4”. At least not in pure CNNs … Things may be a bit different in convolutional “autoencoders” (combinations of convolutional encoders and decoders), but this is another story we will come back to in this blog. Right now we would say that abstract (FCP-) patterns in maps of higher convolutional layers result from intricate filter combinations. These filters may react to certain patterns in an input image – but whether these patterns correspond to entities a human being would use to write down and thereby construct a “4” or an “8” is questionable.
We saw that the abstract information maps at the third layer of our CNN do show some common elements between the images belonging to the same class – and delicate differences with respect to activations resulting from images of other classes. However, the differences reside in details and the situation remains complicated. In the end the MLP-part of a CNN still has a lot of work to do. It must perform its classification task based on the correlation or anti-correlation of “point”-like elements in a multitude of maps – and probably even based on the activation level (i.e. output numbers) at these points.
This is seemingly very different from a conscious consideration process and weighing of alternatives which a human brain performs when it looks at sketches of numbers. When in doubt our brain tries to find traces consistent with a construction process defined for writing down a “4”, i.e. signs of a certain arrangement of straight and curved lines. A human brain, thus, would refer to arrangements of line elements, bows or circles – but not to relations of individual points in an extremely coarse and abstract representation space after some mathematical transformations. You may now argue that we do not need such a process when looking at clear representations of a “4” – we look and just know that its a “4”. I do not doubt that a brain may use maps, too – but I want to point out that a conscious intelligent thought process and conceptual ideas about entities involve constructive operations and not just a passive application of filters. Even from this extremely simplifying point of view CNNs are stupid though efficient algorithms. And authors writing about “features” should avoid any kind of a humanized interpretation.
In the next article
we shall look at the whole procedure again, but then we compare common elements of a “4” with those of a “9” on the 3rd convolutional layer. Then the key question will be: ” What do “4”s have in common on the last convolutional maps which corresponding activations of “9”s do not show – and vice versa.
This will become especially interesting in cases for which a distinction was difficult for pure MLPs. You remember the confusion matrix for the MNIST dataset? See:
A simple Python program for an ANN to cover the MNIST dataset – XI – confusion matrix
We saw at that point in time that pure MLPs had some difficulties to distinct badly written “4”s from “9s”. We will see that the better distinction abilities of CNNs in the end depend on very few point like elements of the eventual activation on the last layer before the MLP.
Further articles in this series
A simple CNN for the MNIST dataset – VII – outline of steps to visualize image patterns which trigger filter maps
A simple CNN for the MNIST dataset – VI – classification by activation patterns and the role of the CNN’s MLP part