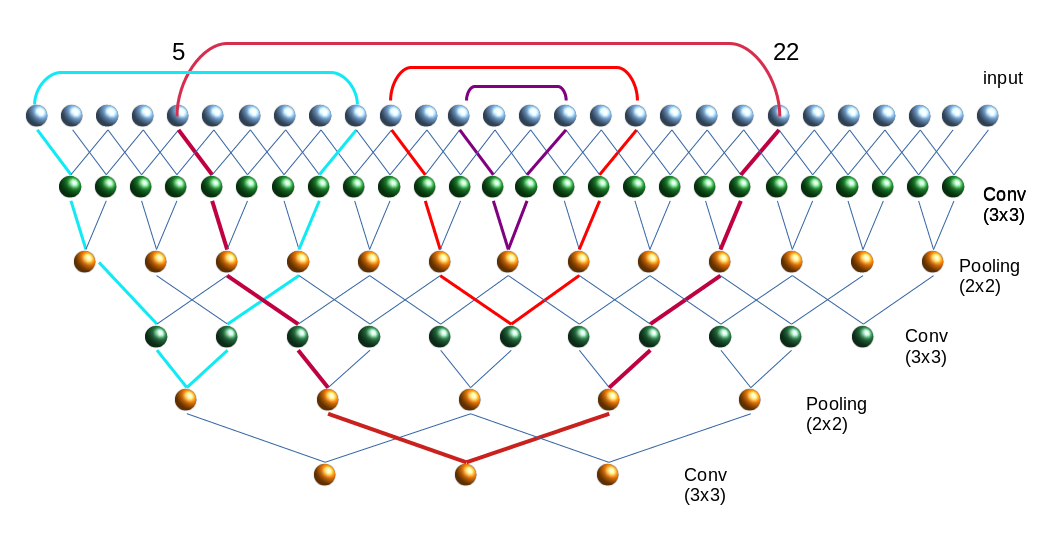
In the last article of this series
A simple CNN for the MNIST datasets – II – building the CNN with Keras and a first test
A simple CNN for the MNIST datasets – I – CNN basics
we saw that we could easily create a Convolutional Neural Network [CNN] with Keras and apply it to the MNIST dataset. But this was only a quick win. Readers who followed me through the MLP-oriented article series may have noticed that we are far, yet, from having the full control over a variety of things we would use to optimize the classification algorithm. After our experience with MLPs you may rightfully argue that
- both a systematic reduction of the learning rate in the vicinity of a (hopefully) global minimum of the multidimensional loss function hyperplane in the {weights, costs}-space
- and the use of a L2- or L1-regularizer
could help to squeeze out the last percentage of accuracy of a CNN. Actually, in our first test run we did not even care about how the RMSProp algorithm invoked a learning rate and what the value(s) may have been. A learning rate “lr” was no parameter of our CNN setup so far. The same is true for the use and parameterization of a regularizer.
I find it interesting that some of the introductory books on “Deep Learning” with Keras do not discuss the handling of learning rates beyond providing a fixed (!) rate parameter on rare occasions to a SGD- or RMSProp-optimizer. Folks out there seem to rely more on the type and basic settings of an optimizer than on a variable learning rate. Within the list of books I appended to the last article, only the book of A. Geron gives you some really useful hints. So, we better find information within the online documentation of the Keras framework to get control over learning rates and regularizers.
Against my original plans, the implementation of schedulers for the learning rate and the usage of a L2-regularizer will be the main topic of this article. But I shall – as a side-step – also cover the use of “callbacks” for some interactive plotting during a training run. This is a bit more interesting than just plotting key data (loss, accuracy) after training – as we did for MLPs.
Learning rate scheduler and momentum
A systematic reduction of the learning rate or the momentum can be handled with Keras via a parameterization of a scheduler. Such a scheduler can e.g. be invoked by a chosen optimizer. As the optimizer came into our CNN with the model.compile()-function we would not be astonished, if we had to include the scheduler there, too. The Keras version of Tensorflow 2 offers a variety of (partially) configurable schedulers. And there are, in addition, multiple different ways how to integrate a systematic decline of the learning rate into a Keras CNN model. I choose a very simple method for this article, which is, however, specific for the TF2 variant of Keras (tensorflow.keras). The book of Geron discusses other alternatives (e.g. the invocation of a scheduler via a so called callback for batches; see below).
The optimizer for the gradient calculation directly accepts parameters for an (initial and constant) learning rate (parameter “learning_rate”) and a momentum (parameter “momentum”). An example with the SGD-optimizer would be:
optimizer = keras.optimizers.SGD(learning_rate=0.001, momentum=0.5) cnn.compile(optimizer=optimizer, ....
If you just provide a parameter “learning_rate=0.001” then the optimizer will use a constant learning rate. However, if we provide a scheduler object – an instance of a scheduler class – then we can control a certain decline of the learning rate. What scheduler classes are available? You find them here:
keras.io/api/optimizers/ learning rate schedules/
A simple scheduler which allows for the “power scheduling” which we implemented for our MLPs is the scheduler “InverseTimeDecay“. It can be found under the library branch optimizers.schedules. We take this scheduler as an example in this article.
But how do we deliver the scheduler object to the optimizer? With tensorflow.keras and TF > 2.0 this is simply done by providing it as the parameter (object) for the learning_rate to the optimizer, as e.g. i the following code example:
from tensorflow.keras import optimizers from tensorflow.keras.optimizers import schedules ... ... scheduled_lr_rate = schedules.InverseTimeDecay(lr_init, lr_decay_steps, lr_decay_rate) optimizer=optimizers.RMSprop(learning_rate=scheduled_lr_rate, momentum=my_momentum) ...
Here “lr_init” defines an initial learning rate (as 0.001), “lr_decay_steps” a number of steps, after which the rate is changed, and lr_decay_rate a decay rate to be applied in the formula of the scheduler. Note that different schedulers take different parameters – so look them up before applying them. Also check that your optimizer accepts a scheduler object as a parameter …
The scheduler instance works as if we had a function returning an output – a learning rate – for an input, which corresponds to a number of “steps“:
learning_rate=scheduler(steps) .
Now what do we mean by “steps”? Epochs? No, actually a step here corresponds practically to the result of an iterator over batches. Each optimization “step” in the sense of a weight adjustment after a gradient analysis for a mini-batch defines a step for the scheduler. If you want to change the learning rate on the level of epochs only, then you must either adjust the “decay_step”-parameter or write a so called “callback-function” invoked after each epoch and control the learning rate yourself.
Momentum
Note, by the way, that in addition to the scheduler we also provided a value for the “momentum” parameter an optimizer may use during the gradient evolution via some specific adaptive formula. How the real weight changes are calculated based on gradient development and momentum parameters is of course specific for an optimizer.
Required import statements
We use the Jupyter cells of the code we built in the last article as far as possible. We need to perform some more imports to work with schedulers, regularizers and plotting. You can exchange the contents of the first Jupyter cell with the following statements:
Jupyter Cell 1:
import numpy as np import scipy import time import sys from sklearn.preprocessing import StandardScaler import tensorflow as tf from tensorflow import keras as K from tensorflow.python.keras import backend as B from tensorflow.keras import models from tensorflow.keras import layers from tensorflow.keras import regularizers from tensorflow.keras import optimizers from tensorflow.keras.optimizers import schedules from tensorflow.keras.utils import to_categorical from tensorflow.keras.datasets import mnist from tensorflow.python.client import device_lib import matplotlib as mpl from matplotlib import pyplot as plt from matplotlib.colors import ListedColormap import matplotlib.patches as mpat import os
Then we have two unchanged cells:
Jupyter Cell 2:
#gpu = False
gpu = True
if gpu:
# GPU = True; CPU = False; num_GPU = 1; num_CPU = 1
GPU = True; CPU = False; num_GPU = 1; num_CPU = 1
else:
GPU = False; CPU = True; num_CPU = 1; num_GPU = 0
config = tf.compat.v1.ConfigProto(intra_op_parallelism_threads=6,
inter_op_parallelism_threads=1,
allow_soft_placement=True,
device_count = {'CPU' : num_CPU,
'GPU' : num_GPU},
log_device_placement=True
)
config.gpu_options.per_process_gpu_memory_fraction=0.35
config.gpu_options.force_gpu_compatible = True
B.set_session(tf.compat.v1.Session(config=config))
and
Jupyter Cell 3:
# load MNIST
# **********
#@tf.function
def load_Mnist():
mnist = K.datasets.mnist
(X_train, y_train), (X_test, y_test) = mnist.load_data()
#print(X_train.shape)
#print(X_test.shape)
# preprocess - flatten
len_train = X_train.shape[0]
len_test = X_test.shape[0]
X_train = X_train.reshape(len_train, 28*28)
X_test = X_test.reshape(len_test, 28*28)
#concatenate
_X = np.concatenate((X_train, X_test), axis=0)
_y = np.concatenate((y_train, y_test), axis=0)
_dim_X = _X.shape[0]
# 32-bit
_X = _X.astype(np.float32)
_y = _y.astype(np.int32)
# normalize
scaler = StandardScaler()
_X = scaler.fit_transform(_X)
# mixing the training indices - MUST happen BEFORE encoding
shuffled_index = np.random.permutation(_dim_X)
_X, _y = _X[shuffled_index], _y[shuffled_index]
# split again
num_test = 10000
num_train = _dim_X - num_test
X_train, X_test, y_train, y_test = _X[:num_train], _X[num_train:], _y[:num_train], _y[num_train:]
# reshape to Keras tensor requirements
train_imgs = X_train.reshape((num_train, 28, 28, 1))
test_imgs = X_test.reshape((num_test, 28, 28, 1))
#print(train_imgs.shape)
#print(test_imgs.shape)
# one-hot-encoding
train_labels = to_categorical(y_train)
test_labels = to_categorical(y_test)
#print(test_labels[4])
return train_imgs, test_imgs, train_labels, test_labels
if gpu:
with tf.device("/GPU:0"):
train_imgs, test_imgs, train_labels, test_labels= load_Mnist()
else:
with tf.device("/CPU:0"):
train_imgs, test_imgs, train_labels, test_labels= load_Mnist()
Include the regularizer via the layer definitions of the CNN
A “regularizer” modifies the loss function via a sum over contributions delivered by a (common) function of the weight at every node. In Keras this sum is split up into contributions at the different layers, which we define via the “model.add()”-function. You can build layers with or without regularization contributions. Already for our present simple CNN case this is very useful:
In a first trial we only want to add weights to the hidden and the output layers of the concluding MLP-part of our CNN. To do this we have to provide a parameter “kernel_regularizer” to “model.add()”, which specifies the type of regularizer to use. We restrict the regularizers in our example to a “l2”- and a “l1”-regularizer, for which Keras provides predefined functions/objects. This leads us to the following change of the function “build_cnn_simple()”, which we used in the last article:
Jupyter Cell 4:
# Sequential layer
model of our CNN
# ***********************************
# just for illustration - the real parameters are fed later
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
li_conv_1 = [32, (3,3), 0]
li_conv_2 = [64, (3,3), 0]
li_conv_3 = [64, (3,3), 0]
li_Conv = [li_conv_1, li_conv_2, li_conv_3]
li_pool_1 = [(2,2)]
li_pool_2 = [(2,2)]
li_Pool = [li_pool_1, li_pool_2]
li_dense_1 = [70, 0]
li_dense_2 = [10, 0]
li_MLP = [li_dense_1, li_dense_2]
input_shape = (28,28,1)
# important !!
# ~~~~~~~~~~~
cnn = None
x_optimizers = None
def build_cnn_simple(li_Conv, li_Pool, li_MLP, input_shape,
my_regularizer=None,
my_reg_param_l2=0.01, my_reg_param_l1=0.01 ):
use_regularizer = True
if my_regularizer == None:
use_regularizer = False
# activation functions to be used in Conv-layers
li_conv_act_funcs = ['relu', 'sigmoid', 'elu', 'tanh']
# activation functions to be used in MLP hidden layers
li_mlp_h_act_funcs = ['relu', 'sigmoid', 'tanh']
# activation functions to be used in MLP output layers
li_mlp_o_act_funcs = ['softmax', 'sigmoid']
# dictionary for regularizer functions
d_reg = {
'l2': regularizers.l2,
'l1': regularizers.l1
}
if use_regularizer:
if my_regularizer not in d_reg:
print("regularizer " + my_regularizer + " not known!")
sys.exit()
else:
regul = d_reg[my_regularizer]
if my_regularizer == 'l2':
reg_param = my_reg_param_l2
elif my_regularizer == 'l1':
reg_param = my_reg_param_l1
# Build the Conv part of the CNN
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
num_conv_layers = len(li_Conv)
num_pool_layers = len(li_Pool)
if num_pool_layers != num_conv_layers - 1:
print("\nNumber of pool layers does not fit to number of Conv-layers")
sys.exit()
rg_il = range(num_conv_layers)
# Define a sequential model
# ~~~~~~~~~~~~~~~~~~~~~~~~~
cnn = models.Sequential()
for il in rg_il:
# add the convolutional layer
num_filters = li_Conv[il][0]
t_fkern_size = li_Conv[il][1]
cact = li_conv_act_funcs[li_Conv[il][2]]
if il==0:
cnn.add(layers.Conv2D(num_filters, t_fkern_size, activation=cact, input_shape=input_shape))
else:
cnn.add(layers.Conv2D(num_filters, t_fkern_size, activation=cact))
# add the pooling layer
if il < num_pool_layers:
t_pkern_size = li_Pool[il][0]
cnn.add(layers.MaxPooling2D(t_pkern_size))
# Build the MLP part of the CNN
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
num_mlp_layers = len(li_MLP)
rg_im = range(num_mlp_layers)
cnn.add(layers.Flatten())
for im in rg_im:
# add the dense layer
n_nodes = li_MLP[im][0]
if im < num_mlp_layers - 1:
m_act = li_mlp_h_act_funcs[li_MLP[im][1]]
if use_regularizer:
cnn.add(layers.Dense(n_nodes, activation=m_act, kernel_regularizer=regul(reg_param)))
else:
cnn.add(layers.Dense(n_nodes, activation=m_act))
else:
m_act = li_mlp_o_act_funcs[li_MLP[im][1]]
if use_regularizer:
cnn.add(layers.Dense(n_nodes, activation=m_act, kernel_regularizer=regul(reg_param)))
else:
cnn.add(layers.Dense(n_nodes, activation=m_act))
return cnn
I have used a dictionary to enable an indirect call of the regularizer object. The invocation of a regularizer happens in
the statements:
cnn.add(layers.Dense(n_nodes, activation=m_act, kernel_regularizer=regul(reg_param)))
We only use a regularizer for MLP-layers in our example.
Note that I used predefined regularizer objects above. If you want to use a regularizer object defined and customized by yourself you can either define a simple regularizer function (which accepts the weights as a parameter) or define a child class of the “keras.regularizers.Regularizer”-class. You find hints how to do this at https://keras.io/api/layers/regularizers/ and in the book of Aurelien Geron (see the literature list at the end of the last article).
Although I have limited the use of a regularizer in the code above to the MLP layers you are of course free to apply a regularizer also to convolutional layers. (The book of A. Geron gives some hints how to avoid code redundancies in chapter 11).
Include a scheduler via a parameter to the optimizer of the CNN
As we saw above a scheduler for the learning rate can be provided as a parameter to the optimizer of a CNN. As the optimizer objects are invoked via the function “model.compile()”, we prepare our own function “my_compile()” with an appropriate interface, which then calls “model.compile()” with the necessary parameters. For indirect calls of predefined scheduler objects we again use a dictionary:
Jupyter Cell 5:
# compilation function for further parameters
def my_compile(cnn,
my_loss='categorical_crossentropy', my_metrics=['accuracy'],
my_optimizer='rmsprop', my_momentum=0.0,
my_lr_sched='powerSched',
my_lr_init=0.01, my_lr_decay_steps=1, my_lr_decay_rate=0.00001 ):
# dictionary for the indirct call of optimizers
d_optimizer = {
'rmsprop': optimizers.RMSprop,
'nadam': optimizers.Nadam,
'adamax': optimizers.Adamax
}
if my_optimizer not in d_optimizer:
print("my_optimzer" + my_optimizer + " not known!")
sys.exit()
else:
optim = d_optimizer[my_optimizer]
use_scheduler = True
if my_lr_sched == None:
use_scheduler = False
print("\n No scheduler will be used")
# dictionary for the indirct call of scheduler funcs
d_sched = {
'powerSched' : schedules.InverseTimeDecay,
'exponential': schedules.ExponentialDecay
}
if use_scheduler:
if my_lr_sched not in d_sched:
print("lr scheduler " + my_lr_sched + " not known!")
sys.exit()
else:
sched = d_sched[my_lr_sched]
if use_scheduler:
print("\n lr_init = ", my_lr_init, " decay_steps = ", my_lr_decay_steps, " decay_rate = ", my_lr_decay_rate)
scheduled_lr_rate = sched(my_lr_init, my_lr_decay_steps, my_lr_decay_rate)
optimizer = optim(learning_rate=scheduled_lr_rate, momentum=my_momentum)
else:
print("\n lr_init = ", my_lr_init)
optimizer=optimizers.RMSprop(learning_rate=my_lr_init, momentum=my_momentum)
cnn.compile(optimizer=optimizer, loss=my_loss, metrics=my_metrics)
return cnn
You see that, for the time being, I only offered the choice between the predefined schedulers “InverseTimeDecay” and “ExponentialDecay“. If no scheduler name is provided then only a constant learning rate is delivered to the chosen optimizer.
You find some basic information on optimizers and schedulers here:
https://keras.io/api/optimizers/
nhttps://keras.io/api/optimizers/ learning rate schedules/.
Note that you can build your own scheduler object via defining a child class of “keras.callbacks.LearningRateScheduler” and provide with a list of other callbacks to the “model.fit()“-function; see
https://keras.io/api/callbacks/ learning rate scheduler/.
or the book of Aurelien Geron for more information.
Two callbacks – one for printing the learning rate and an other for plotting during training
As I am interested in the changes of the learning rate with the steps during an epoch I want to print them after each epoch during training. In addition I want to plot key data produced during the training of a CNN model.
For both purposes we can use a convenient mechanism Keras offers – namely “callbacks”, which can either be invoked at the beginning/end of the treatment of a mini-batch or at the beginning/end of each epoch. Information on callbacks is provided here:
https://keras.io/api/callbacks/
https://keras.io/guides/ writing your own callbacks/
It is best to just look at examples to get the basic points; below we invoke two callback objects which provide data for us at the end of each epoch.
Jupyter Cell 6:
# Preparing some callbacks
# **************************
# Callback class to print information on the iteration and the present learning rate
class LrHistory(K.callbacks.Callback):
def __init__(self, use_scheduler):
self.use_scheduler = use_scheduler
def on_train_begin(self, logs={}):
self.lr = []
def on_epoch_end(self, epoch, logs={}):
if self.use_scheduler:
optimizer = self.model.optimizer
iterations = optimizer.iterations
present_lr = optimizer.lr(iterations)
else:
present_lr = optimizer.lr
self.lr.append(present_lr)
#print("\npresent lr:", present_lr)
tf.print("\npresent lr: ", present_lr)
tf.print("present iteration:", iterations)
# Callback class to support interactive printing
class EpochPlot(K.callbacks.Callback):
def __init__(self, epochs, fig1, ax1_1, ax2_2):
#def __init__(self, epochs):
self.fig1 = fig1
self.ax1_1 = ax1_1
self.ax1_2 = ax1_2
self.epochs = epochs
rg_i = range(epochs)
self.lin92 = []
for i in rg_i:
self.lin92.append(0.992)
def on_train_begin(self, logs={}):
self.train_loss = []
self.train_acc = []
self.val_loss = []
self.val_acc = []
def on_epoch_end(self, epoch, logs={}):
if epoch == 0:
for key in logs.keys():
print(key)
self.train_loss.append(logs.get('loss'))
self.train_acc.append(logs.get('accuracy'))
self.val_loss.append(logs.get('val_loss'))
self.val_acc.append(logs.get('val_accuracy'))
if len(self.train_acc) > 0:
self.ax1_1.clear()
self.ax1_1.set_xlim (0, self.epochs+1)
self.ax1_1.set_ylim (0.97, 1.001)
self.ax1_1.plot(self.train_acc, 'g' )
self.ax1_1.plot(self.val_acc, 'r' )
self.ax1_1.plot(self.lin92, 'b', linestyle='--' )
self.ax1_2.clear()
self.ax1_2.set_xlim (0, self.epochs+1)
self.ax1_2.set_ylim (0.0, 0.2)
self.ax1_2.plot(self.train_loss, 'g' )
self.ax1_2.plot(self.val_loss, 'r' )
self.fig1.canvas.draw()
From looking at the code we see that a callback can be defined as a child class of a base class “keras.callbacks.Callback” or of some specialized predefined callback classes listed under “https://keras.io/api/callbacks/“. They are all useful, but perhaps the classes “ModelCheckpoint”, “LearningRateScheduler” (see above) and “EarlyStopping” are of direct practical interest for people who want to move beyond the scope of this article. In the above example I, however, only used the base class “callbacks.Callback”.
Printing steps, iterations and the learning rate
Let us look at the first callback, which provides a printout of the learning rate. This is a bit intricate: Regarding the learning rate I have already mentioned that a scheduler changes it after each operation on a batch; such a step corresponds to a variable “iterations” which is counted up by the optimizer during training after the treatment of each mini-batch. [Readers from my series on MLPs remember that gradient descent can be based on a gradient evaluation over the samples of mini-batches – instead of using a gradient evaluation based on all samples (batch gradient descent or full gradient descent) or of each individual sample (stochastic gradient descent).]
As we defined a batch size = 64 for the fit()-method of our CNN in the last article the number of optimizer iterations (=steps) per epoch is quite big: 60000/64 => 938 (with a smaller last batch).
Normally, the constant initial learning rate of an optimizer could be retrieved in a callback by a property “lr” as in “self.model.optimizer.lr”. However, in our example this attribute now points to a method “schedule()” of the scheduler object. (See schedule() method of the LearningRateScheduler class). We must provide the number of “iterations” (= number of steps) as a parameter to this schedule()-function and take the returned result as the present lr-value.
Our own callback class “LrHistory(K.callbacks.Callback)” gets three methods which are called at different types of “events” (Javascript developers should feel at home 🙂 ):
- At initialization (__init__()) we provide a parameter which defines whether we used a scheduler at all, or a constant lr.
- At the beginning of the training (on_train_begin()) we just instantiate a list, which we later fill with retrieved lr-values; we could use this list for some evaluation e.g. at the end of the training.
- At the end of each epoch (on_epoch-end()) we determine the present learning rate via calling the scheduler (behind the “lr” attribute) – if we used one – for the number of iterations passed.
This explains the following statements:
optimizer = self.model.optimizer iterations = optimizer.iterations present_lr = optimizer.lr(iterations)
Otherwise we just print the constant learning rate used by the optimizer (e.g. a default value).
Note that we use the “tf.print()” method to do the printing and not the Python “print()” function. This is basically done for convenience reasons: We thus avoid a special evaluation of the tensor and a manual transformation to a Numpy value. Do not forget: All data in Keras/TF are basically tensors and, therefore, the Python print() function would print extra information besides the expected numerical value!
Plotting at the end of an epoch
With our MLP we never used interactive plots in our Jupyter notebooks. We always plotted after training based on arrays or lists filled during
the training iterations with intermediately achieved values of accuracy and loss. But seeing the evolution of key metric data during training is a bit more rewarding; we get a feeling for what happens and can stop a training run with problematic hyperparameters before we waste too much valuable GPU or CPU time. So, how do we do change plots during training in our Jupyter environment?
The first point is that we need to prepare the environment of our Jupyter notebook; we do this before we start the training run. Below you find the initial statements of the respective Jupyter cell; the really important statement is the call of the ion-function: “plt.ion()”. For some more information see
https://linux-blog.anracom.com/2019/12/26/matplotlib-jupyter-and-updating-multiple-interactive-plots/
and links at the end of the article.
# Perform a training run # ******************** # Prepare the plotting # The really important commands for interactive (=intermediate) plot updating %matplotlib notebook plt.ion() #sizing fig_size = plt.rcParams["figure.figsize"] fig_size[0] = 8 fig_size[1] = 3 # One figure # ----------- fig1 = plt.figure(1) #fig2 = plt.figure(2) # first figure with two plot-areas with axes # -------------------------------------------- ax1_1 = fig1.add_subplot(121) ax1_2 = fig1.add_subplot(122) fig1.canvas.draw()
The interesting statements are the first two. The statements for setting up the plot figures should be familiar to my readers already. The statement “fig1.canvas.draw()” leads to the display of two basic coordinate systems at the beginning of the run.
The real plotting is, however, done with the help of our second callback “EpochPlot()” (see the code given above for Jupyter cell 6). We initialize our class with the number of epochs for which our training run shall be performed. We also provide the addresses of the plot figures and their two coordinate systems (ax1_1 and ax1_2). At the beginning of the training we provide empty lists for some key data which we later (after each epoch) fill with calculated values.
Which key or metric data can the training loop of Keras provide us with?
Keras fills a dictionary “logs” with metric data and loss values. What the metric data are depends on the loss function. At the following link you find a discussion of the metrics you can use with respect to different kinds of ML problems: machinelearningmastery.com custom-metrics-deep-learning-keras-python/.
For categorical_crossentropy and our metrics list [‘accuracy’] we get the following keys
loss, accuracy, val_loss, val_accuracy
as we also provided validation data. We print this information out at epoch == 0. (You will note however, that epoch 0 is elsewhere printed out as epoch 1. So, the telling is a bit different inside Keras and outside).
Then have a look at the parameterization of the method on_epoch_end(self, epoch, logs={}). This should not disturb you; logs is only empty at the beginning, later on a filled logs-dictionary is automatically provided.
As you see form the code we retrieve the data from the logs-dictionary at the end of each epoch. The rest are plain matplot-commands The “clear()”-statements empty the coordinate areas; the statement “self.fig1.canvas.draw()” triggers a redrawing.
The new training function
To do things properly we now need to extend our training function which invokes the other already defined functions as needed:
Jupyter Cell 7:
# Training 2 - with test data integrated
# *****************************************
def train( cnn, build, train_imgs, train_labels,
li_Conv, li_Pool, li_MLP, input_shape,
reset=True, epochs=5, batch_size=64,
my_loss='categorical_crossentropy', my_metrics=['accuracy'],
my_regularizer=None,
my_reg_param_l2=0.01, my_reg_param_l1=0.01,
my_optimizer='rmsprop', my_momentum=0.0,
my_lr_sched=None,
my_lr_init=0.001, my_lr_decay_steps=1, my_lr_decay_rate=0.00001,
fig1=None, ax1_1=None, ax1_2=None
):
if build:
# build cnn layers - now with regularizer - 200603 by ralph
cnn = build_cnn_simple( li_Conv, li_Pool, li_MLP, input_shape,
my_regularizer = my_regularizer,
my_reg_param_l2 = my_reg_param_l2, my_reg_param_l1 = my_reg_param_l1)
# compile - now with lr_scheduler - 200603 by ralph
cnn = my_compile(cnn=cnn,
my_loss=my_loss, my_metrics=my_metrics,
my_optimizer=my_optimizer, my_momentum=my_momentum,
my_lr_sched=my_lr_sched,
my_lr_init=my_lr_init, my_lr_decay_steps=my_lr_decay_steps,
my_lr_decay_rate=my_lr_decay_rate)
# save the inital (!) weights to be able to restore them
cnn.save_weights('cnn_weights.h5') # save the initial weights
# reset weights(standard)
if reset and not build:
cnn.load_weights('cnn_weights.h5')
# Callback list
# ~~~~~~~~~~~~~
use_scheduler = True
if my_lr_sched == None:
use_scheduler = False
lr_history = LrHistory(use_scheduler)
callbacks_list = [lr_history]
if fig1 != None:
epoch_plot = EpochPlot(epochs, fig1, ax1_1, ax1_2)
callbacks_list.append(epoch_plot)
start_t = time.perf_counter()
if reset:
history = cnn.fit(train_imgs, train_labels, initial_epoch=0, epochs=epochs, batch_size=batch_size, verbose=1, shuffle=True,
validation_data=(test_imgs, test_labels), callbacks=callbacks_list)
else:
history = cnn.fit(train_imgs, train_labels, epochs=epochs, batch_size=batch_size, verbose=1, shuffle=True,
validation_data=(test_imgs, test_labels), callbacks=callbacks_list )
end_t = time.perf_counter()
fit_t = end_t - start_t
return cnn, fit_t, history, x_optimizer # we return cnn to be able to use it by other functions in the Jupyter later on
Note, how big our interface became; there are a lot of parameters for the control of our training. Our set of configuration parameters is now very similar to what we used for MLP training runs before.
Note also how we provided a list of our callbacks to the “model.fit()”-function.
Setting up a training run
Now , we are only a small step away from testing our modified CNN-setup. We only need one further cell:
Jupyter Cell 8:
# Perform a training run
# ********************
# Prepare the plotting
# The really important command for interactive (=interediate) plot updating
%matplotlib notebook
plt.ion()
#sizing
fig_size = plt.rcParams["figure.figsize"]
fig_size[0] = 8
fig_size[1] = 3
# One figure
# -----------
fig1 = plt.figure(1)
#fig2 = plt.figure(2)
# first figure with two plot-areas with axes
# --------------------------------------------
ax1_1 = fig1.add_subplot(121)
ax1_2 = fig1.add_subplot(122)
nfig1.canvas.draw()
# second figure with just one plot area with axes
# -------------------------------------------------
#ax2 = fig2.add_subplot(121)
#ax2_1 = fig2.add_subplot(121)
#ax2_2 = fig2.add_subplot(122)
#fig2.canvas.draw()
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
# Parameterization of the training run
build = False
build = True
if cnn == None:
build = True
x_optimizer = None
batch_size=64
epochs=80
reset = False # we want training to start again with the initial weights
my_loss ='categorical_crossentropy'
my_metrics =['accuracy']
my_regularizer = None
my_regularizer = 'l2'
my_reg_param_l2 = 0.0009
my_reg_param_l1 = 0.01
my_optimizer = 'rmsprop' # Present alternatives: rmsprop, nadam, adamax
my_momentum = 0.6 # momentum value
my_lr_sched = 'powerSched' # Present alternatrives: None, powerSched, exponential
#my_lr_sched = None # Present alternatrives: None, powerSched, exponential
my_lr_init = 0.001 # initial leaning rate
my_lr_decay_steps = 1 # decay steps = 1
my_lr_decay_rate = 0.001 # decay rate
li_conv_1 = [64, (3,3), 0]
li_conv_2 = [64, (3,3), 0]
li_conv_3 = [128, (3,3), 0]
li_Conv = [li_conv_1, li_conv_2, li_conv_3]
li_pool_1 = [(2,2)]
li_pool_2 = [(2,2)]
li_Pool = [li_pool_1, li_pool_2]
li_dense_1 = [120, 0]
#li_dense_2 = [30, 0]
li_dense_3 = [10, 0]
li_MLP = [li_dense_1, li_dense_2, li_dense_3]
li_MLP = [li_dense_1, li_dense_3]
input_shape = (28,28,1)
try:
if gpu:
with tf.device("/GPU:0"):
cnn, fit_time, history, x_optimizer = train( cnn, build, train_imgs, train_labels,
li_Conv, li_Pool, li_MLP, input_shape,
reset, epochs, batch_size,
my_loss=my_loss, my_metrics=my_metrics,
my_regularizer=my_regularizer,
my_reg_param_l2=my_reg_param_l2, my_reg_param_l1=my_reg_param_l1,
my_optimizer=my_optimizer, my_momentum = 0.8,
my_lr_sched=my_lr_sched,
my_lr_init=my_lr_init, my_lr_decay_steps=my_lr_decay_steps,
my_lr_decay_rate=my_lr_decay_rate,
fig1=fig1, ax1_1=ax1_1, ax1_2=ax1_2
)
print('Time_GPU: ', fit_time)
else:
with tf.device("/CPU:0"):
cnn, fit_time, history = train( cnn, build, train_imgs, train_labels,
li_Conv, li_Pool, li_MLP, input_shape,
reset, epochs, batch_size,
my_loss=my_loss, my_metrics=my_metrics,
my_regularizer=my_regularizer,
my_reg_param_l2=my_reg_param_l2, my_reg_param_l1=my_reg_param_l1,
my_optimizer=my_optimizer, my_momentum = 0.8,
my_lr_sched=my_lr_sched,
my_lr_init=my_lr_init, my_lr_decay_steps=my_lr_decay_steps,
my_lr_decay_rate=my_lr_decay_rate,
fig1=fig1, ax1_1=ax1_1, ax1_2=ax1_2
)
print('Time_CPU: ', fit_time)
except SystemExit:
print("stopped due to exception")
n
Results
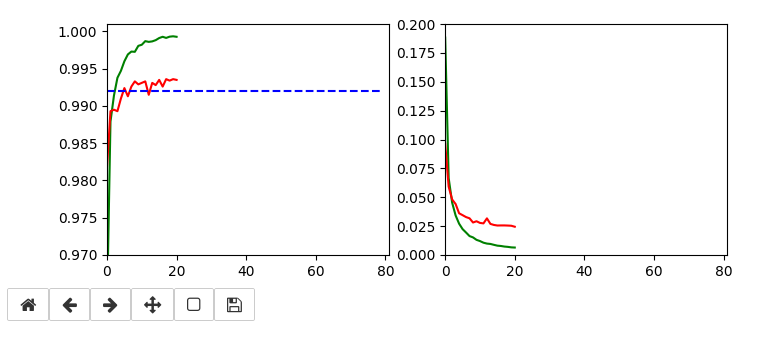
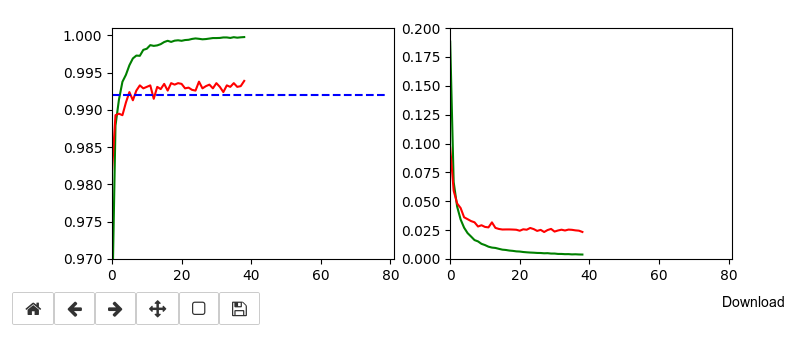
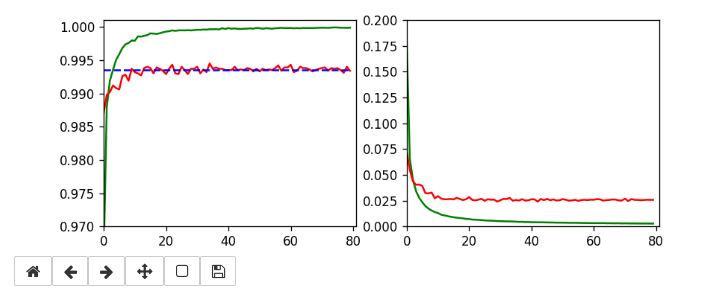
Below I show screenshots taken during training for the parameters defined above.
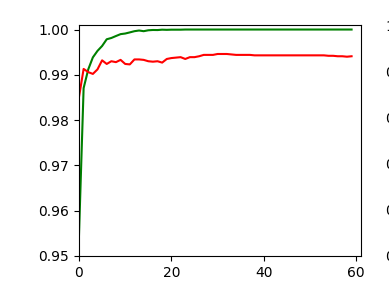
The left plot shows the achieved accuracy; red for the validation set (around 99.32%). The right plot shows the decline of the loss function relative to the original value. The green lines are for the training data, the red for the validation data.
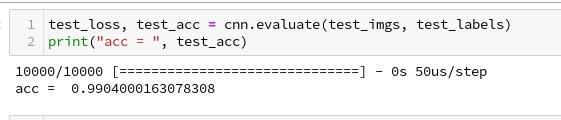
Besides the effect of seeing the data change and the plots evolve during data, we can also take the result with us that we have improved the accuracy already from 99.0% to 99.32%. When you play around with the available hyperparameters a bit you may find that 99.25% is a reproducible accuracy. But in some cases you may reach 99.4% as best values.
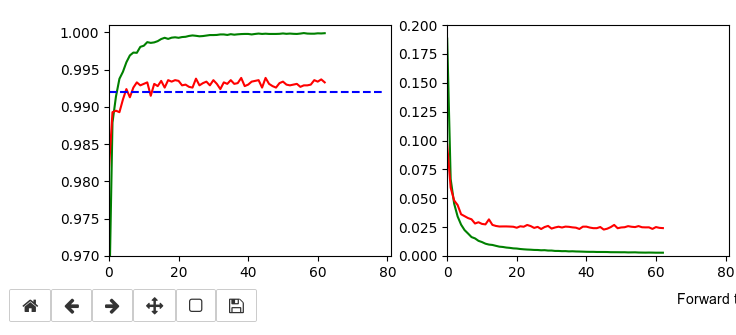
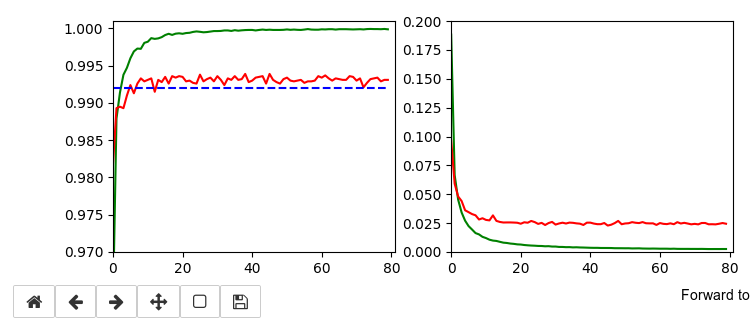
The following plots have best values around 99.4% with an average above 99.35%.
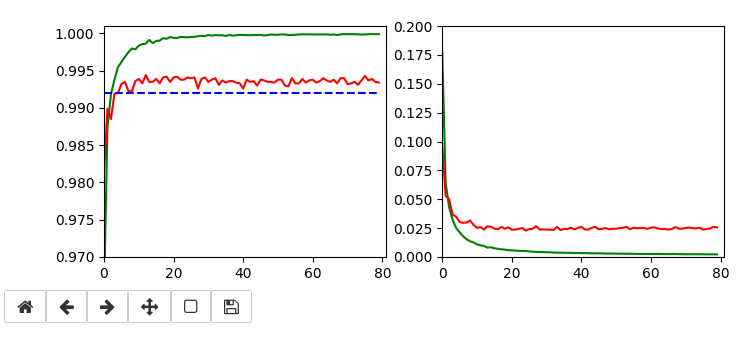
Epoch 13/80 935/938 [============================>.] - ETA: 0s - loss: 0.0096 - accuracy: 0.9986 present lr: 7.57920279e-05 present iteration: 12194 938/938 [==============================] - 6s 6ms/step - loss: 0.0096 - accuracy: 0.9986 - val_loss: 0.0238 - val_accuracy: 0.9944 Epoch 19/80 937/938 [============================>.] - ETA: 0s - loss: 0.0064 - accuracy: 0.9993 present lr: 5.3129319e-05 present iteration: 17822 938/938 [==============================] - 6s 6ms/step - loss: 0.0064 - accuracy: 0.9993 - val_loss: 0.0245 - val_accuracy: 0.9942 Epoch 22/80 930/938 [============================>.] - ETA: 0s - loss: 0.0056 - accuracy: 0.9994 present lr: 4.6219262e-05 present iteration: 20636 938/938 [==============================] - 6s 6ms/step - loss: 0.0056 - accuracy: 0.9994 - val_loss: 0.0238 - val_accuracy: 0.9942 Epoch 30/80 937/938 [============================>.] - ETA: 0s - loss: 0.0043 - accuracy: 0.9996 present lr: 3.43170905e-05 present iteration: 28140 938/938 [==============================] - 6s 6ms/step - loss: 0.0043 - accuracy: 0.9997 - val_loss: 0.0239 - val_accuracy: 0.9941 Epoch 55/80 937/938 [============================>.] - ETA: 0s - loss: 0.0028 - accuracy: 0.9998 present lr: 1. 90150222e-05 present iteration: 51590 938/938 [==============================] - 6s 6ms/step - loss: 0.0028 - accuracy: 0.9998 - val_loss: 0.0240 - val_accuracy: 0.9940 Epoch 69/80 936/938 [============================>.] - ETA: 0s - loss: 0.0025 - accuracy: 0.9999 present lr: 1.52156063e-05 present iteration: 64722 938/938 [==============================] - 6s 6ms/step - loss: 0.0025 - accuracy: 0.9999 - val_loss: 0.0245 - val_accuracy: 0.9940 Epoch 70/80 933/938 [============================>.] - ETA: 0s - loss: 0.0024 - accuracy: 0.9999 present lr: 1.50015e-05 present iteration: 65660 938/938 [==============================] - 6s 6ms/step - loss: 0.0024 - accuracy: 0.9999 - val_loss: 0.0245 - val_accuracy: 0.9940 Epoch 76/80 937/938 [============================>.] - ETA: 0s - loss: 0.0024 - accuracy: 0.9999 present lr: 1.38335554e-05 present iteration: 71288 938/938 [==============================] - 6s 6ms/step - loss: 0.0024 - accuracy: 0.9999 - val_loss: 0.0237 - val_accuracy: 0.9943
Parameters of the last run were
build = True
if cnn == None:
build = True
x_optimizer = None
batch_size=64
epochs=80
reset = True
my_loss ='categorical_crossentropy'
my_metrics =['accuracy']
my_regularizer = None
my_regularizer = 'l2'
my_reg_param_l2 = 0.0008
my_reg_param_l1 = 0.01
my_optimizer = 'rmsprop' # Present alternatives: rmsprop, nadam, adamax
my_momentum = 0.5 # momentum value
my_lr_sched = 'powerSched' # Present alternatrives: None, powerSched, exponential
#my_lr_sched = None # Present alternatrives: None, powerSched, exponential
my_lr_init = 0.001 # initial leaning rate
my_lr_decay_steps = 1 # decay steps = 1
my_lr_decay_rate = 0.001 # decay rate
li_conv_1 = [64, (3,3), 0]
li_conv_2 = [64, (3,3), 0]
li_conv_3 = [128, (3,3), 0]
li_Conv = [li_conv_1, li_conv_2, li_conv_3]
li_pool_1 = [(2,2)]
li_pool_2 = [(2,2)]
li_Pool = [li_pool_1, li_pool_2]
li_dense_1 = [120, 0]
#li_dense_2 = [30, 0]
li_dense_3 = [10, 0]
li_MLP = [li_dense_1, li_dense_2, li_dense_3]
li_MLP = [li_dense_1, li_dense_3]
input_shape = (28,28,1)
It is interesting that RMSProp only requires small values of the L2-regularizer for a sufficient stabilization – such that the loss curve for the validation data does not rise again substantially. Instead we observe an oscillation around a minimum after the learning rate as decreased sufficiently.
Addendum, 15.06.2020
An attentive reader has found out that I have cheated a bit: I have used 64 maps at the first convolutional layer instead of 32 according to the setup in the previous article. Yes, sorry. To compensate for this I include the plot of a run with 32 maps at the first convolution. The blue line marks an accuracy of 99.35%. You see that we can get above it with 32 maps, too.
The parameters were:
build = True
if cnn == None:
build = True
x_optimizer = None
batch_size=64
epochs=80
reset = False # reset the initial weight values to those saved?
my_loss ='categorical_crossentropy'
my_metrics =['accuracy']
my_regularizer = None
my_regularizer = 'l2'
my_reg_param_l2 = 0.001
#my_reg_param_l2 = 0.01
my_reg_param_l1 = 0.01
my_
optimizer = 'rmsprop' # Present alternatives: rmsprop, nadam, adamax
my_momentum = 0.9 # momentum value
my_lr_sched = 'powerSched' # Present alternatrives: None, powerSched, exponential
#my_lr_sched = None # Present alternatrives: None, powerSched, exponential
my_lr_init = 0.001 # initial leaning rate
my_lr_decay_steps = 1 # decay steps = 1
my_lr_decay_rate = 0.001 # decay rate
li_conv_1 = [32, (3,3), 0]
li_conv_2 = [64, (3,3), 0]
li_conv_3 = [128, (3,3), 0]
li_Conv = [li_conv_1, li_conv_2, li_conv_3]
li_Conv_Name = ["Conv2D_1", "Conv2D_2", "Conv2D_3"]
li_pool_1 = [(2,2)]
li_pool_2 = [(2,2)]
li_Pool = [li_pool_1, li_pool_2]
li_Pool_Name = ["Max_Pool_1", "Max_Pool_2", "Max_Pool_3"]
li_dense_1 = [120, 0]
#li_dense_2 = [30, 0]
li_dense_3 = [10, 0]
li_MLP = [li_dense_1, li_dense_2, li_dense_3]
li_MLP = [li_dense_1, li_dense_3]
input_shape = (28,28,1)
Epoch 15/80 926/938 [============================>.] - ETA: 0s - loss: 0.0095 - accuracy: 0.9988 present lr: 6.6357e-05 present iteration: 14070 938/938 [==============================] - 4s 5ms/step - loss: 0.0095 - accuracy: 0.9988 - val_loss: 0.0268 - val_accuracy: 0.9940 Epoch 23/80 935/938 [============================>.] - ETA: 0s - loss: 0.0067 - accuracy: 0.9995 present lr: 4.42987512e-05 present iteration: 21574 938/938 [==============================] - 4s 5ms/step - loss: 0.0066 - accuracy: 0.9995 - val_loss: 0.0254 - val_accuracy: 0.9943 Epoch 35/80 936/938 [============================>.] - ETA: 0s - loss: 0.0049 - accuracy: 0.9996 present lr: 2.95595619e-05 present iteration: 32830 938/938 [==============================] - 4s 5ms/step - loss: 0.0049 - accuracy: 0.9996 - val_loss: 0.0251 - val_accuracy: 0.9945
Another stupid thing, which I should have mentioned:
I have not yet found a simple way of how to explicitly reset the number of iterations, only,
- without a recompilation
- or reloading a fully saved model at iteration 0 instead of reloading only the weights
- or writing my own scheduler class based on epochs or batches.
With the present code you would have to (re-) build the model to avoid starting with a large number of “iterations” – and thus a small learning-rate in a second training run. My “reset”-parameter alone does not help.
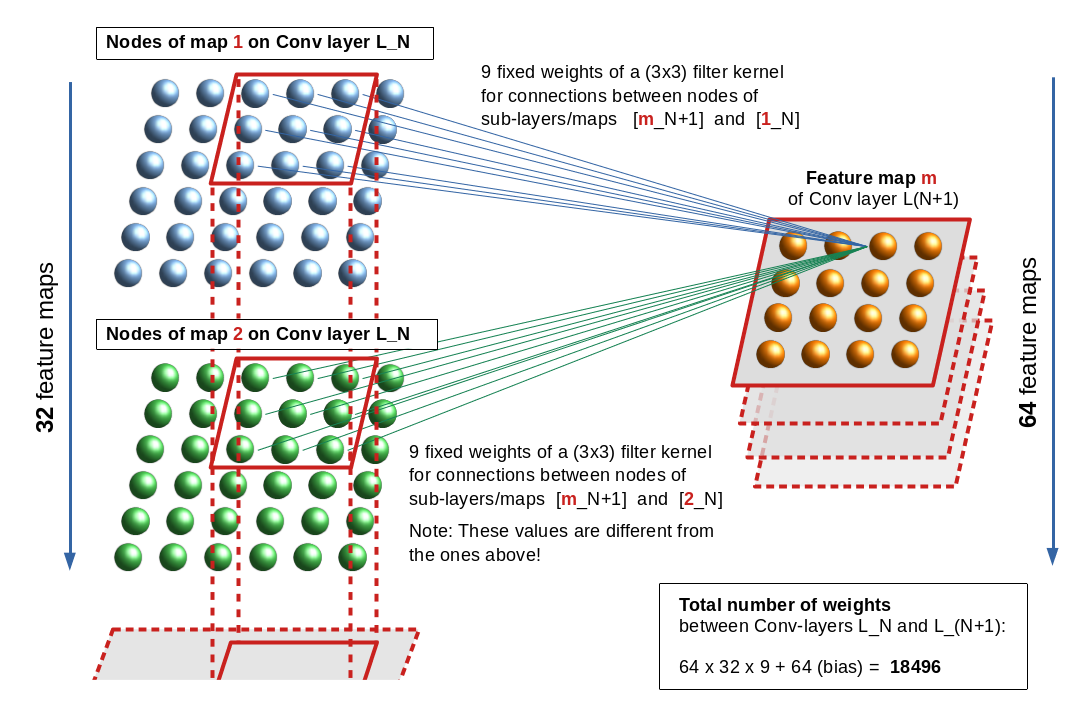
By the way: Shape of the weight matrices
Some of my readers may be tempted to have a look at the weight tensors. This is possible via the optimizer and a callback. Then they may wonder about the dimensions, which are logically a bit different from the weight matrices I have used in my code for MLPs in another article series.
In the last article of this CNN series I have mentioned that Keras takes care of configuring the weight matrices itself as soon as it got all information about the layers and layers’ nodes (or units). Now, this still leaves some open degrees of handling the enumeration of layers and matrices; it is plausible that the logic of the layer and weight association must follow a convention. E.g., you can associate a weight matrix with the receiving layer in forward propagation direction. This is what Keras and TF2 do and it is different from what I did in my MLP code. Even if you have fixed this point, then you still can discuss the row/column-layout (shape) of the weight matrix:
Keras and TF2 require the “input_shape” i.e. the shape of the tensor which is fed into the present layer. Regarding the weight matrix it is the number of nodes (units) of the previous layer (in FW direction) which is interesting – normally this is given by the 2nd dimension number of the tensors shape. This has to be combined with the number of units in the present
layer. The systematics is that TF2 and Keras define the weight matrix between two layers according to the following convention for two adjacent layers L_N and L_(N+1) in FW direction:
L_N with A nodes (6 units) and L_(N+1) with B nodes (4 units) give a shape of the weight matrix as (A,B). [(6,4)]
See the references at the bottom of the article.
Note that this is NOT what we would expect from our handling of the MLP. However, the amount of information kept in the matrix is, of course, the same. It is just a matter of convention and array transposition for the required math operations during forward and error backward propagation. If you (for reasons of handling gradients) concentrate on BW propagation then the TF2/Keras convention is quite reasonable.
Conclusion
The Keras framework gives you access to a variety of hyperparameters you can use to control the use of a regularizer, the optimizer, the decline of the learning rate with batches/epochs, the momentum. The handling is a bit unconventional, but one gets pretty fast used to it.
We once again learned that it pays to invest a bit in a variation of such parameters, when you want to fight for the last tenth of percents beyond 99.0% accuracy. In the case of MNIST we can drive accuracy to 99.35%.
With the next article of this series
we turn more to the question of visualizing some data at the convolutional layers to better understand their working.
Links
Layers and the shape of the weight matrices
https://keras.io/guides/sequential_model/
https://stackoverflow.com/questions/ 44791014/ understanding-keras-weight-matrix-of-each-layers
https://keras.io/guides/ making new layers and models via subclassing/
Learning rate and adaptive optimizers
https://towardsdatascience.com/learning-rate-schedules-and-adaptive-learning-rate-methods-for-deep-learning-2c8f433990d1
https://machinelearningmastery.com/ understand-the-dynamics-of-learning-rate-on-deep-learning-neural-networks/
https://faroit.com/keras-docs/2.0.8/optimizers/
https://keras.io/api/optimizers/
https://keras.io/api/optimizers/ learning_rate_schedules/
https://www.jeremyjordan.me/nn-learning-rate/
Keras Callbacks
https://keras.io/api/callbacks/
https://keras.io/guides/ writing_your_own_callbacks/
https://keras.io/api/callbacks/ learning_rate_scheduler/
Interactive plotting
https://stackoverflow.com/questions/ 39428347/ interactive-matplotlib-plots-in-jupyter-notebook
https://matplotlib.org/3.2.1/api/ _as_gen/ matplotlib.pyplot.ion.html
https://matplotlib.org/3.1.3/tutorials/ introductory/ usage.html
Further articles in this series
A simple CNN for the MNIST dataset – XI – Python code for filter visualization and OIP detection
A simple CNN for the MNIST dataset – X – filling some gaps in filter visualization
A simple CNN for the MNIST dataset – IX – filter visualization at a convolutional layer
A simple CNN for the MNIST dataset – VIII – filters and features – Python code to visualize patterns which activate a map strongly
A simple CNN for the MNIST dataset – VII – outline of steps to visualize image patterns which trigger filter maps
A simple CNN for the MNIST dataset – VI – classification by activation patterns and the role of the CNN’s MLP part
A simple CNN for the MNIST dataset – V – about the difference of activation patterns and features